Advancing Science with DNA Sequence Microbial Genome Annotation

Advancing Science with DNA Sequence Microbial Genome Annotation Nikos Kyrpides DOE Joint Genome institute

Advancing Science with DNA Sequence Two main goals of genome analysis: • Evolutionary analysis – How does an organism compare to the rest? • Metabolic reconstruction – What can an organism do and how?
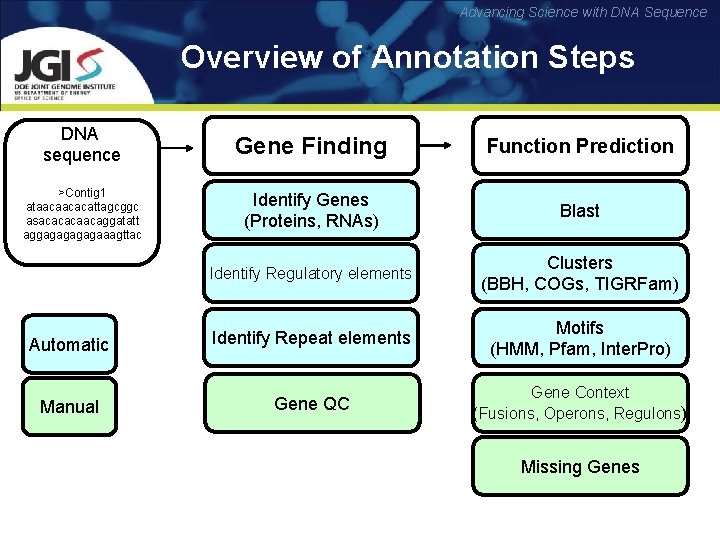
Advancing Science with DNA Sequence Overview of Annotation Steps DNA sequence Gene Finding Function Prediction >Contig 1 ataacaacacattagcggc asacacacaacaggatatt aggagagaaagttac Identify Genes (Proteins, RNAs) Blast Identify Regulatory elements Clusters (BBH, COGs, TIGRFam) Automatic Identify Repeat elements Motifs (HMM, Pfam, Inter. Pro) Manual Gene QC Gene Context (Fusions, Operons, Regulons) Missing Genes

Advancing Science with DNA Sequence 1. Finding the genes in microbial genomes 1. Introduction 2. Tools out there 3. Basic principles behind tools 4. Known problems of the tools: why you may need manual curation

Advancing Science with DNA Sequence Finding the genes in microbial genomes features Sequence features in prokaryotic genomes: § stable RNA-coding genes (r. RNAs, t. RNAs, RNA component of RNase. P, tm. RNA) § protein-coding genes (CDSs) § transcriptional features (m. RNAs, operons, promoters, terminators, protein-binding sites, DNA bends) § translational features (RBS, regulatory antisense RNAs, m. RNA secondary structures, translational recoding and programmed frameshifts, inteins) § pseudogenes (t. RNA and protein-coding genes) §…
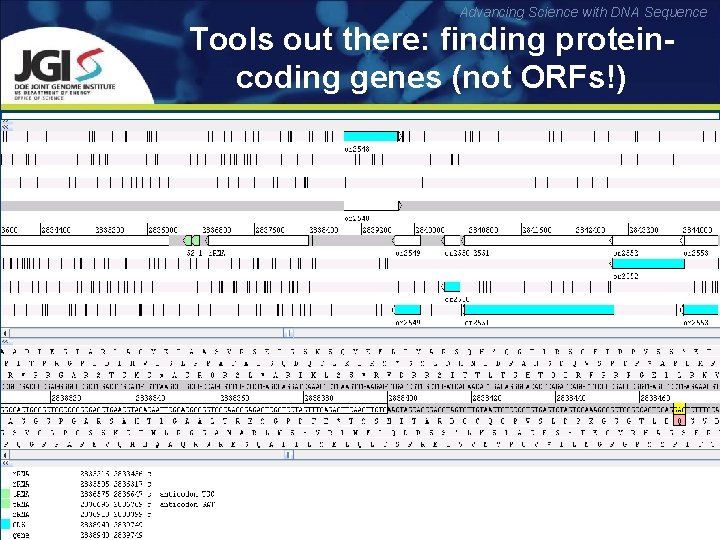
Advancing Science with DNA Sequence Tools out there: finding proteincoding genes (not ORFs!) Reading frames: translations of the nucleotide sequence with an offset of 0, 1 and 2 nucleotides (three possible translations in each direction) Open reading frame (ORF): reading frame between a start and stop codon
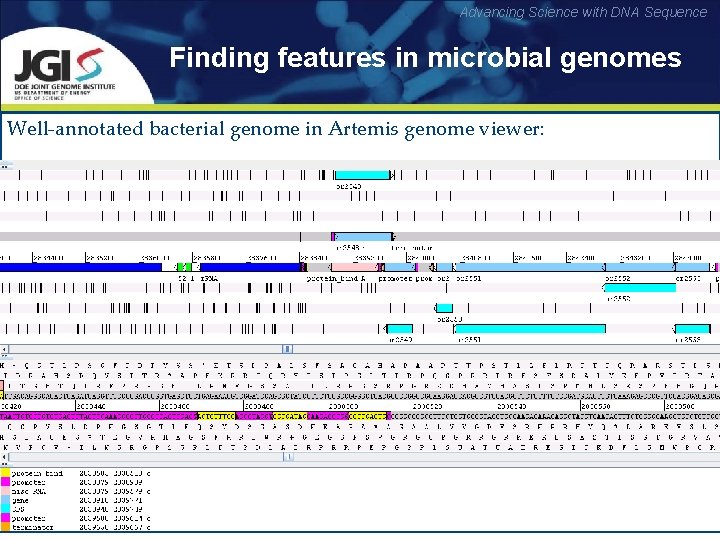
Advancing Science with DNA Sequence Finding features in microbial genomes Well-annotated bacterial genome in Artemis genome viewer:

Advancing Science with DNA Sequence Finding the genes in microbial genomes 1. Introduction 2. Tools out there 3. Basic principles behind tools 4. Known problems of the tools: why you may need manual curation

Advancing Science with DNA Sequence Tools out there: servers for microbial genome annotation - I • IMG-ER http: //img. jgi. doe. gov/er IMG-ER submission page: http: //img. jgi. doe. gov/submit • RAST http: //rast. nmpdr. org/ • JCVI Annotation Service Output: üstable RNA-encoding genes, üCDSs, üfunctional annotations üoutput in Gen. Bank format Output: ür. RNAs and t. RNAs, üCDSs, üfunctional annotations üoutput in several formats http: //www. jcvi. org/cms/research/projects/annotation-service/overview/ Output: üCDSs, stable RNAs? üfunctional annotations üformat?

Advancing Science with DNA Sequence Tools out there: servers for microbial genome annotation - II • AMIGENE http: //www. genoscope. cns. fr/agc/tools/amiga/Form/form. php • Ref. Seq Output: CDSs, output in gff format http: //www. ncbi. nlm. nih. gov/genomes/MICROBES/genemark. cgi http: //www. ncbi. nlm. nih. gov/genomes/MICROBES/glimmer_3. cgi • Easy. Gene Output: CDSs, output in tbl format http: //www. cbs. dtu. dk/services/Easy. Gene/ Output: CDSs, size restriction <1 Mb
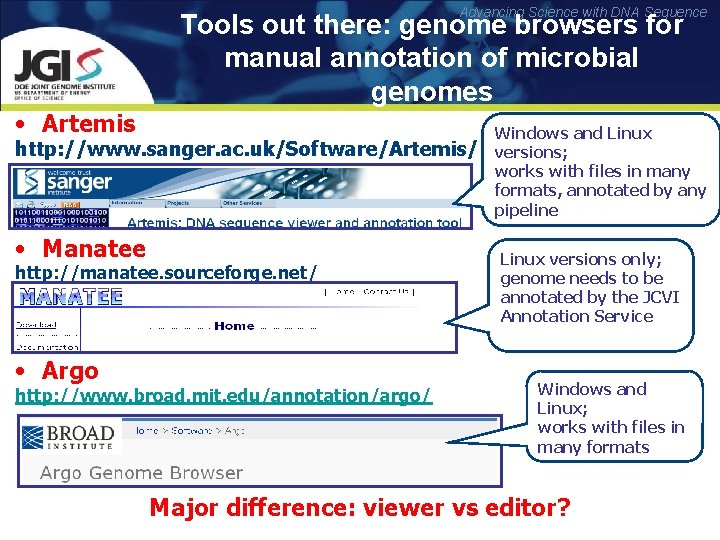
Advancing Science with DNA Sequence Tools out there: genome browsers for manual annotation of microbial genomes • Artemis http: //www. sanger. ac. uk/Software/Artemis/ • Manatee http: //manatee. sourceforge. net/ • Argo http: //www. broad. mit. edu/annotation/argo/ Windows and Linux versions; works with files in many formats, annotated by any pipeline Linux versions only; genome needs to be annotated by the JCVI Annotation Service Windows and Linux; works with files in many formats Major difference: viewer vs editor?

Advancing Science with DNA Sequence Tools out there: tools for finding stable (“non-coding”) RNAs - I • Large structural RNAs (23 S and 16 S r. RNAs) RNAmmer http: //www. cbs. dtu. dk/services/RNAmmer/ • Small structural RNAs (5 S r. RNA, t. RNAs, tm. RNA, RNase. P RNA component) Rfam database, INFERNAL search tool http: //www. sanger. ac. uk/Software/Rfam/ http: //rfam. janelia. org/ http: //infernal. janelia. org/ Web service: sequence search is limited to 2 kb ARAGORN Web service: sequence http: //130. 235. 46. 10/ARAGORN 1. 1/HTML/aragorn 1. 2. html t. RNAScan-SE http: //lowelab. ucsc. edu/t. RNAscan-SE/ search is limited to 15 kb, finds t. RNAs and tm. RNAs only Web service: sequence search is limited to 5 Mb, finds t. RNAs only

Advancing Science with DNA Sequence Tools out there: tools for finding “non-coding” RNAs - II • Short regulatory RNAs Rfam database, INFERNAL search tool http: //www. sanger. ac. uk/Software/Rfam/ http: //rfam. janelia. org/ http: //infernal. janelia. org/ Web service: sequence search is limited to 2 kb; Provides list of precalculated RNAs for publicly available genomes Other (less popular) tools: Pipeline for discovering cis-regulatory nc. RNA motifs: http: //bio. cs. washington. edu/supplements/yzizhen/pipeline/ RNAz http: //www. tbi. univie. ac. at/~wash/RNAz/

Advancing Science with DNA Sequence Tools out there: most popular CDS-finding tools • CRITICA • Glimmer family (Glimmer 2, Glimmer 3, RBS finder) http: //glimmer. sourceforge. net/ • Gene. Mark family (Gene. Mark-hmm, Gene. Mark. S) http: //exon. gatech. edu/Gene. Mark/ • Easy. Gene • AMIGENE • PRODIGAL (default JGI gene finder) http: //compbio. ornl. gov/prodigal/ Combinations and variations of the above • RAST (Glimmer 2 + pre- and post-processing)

Advancing Science with DNA Sequence Basic principles: finding CDSs using evidence-based vs ab initio algorithms Two major approaches to prediction of protein-coding genes: • “evidence-based” (ORFs with translations homologous to the known proteins are CDSs) Advantages: finds “unusual” genes (e. g. horizontally transferred); relatively low rate of false positive predictions Limitations: cannot find “unique” genes; low sensitivity on short genes; prone to propagation of false positive results of ab initio annotation tools • ab initio (ORFs with nucleotide composition similar to CDSs are also CDSs) Advantages: finds “unique” genes; high sensitivity Limitations: often misses “unusual” genes; high rate of false positives

Advancing Science with DNA Sequence Finding the genes in microbial genomes 1. Introduction 2. Tools out there 3. Basic principles behind tools 4. Known problems of the tools: why you may need manual curation

Advancing Science with DNA Sequence Known problems: CDSs § Short CDSs: many are missed, others are overpredicted ü short ribosomal proteins (30 -40 aa long) are often missed ü short proteins in the promoter region are often overpredicted § N-terminal sequences are often inaccurate (many features of the sequence around start codon are not accounted for) ü Glimmer 2. 0 is calling genes longer than they should be ü Gene. Mark, Glimmer 3. 0 mostly call genes shorter § Pseudogenes and sequencing errors (artificial frameshift) ü all tools are looking for ORFs (needs valid start and stop codons) ü “unique” genes are often predicted on the opposite strand of a pseudogene or a gene with a sequencing error § Proteins with unusual translational features (recoding, programmed frameshifts) ü these genes are often mistaken for pseudogenes ü see pseudogenes

Advancing Science with DNA Sequence Known problems: CDSs Lack of Standards
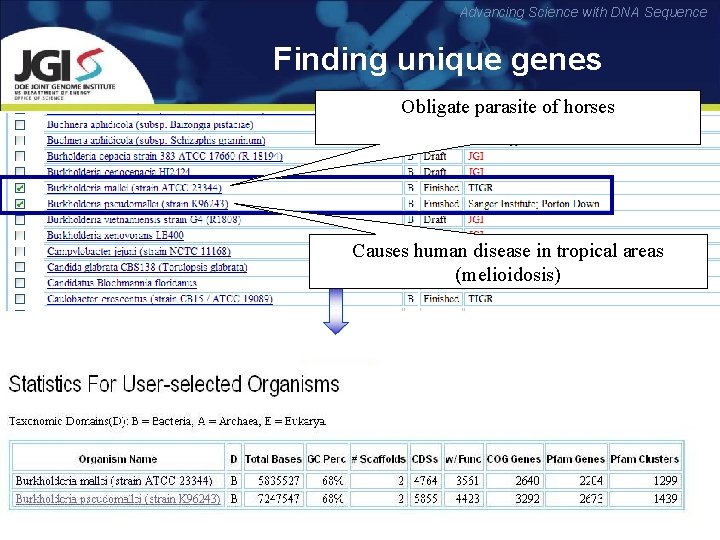
Advancing Science with DNA Sequence Finding unique genes Obligate parasite of horses Causes human disease in tropical areas (melioidosis)

Advancing Science with DNA Sequence • Phylogenetic profiler finds 548 unique genes in B. mallei • However, 497 of them in fact exist in B. pseudomallei, but they have not been called as real genes. The difference in gene models reveals 89. 2% error rate in unique genes •
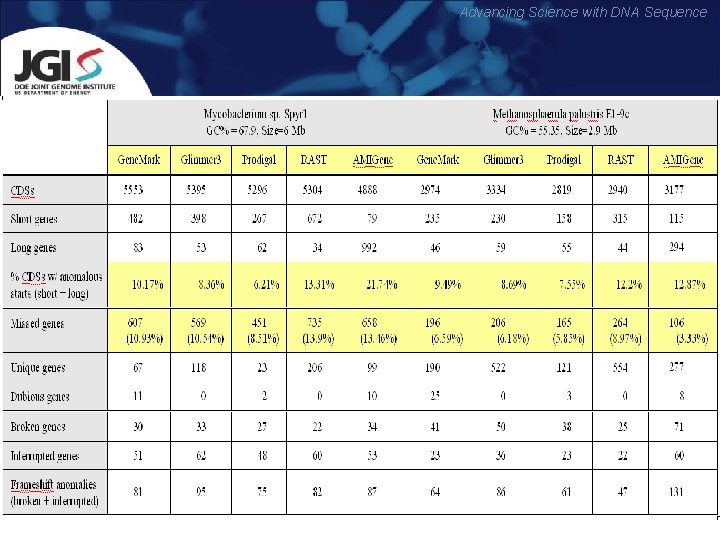
Advancing Science with DNA Sequence
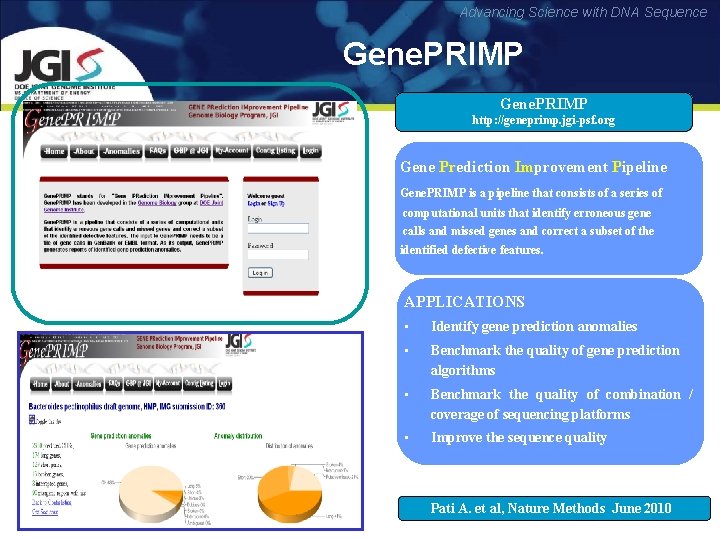
Advancing Science with DNA Sequence Gene. PRIMP http: //geneprimp. jgi-psf. org Gene Prediction Improvement Pipeline Pr Im Gene. PRIMP is a pipeline that consists of a series of computational units that identify erroneous gene calls and missed genes and correct a subset of the identified defective features. APPLICATIONS • Identify gene prediction anomalies • Benchmark the quality of gene prediction algorithms • Benchmark the quality of combination / coverage of sequencing platforms • Improve the sequence quality Pati A. et al, Nature Methods June 2010
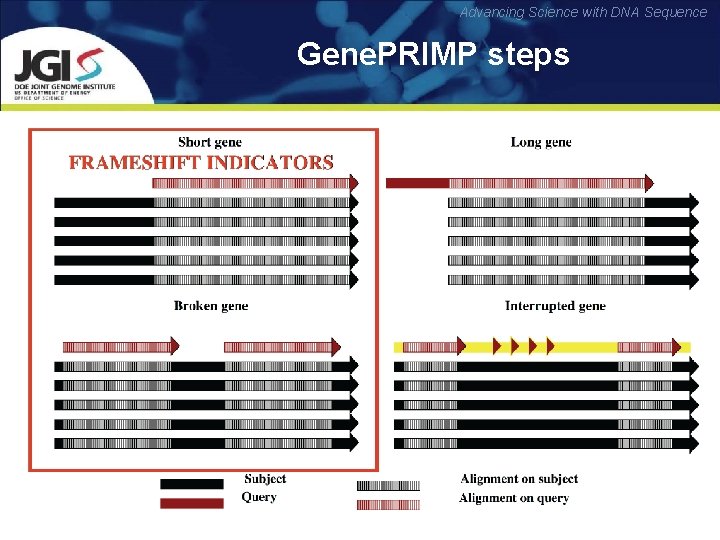
Advancing Science with DNA Sequence Gene. PRIMP steps
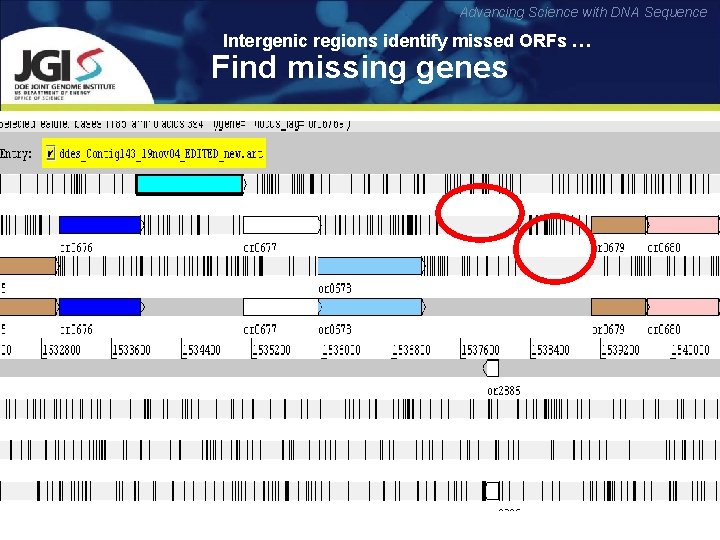
Advancing Science with DNA Sequence Intergenic regions identify missed ORFs … Find missing genes
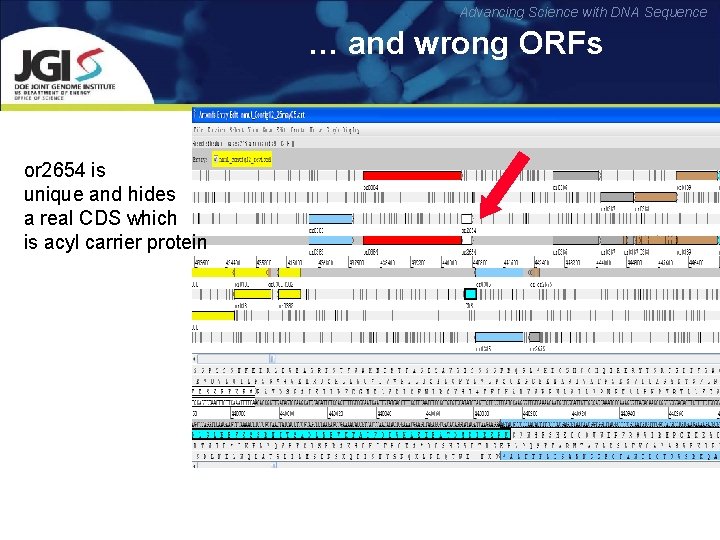
Advancing Science with DNA Sequence … and wrong ORFs or 2654 is unique and hides a real CDS which is acyl carrier protein
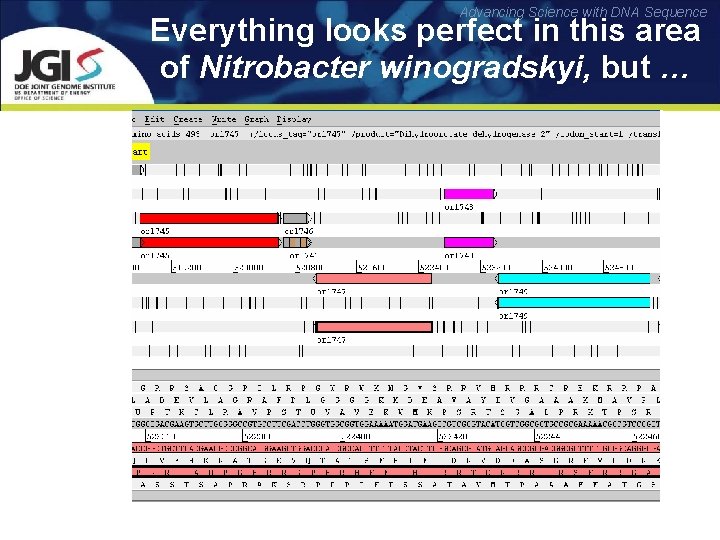
Advancing Science with DNA Sequence Everything looks perfect in this area of Nitrobacter winogradskyi, but …

Advancing Science with DNA Sequence … hides a real ORF

Guinness Book of protein-coding genes Advancing Science with DNA Sequence ü The longest human gene is 2, 220, 223 nucleotides long. It has 79 exons, with a total of only 11, 058 nucleotides, which specify the sequence of the 3, 685 amino acids and codes for a protein dystrophin. It is part of a protein complex located in the cell membrane, which transfers the force generated by the actinmyosin structure inside the muscle fiber to the entire fiber. ü The smallest human gene is 252 nucleotides long, it specifies a polypeptide of 67 amino acids and codes for an insulin-like growth factor II. ü The longest bacterial gene is 110, 418 nucleotides long, which specify the sequence of 36, 805 amino acids. Its function is unknown, most likely a surface protein. ü The smallest bacterial gene is 54 nucleotides long, it specifies a polypeptide of 17 amino acids and codes for a regulatory protein in cyanobacteria

Advancing Science with DNA Sequence False positives Genome name CDSs with no hits < 100 aa % with t. BLASTn hit % t. BLASTn hits with frameshifts/stop codons Prochlorococcus AS 9601 18 88. 9 68. 8 Prochlorococcus MIT 9211 62 40. 3 80 Prochlorococcus MIT 9215 24 58. 3 64. 2 Prochlorococus MIT 9301 12 75 66. 7 Prochlorococcus MIT 9303 501 83 61. 8 Prochlorococcus MIT 9313 35 8. 6 66. 7 Prochlrococcus MIT 9515 32 81. 3 50 Prochlorococcus NATL 1 A 209 95. 2 48. 2 Prochlorococcus CCMP 1375 50 34 82. 4 Synechococcus PCC 7942 39 0 0 Synechococcus CC 9311 313 11. 5 83. 3 Synethococcus CC 9605 83 38. 6 81. 3 Synechococcus CC 9902 21 57. 1 100 Synechococcus JA-2 -3 Ba 176 26. 7 85. 1 Synechococcus JA-3 -3 Ab 142 35. 2 92 Synechococcus PCC 7002 93 17. 2 56. 3 Synechococcus RCC 307 184 10. 3 68. 4 S Synechococcus WH 7803 32 18. 8 83. 3 Synechococcus WH 8102 39 38. 4 46. 7

Advancing Science with DNA Sequence 2. Finding the functions in microbial genomes 1. Introduction 2. Tools out there 3. Known problems

Advancing Science with DNA Sequence what is function? cobalamin biosynthetic enzyme, cobalt-precorrin-4 methyltransferase (Cbi. F) • molecular/enzymatic (methyltransferase) – Reaction (methylation) – Substrate (cobalt-precorrin-4) – Ligand (S-adenosyl-L-methionine) • metabolic (cobalamin biosynthesis) • physiological (maintenance of healthy nerve and red blood cells, through B 12).

Advancing Science with DNA Sequence Functional characterization

Advancing Science with DNA Sequence Computational approaches to Functional characterization
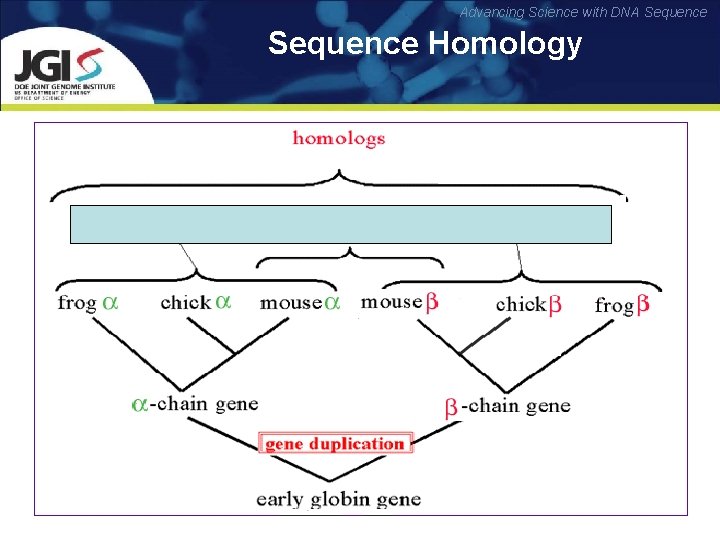
Advancing Science with DNA Sequence Homology Two sequences are homologous, if there existed a molecule in the past that is ancestral to both of the sequences. Types of Homology: üOrthology: bifurcation in molecular tree reflects speciation üParalogy: bifurcation in molecular tree reflects gene duplication

Advancing Science with DNA Sequence Homology & analogy • The term homology is confounded & abused in the literature! – sequences are homologous if they’re related by divergence from a common ancestor – analogy relates to the acquisition of common features from unrelated ancestors via convergent evolution • e. g. , b-barrels occur in soluble serine proteases & integral membrane porins; chymotrypsin & subtilisin share groups of catalytic residues, with near identical spatial geometries, but no other similarities • Homology is not a measure of similarity & is not quantifiable – it is an absolute statement that sequences have a divergent rather than a convergent relationship – the phrases "the level of homology is high" or "the sequences show 50% homology", or any like them, are strictly meaningless!
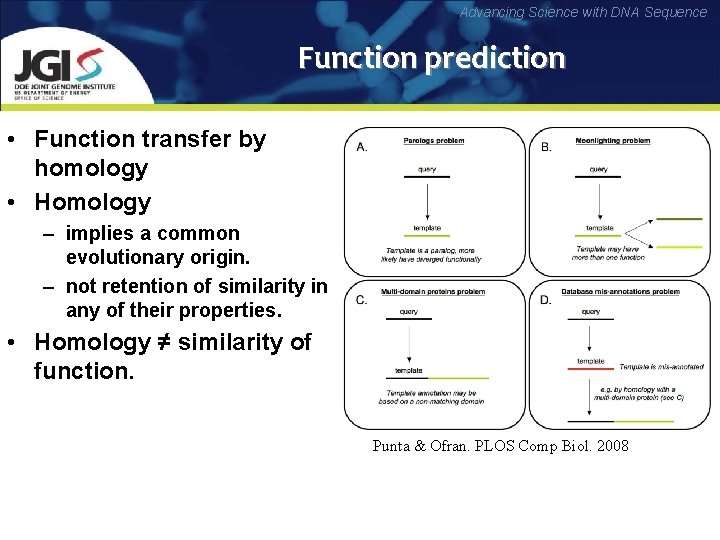
Advancing Science with DNA Sequence Function prediction • Function transfer by homology • Homology – implies a common evolutionary origin. – not retention of similarity in any of their properties. • Homology ≠ similarity of function. Punta & Ofran. PLOS Comp Biol. 2008
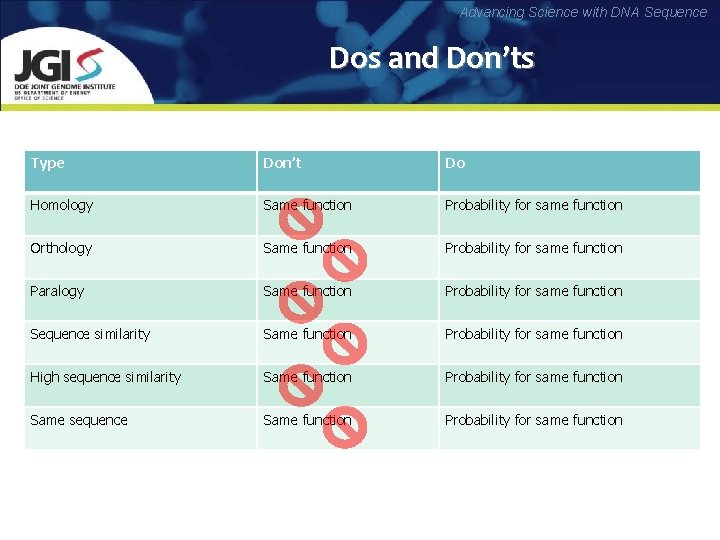
Advancing Science with DNA Sequence Dos and Don’ts Type Don’t Do Homology Same function Probability for same function Orthology Same function Probability for same function Paralogy Same function Probability for same function Sequence similarity Same function Probability for same function High sequence similarity Same function Probability for same function Same sequence Same function Probability for same function

Advancing Science with DNA Sequence Application areas of analysis tools • • The scale indicates % identity between aligned sequences Alignment of 2 random seqs can produce ~20% identity – less than 20% does not constitute a significant alignment – around this threshold is the Twilight Zone, where alignments may appear plausible to the eye, but can’t be proved by conventional methods

Advancing Science with DNA Sequence Finding the functions in microbial genomes 1. Introduction 2. Tools out there 3. Known problems

Advancing Science with DNA Sequence Function prediction • Similarity searches (BLAST). • Domain identification(Pfam). • Small sequence identification(PROSITE).
Advancing Science with DNA Sequence What if nothing is similar ? • • Subcellular localization Gene context Structure Prediction of binding residues (DISIS, bind. N) S~S Periplasm Cytoplasm
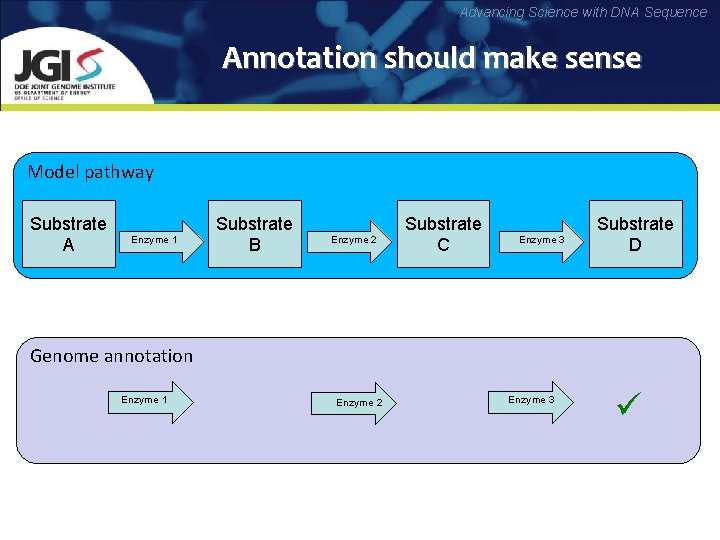
Advancing Science with DNA Sequence Annotation should make sense Model pathway Substrate A Enzyme 1 Substrate B Enzyme 2 Substrate C Enzyme 3 Substrate D Genome annotation ? Enzyme 1 Enzyme 2 ? Enzyme 3 ü

Advancing Science with DNA Sequence Annotation should make sense

Advancing Science with DNA Sequence Databases • Databases used for the analysis of biological molecules. • Databases contain information organized in a way that allows users/researchers to retrieve and exploit it. • Why bother? – – – Store information. Organize data. Predict features (genes, functions. . . ). Predict the functional role of a feature (annotation). Understand relationships (metabolic reconstruction).
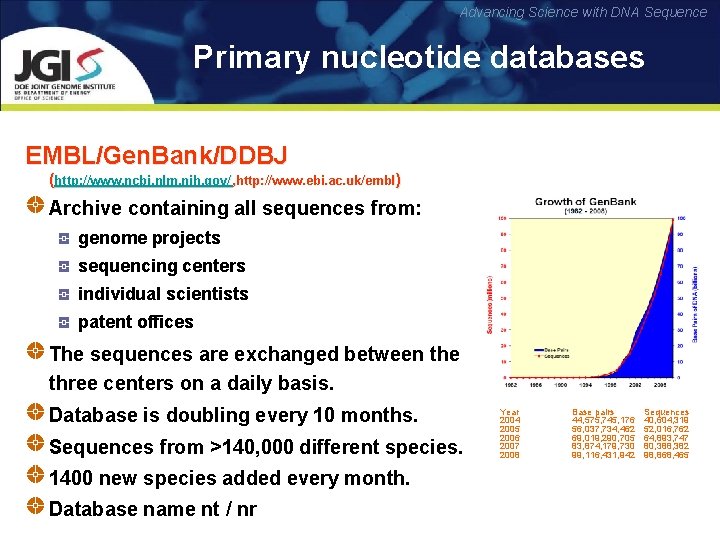
Advancing Science with DNA Sequence Primary nucleotide databases EMBL/Gen. Bank/DDBJ (http: //www. ncbi. nlm. nih. gov/ , http: //www. ebi. ac. uk/embl ) Archive containing all sequences from: genome projects sequencing centers individual scientists patent offices The sequences are exchanged between the three centers on a daily basis. Database is doubling every 10 months. Sequences from >140, 000 different species. 1400 new species added every month. Database name nt / nr Year 2004 2005 2006 2007 2008 Base pairs 44, 575, 745, 176 56, 037, 734, 462 69, 019, 290, 705 83, 874, 179, 730 99, 116, 431, 942 Sequences 40, 604, 319 52, 016, 762 64, 893, 747 80, 388, 382 98, 868, 465

Advancing Science with DNA Sequence Primary protein sequence databases • Contain coding sequences derived from the translation of nucleotide sequences – Gen. Bank • Valid translations (CDS) from nt Gen. Bank entries. – Uni. Prot. KB/Tr. EMBL (1996) • Automatic CDS translations from EMBL. • Tr. EMBL Release 40. 3 (26 -May-2009) contains 7, 916, 844 entries.

Advancing Science with DNA Sequence Ref. Seq Curated transcripts and proteins. reviewed by NCBI staff. Model transcripts and proteins. generated by computer algorithms. Assembled Genomic Regions (contigs). Chromosome records.

Advancing Science with DNA Sequence Classification databases Groups (families/clusters) of proteins based on… Overall sequence similarity. Local sequence similarity. Presence / absence of specific features. Structural similarity. . These groups contain proteins with similar properties. Specific function, enzymatic activity. Broad function. Evolutionary relationship. …
Advancing Science with DNA Sequence Overall sequence similarity
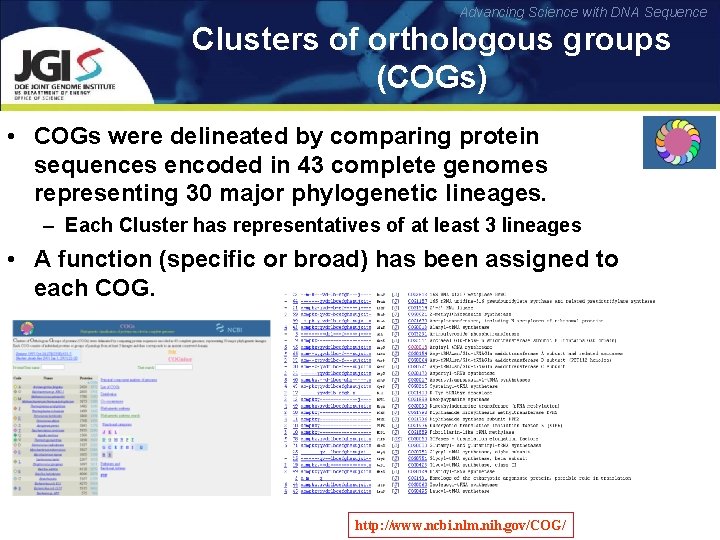
Advancing Science with DNA Sequence Clusters of orthologous groups (COGs) • COGs were delineated by comparing protein sequences encoded in 43 complete genomes representing 30 major phylogenetic lineages. – Each Cluster has representatives of at least 3 lineages • A function (specific or broad) has been assigned to each COG. http: //www. ncbi. nlm. nih. gov/COG/
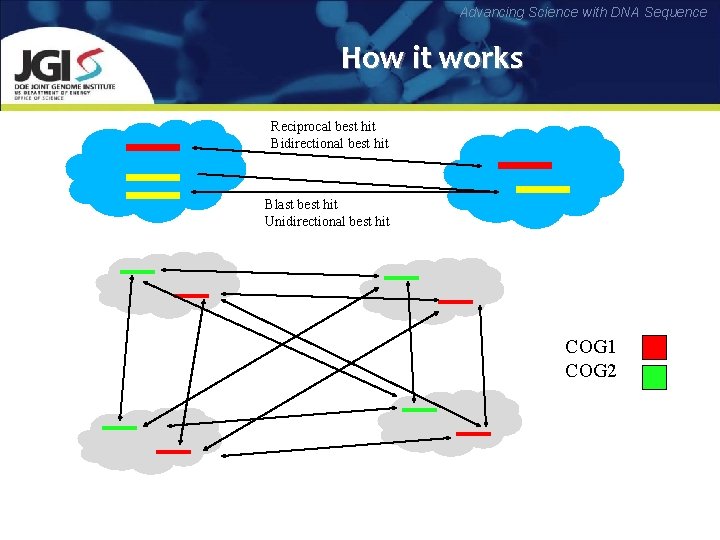
Advancing Science with DNA Sequence How it works Reciprocal best hit Bidirectional best hit Blast best hit Unidirectional best hit COG 1 COG 2

Advancing Science with DNA Sequence Profiles & Pfam • A method for classifying proteins into groups exploits region similarities, which contain valuable information (domains/profiles). • These domains/profiles can be used to detect distant relationships, where only few residues are conserved.
Advancing Science with DNA Sequence Regions similarity

Advancing Science with DNA Sequence Pfam http: //pfam. sanger. ac. uk HMMs of protein alignments (local) for domains, or global (cover whole protein)
![Advancing Science with DNA Sequence PROSITE http: //au. expasy. org/prosite/ R-Y-x-[DT]-W-x-[LIVM]-[ST]-T-P-[LIVM](3) Advancing Science with DNA Sequence PROSITE http: //au. expasy. org/prosite/ R-Y-x-[DT]-W-x-[LIVM]-[ST]-T-P-[LIVM](3)](http://slidetodoc.com/presentation_image_h2/0c4a0fff210f079d053c040e5e11e96f/image-55.jpg)
Advancing Science with DNA Sequence PROSITE http: //au. expasy. org/prosite/ R-Y-x-[DT]-W-x-[LIVM]-[ST]-T-P-[LIVM](3)
Advancing Science with DNA Sequence KEGG orthology

Advancing Science with DNA Sequence Composite pattern databases • To simplify sequence analysis, the family databases are being integrated to create a unified annotation resource – Inter. Pro – – Release 32. 0 (Apr 11) contains 21516 entries Central annotation resource, with pointers to its satellite dbs http: //www. ebi. ac. uk/interpro/
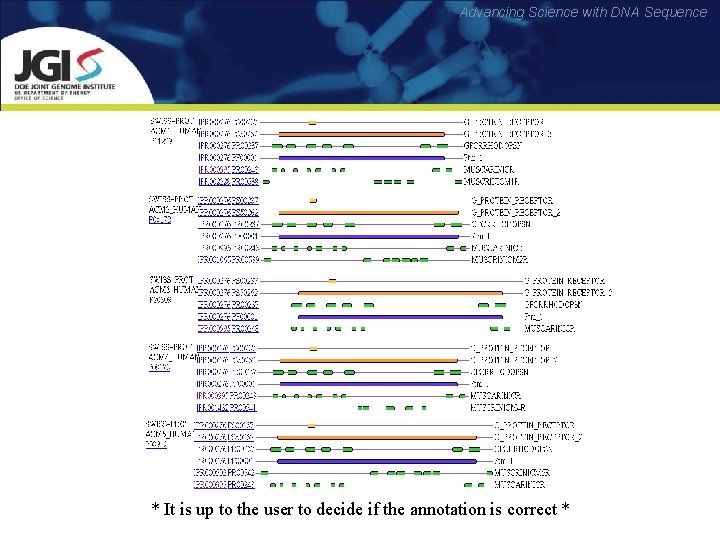
Advancing Science with DNA Sequence * It is up to the user to decide if the annotation is correct *
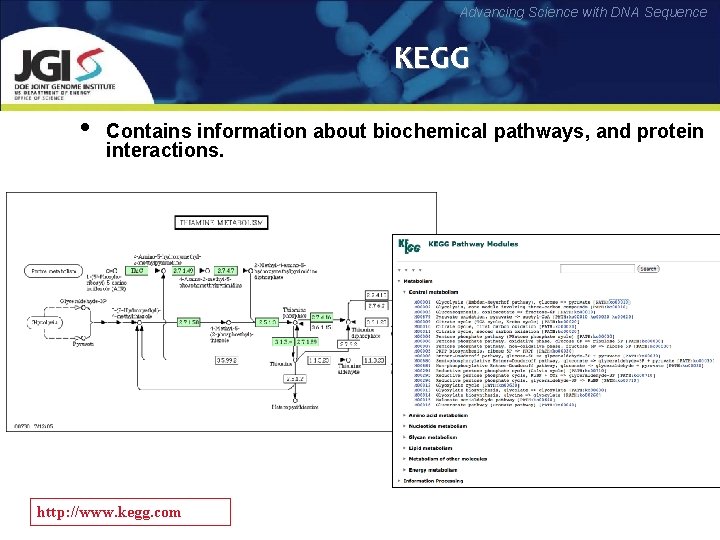
Advancing Science with DNA Sequence KEGG • Contains information about biochemical pathways, and protein interactions. http: //www. kegg. com

Advancing Science with DNA Sequence Summary • We have main archives (Genbank), and currated databases (Refseq, Swiss. Prot), and protein classification database (COG, Pfam). • This is the tip of the iceberg of databases. • They help predict the function, or the network of functions. • Systems that integrate the information from several databases, visualize and allow handling of data in an intuitive way are required
Advancing Science with DNA Sequence Functional annotation in IMG • Automated protein product assignment pipeline • Functional context in IMG • KEGG Pathways, Modules, KEGG Orthology • Meta. Cyc Pathways • IMG Pathways No longer maintained: • TIGR Role Categories • TIGR Genome Properties • COG Functional Categories
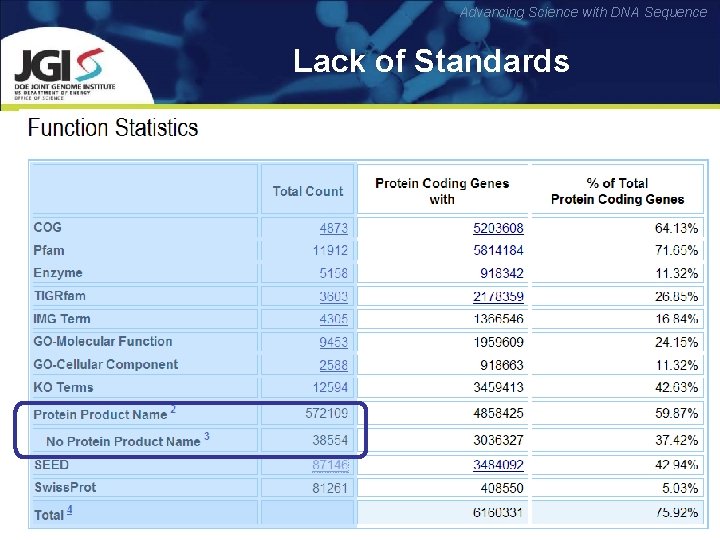
Advancing Science with DNA Sequence Lack of Standards
- Slides: 62