Advances in Biology and Pathophysiology of Multiple Myeloma

Advances in Biology and Pathophysiology of Multiple Myeloma Amer G. Rassam, MD

History of Multiple Myeloma Ø First case, a London grocer “Thomas Alexander Mc. Bean” Ø Jumped from a cave in 1844 Ø According to Drs. Thomas Watson and William Mac. Intyre, Mr. Mc. Bean had “Mollities et Fragilitas Ossium” Ø Mr. Mc. Bean died on New Year’s day in 1846
History of Multiple Myeloma Ø Urine sample presented to “Henry Bence Jones” Ø Large amount of protein was found in the sample Ø The protein has became known as Bence Jones Protein

History of Multiple Myeloma In 1890 s, Paul Unna and Ramon Cajal identified the plasma cell as a cell type and the cause of Multiple Myeloma Paul Gerson Unna Santiago Ramon Y. Cajal 1850 -1929 1852 -1934

History of Multiple Myeloma Ø In 1873, Rustizky introduced the name Multiple Myeloma Ø In 1922, Bayne-Jones and Wilson identified 2 distinct groups of Bence Jones protein Ø In 1956, Korngold and Lipari identified the relationship between Bence Jones protein and serum proteins

Epidemiology of Multiple Myeloma Ø Prevalence (at any one time) : 40000 Ø Incidence: 14000 diagnosed each year Ø Median age: 65 Ø Median survival: 33 months Ø M: F 53: 47 Ø 1. 1% of all cancer diagnosis Ø 2% of all cancer deaths
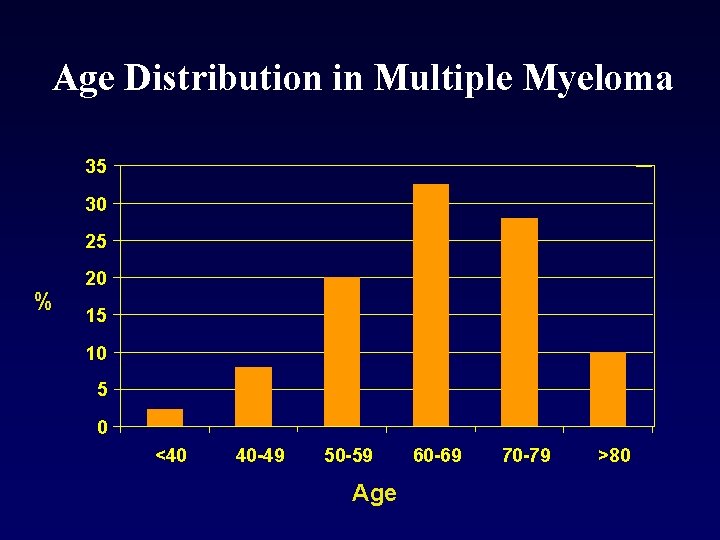
Age Distribution in Multiple Myeloma 35 30 25 % 20 15 10 5 0 <40 40 -49 50 -59 Age 60 -69 70 -79 >80

Monoclonal Gammopathies – Mayo clinic Extramedullary 1% (8) SMM 4% (39) Macro 3% (30) Other 3% (33) LP 3% (37) AL 8% (90) MGUS 62% (659) MM 16% (172)

Immunophenotype of Multiple Myeloma Marker Features CD 10 Subset CD 19 & CD 20 Rarely expressed CD 28 & CD 86 Occurs with progressive disease CD 34 Not expressed by malignant clone CD 38 High expression of most but not all malignant cells CD 56 (N-CAM) Absent in MGUS and PCL CD 138 Syndecan-1 is over expressed
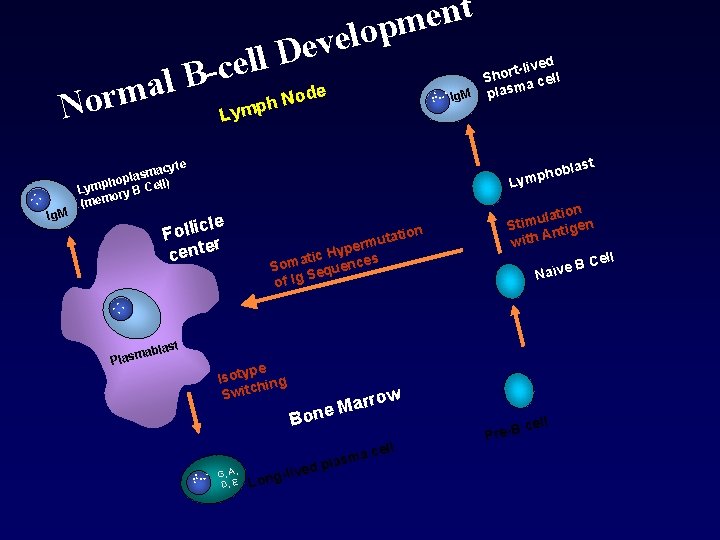
D l l e c B l a Ig. M lived t r o Sh cell a m. s. pla : : . Ig. M ode N h Lymp last b o h Lymp te macy s a l p ho l) Lymp ry B Cel o (mem le Follic r cente on utati m r e p ic Hy ces t a m So quen e S g of I tion a l u m Sti tigen n A h wit e. B Na ï v : : Norm t n e m p o l eve t ablas Plasm pe Isoty ing h Switc ow arr M e n Bo l a cel . . : : . G, A, D, E Long liv asm ed pl Pre-B cell Cell
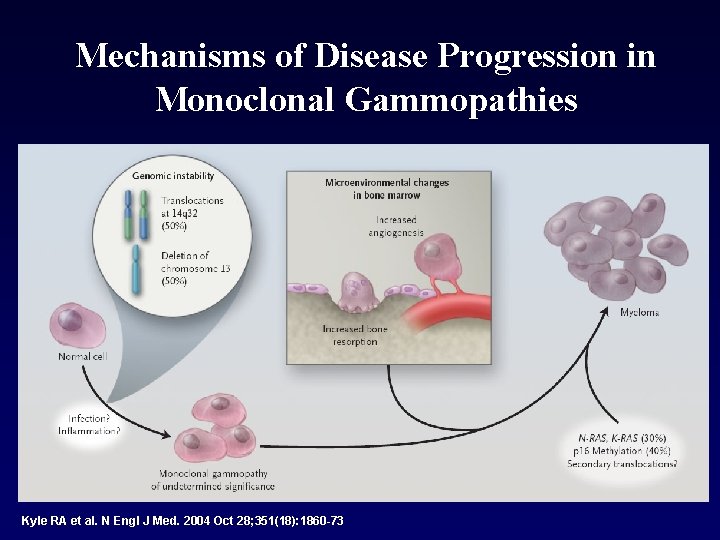
Mechanisms of Disease Progression in Monoclonal Gammopathies Kyle RA et al. N Engl J Med. 2004 Oct 28; 351(18): 1860 -73

Chromosomal Abnormalities in MM Translocations (listed in order of frequency) 14 q 32 with 11 q 13 (cyclin D, other new fibroblastic growth factors) 4 p 16 (FGFR 3) 6 p 25 (Interferon regulatory factor 4) 16 q 23 (C-MAF transcription factor) 8 q 24 (C-MYC) 18 q 21 (BCL-2) 1 q with 5, 8, 12, 14, 15, 16, 17, 19 q, 21, 22 Losses 6 q, 13 q Gains 3, 5, 7, 9 q, 11 q, 12 q, 15 q, 17 q, 18, 19, 21, 22 q
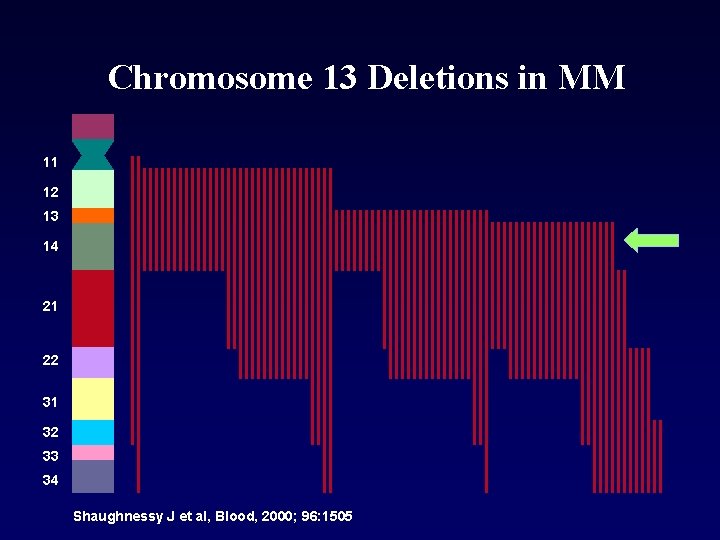
Chromosome 13 Deletions in MM 11 12 13 14 21 22 31 32 33 34 Shaughnessy J et al, Blood, 2000; 96: 1505

Pathogenesis of Multiple Myeloma Two pathways involved in the early pathogenesis of MGUS and MM 50% Hyperdiploid 50% non-hyperdiploid Infrequent Ig. H Translocations Multiple trisomies of chromosomes 3, 5, 7, 9, 11, 15, 19 and 21 Hideshima et al, Blood, August 2004, 607 -618 11 q 13 (cyclin D 1) 4 p 16 FGFR 3+ MMSET 16 q 23 (cmaf) 6 p 21 (cyclin D 3) 20 q 11 (maf. B)

Prevelance of Ig. H Translocations Pathogenesis of Multiple Myeloma 100 90 80 70 60 50 40 30 20 10 0 MGUS Hideshima et al, Blood, August 2004, 607 -618 MM PPCL HMCLs

Prevalence of Ig. H Translocations 4 p 16 or 16 q 23 Ø Lower incidence with MGUS/SMM Ø de novo MM Ø Rapid progression of MGUS to MM Ø Extremely poor prognosis
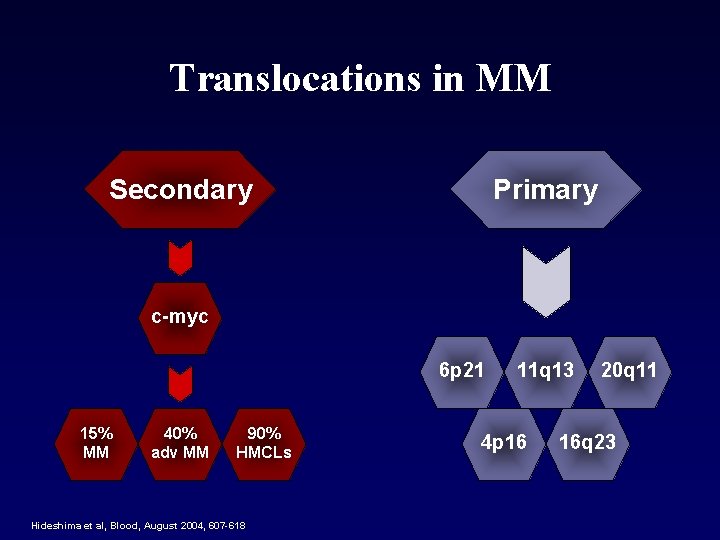
Translocations in MM Secondary Primary c-myc 6 p 21 15% MM 40% adv MM 90% HMCLs Hideshima et al, Blood, August 2004, 607 -618 11 q 13 4 p 16 20 q 11 16 q 23
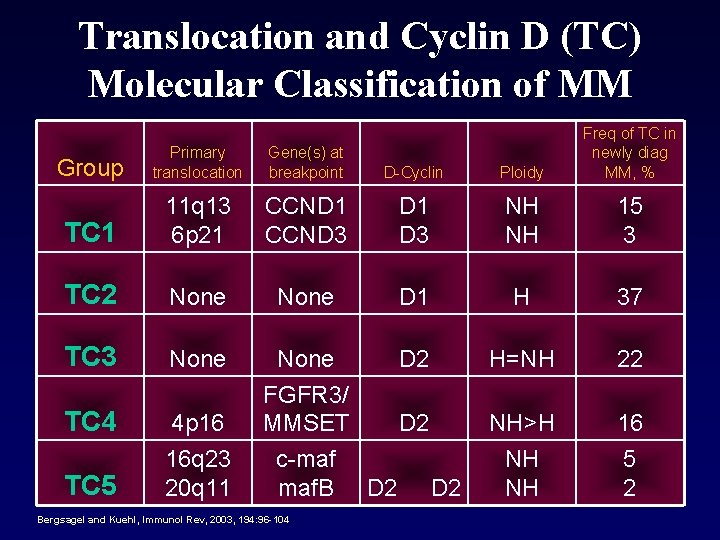
Translocation and Cyclin D (TC) Molecular Classification of MM Group Primary translocation Gene(s) at breakpoint D-Cyclin Ploidy Freq of TC in newly diag MM, % TC 1 11 q 13 6 p 21 CCND 3 D 1 D 3 NH NH 15 3 TC 2 None D 1 H 37 TC 3 None H=NH 22 TC 4 4 p 16 16 q 23 20 q 11 NH>H NH NH 16 5 2 TC 5 None D 2 FGFR 3/ MMSET D 2 c-maf maf. B D 2 Bergsagel and Kuehl, Immunol Rev, 2003, 194: 96 -104
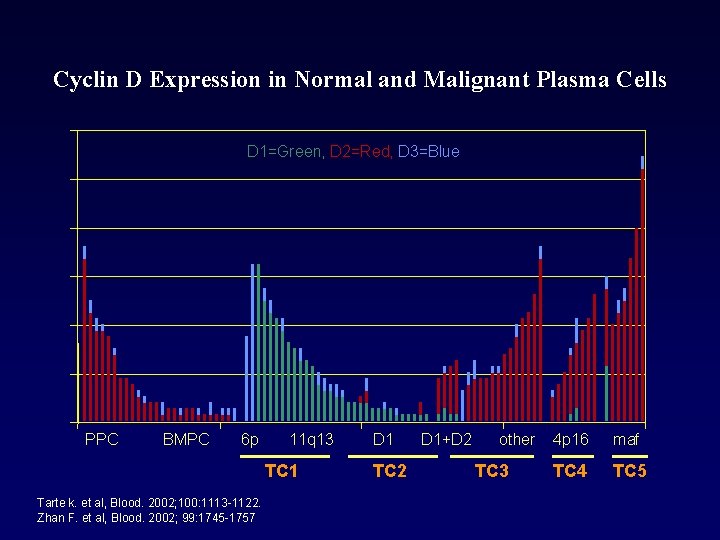
Cyclin D Expression in Normal and Malignant Plasma Cells D 1=Green, D 2=Red, D 3=Blue PPC BMPC 6 p 11 q 13 TC 1 Tarte k. et al, Blood. 2002; 100: 1113 -1122. Zhan F. et al, Blood. 2002; 99: 1745 -1757 D 1 TC 2 D 1+D 2 other TC 3 4 p 16 maf TC 4 TC 5

Dysregulation of cyclin D 1, D 2, D 3 “a unifying oncogenic event in MM” Ø MGUS and MM appear closer to normal PCs than to normal PBs Ø >30% of cells can be in S phase Ø Expression level of cyclin D 1, D 2, D 3 m. RNA in MM and MGUS is distinctly higher than normal PCs Ø Expression level of cyclin D 2 m. RNA is comparable with that expressed in normal proliferating PBs

Dysregulation of cyclin D 1, D 2, D 3 “a unifying oncogenic event in MM” Ø Cyclin D 1 is not expressed in normal hemopoitic cells Ø Cyclin D 1 expressed in 40% of MM lacking a t(11; 14) translocation Ø Ig translocations that dysregulate cyclin D 1 or D 3 occur in about 20% of MM tumors Ø Therefore, almost all MM tumors dysregulate at least one of the cyclin D genes
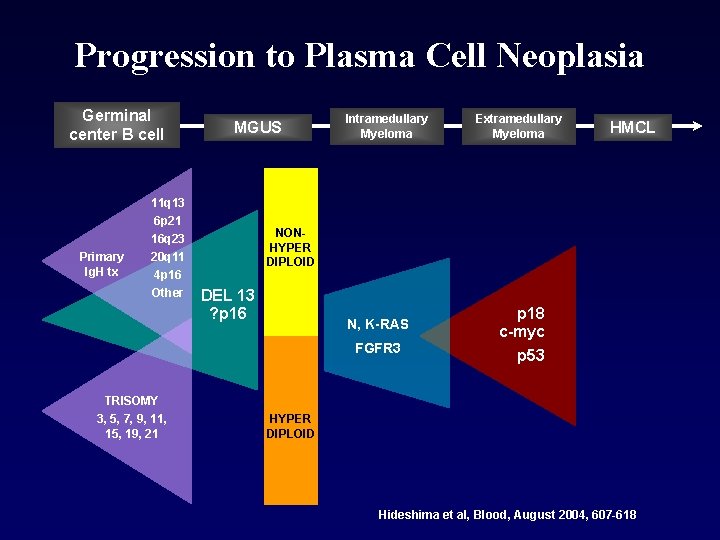
Progression to Plasma Cell Neoplasia Germinal center B cell Primary Ig. H tx MGUS 11 q 13 6 p 21 16 q 23 20 q 11 4 p 16 Other Intramedullary Myeloma HMCL NONHYPER DIPLOID DEL 13 ? p 16 N, K-RAS FGFR 3 TRISOMY 3, 5, 7, 9, 11, 15, 19, 21 Extramedullary Myeloma p 18 c-myc p 53 HYPER DIPLOID Hideshima et al, Blood, August 2004, 607 -618
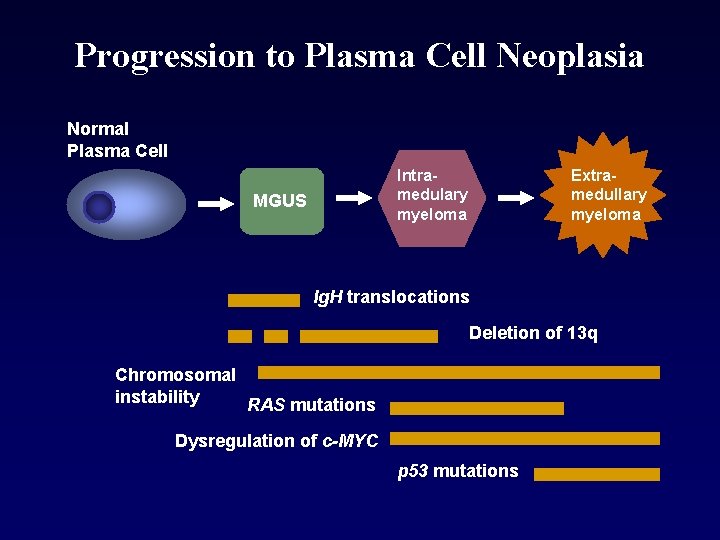
Progression to Plasma Cell Neoplasia Normal Plasma Cell Intramedulary myeloma MGUS Extramedullary myeloma Ig. H translocations Deletion of 13 q Chromosomal instability RAS mutations Dysregulation of c-MYC p 53 mutations

The TC Molecular Classification Predicts Prognosis and Response to Therapies Deletion of p 53 Increased PC Labeling Index Monosomy of chro 13/13 q Hypodiploidy Activating Mutations of K -Ras t(14; 16) TC 5 Lack of Cyclin D 1 Expression Bad prognosis Monosomy of chro 17 Tumor Cells with Abnormal Karyotype t(4; 14) TC 4
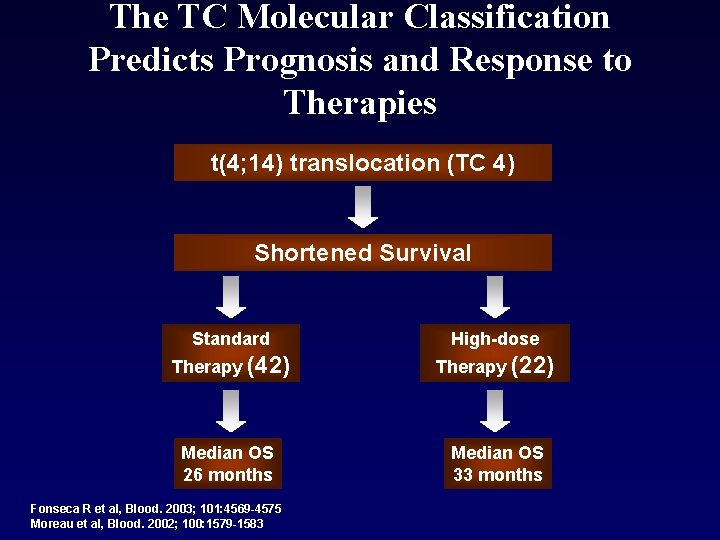
The TC Molecular Classification Predicts Prognosis and Response to Therapies t(4; 14) translocation (TC 4) Shortened Survival Standard High-dose Therapy (42) Therapy (22) Median OS 26 months Median OS 33 months Fonseca R et al, Blood. 2003; 101: 4569 -4575 Moreau et al, Blood. 2002; 100: 1579 -1583

The TC Molecular Classification Predicts Prognosis and Response to Therapies t(14; 16) translocation (TC 5) Shortened Survival (worse Prognosis) Standard Therapy (15) Median OS 16 months Fonseca R et al, Blood. 2003; 101: 4569 -4575
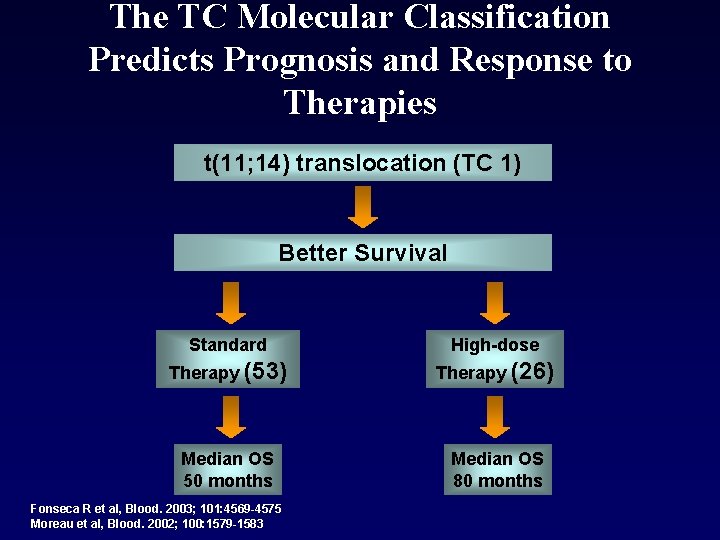
The TC Molecular Classification Predicts Prognosis and Response to Therapies t(11; 14) translocation (TC 1) Better Survival Standard High-dose Therapy (53) Therapy (26) Median OS 50 months Median OS 80 months Fonseca R et al, Blood. 2003; 101: 4569 -4575 Moreau et al, Blood. 2002; 100: 1579 -1583

The TC Molecular Classification Predicts Prognosis and Response to Therapies v The TC classification may be clinically useful way to classify patients into groups that have distinct subtypes of MM (and MGUS) tumors. v The TC classification identifies clinically important molecular subtypes of MM with different prognosis and with unique responses to different treatments.

The TC Molecular Classification Predicts Prognosis and Response to Therapies v High dose therapy and TC 1 v Microenvironment-directed therapy and TC 2 v FGFR 3 inhibitor and TC 4 v maf dominant-negative and TC 5
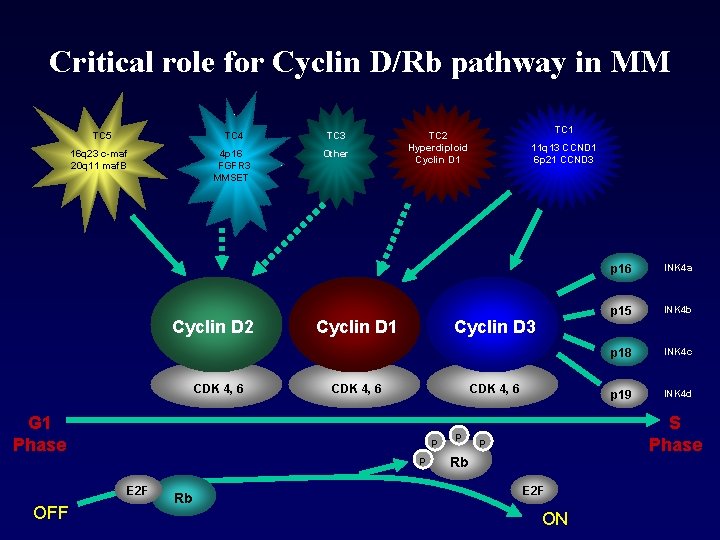
Critical role for Cyclin D/Rb pathway in MM TC 5 TC 4 TC 3 16 q 23 c-maf 20 q 11 maf. B 4 p 16 FGFR 3 MMSET Other Cyclin D 2 CDK 4, 6 G 1 Phase OFF Cyclin D 1 CDK 4, 6 p Rb 11 q 13 CCND 1 6 p 21 CCND 3 Cyclin D 3 p E 2 F TC 1 TC 2 Hyperdiploid Cyclin D 1 p p 16 INK 4 a p 15 INK 4 b p 18 INK 4 c p 19 INK 4 d S Phase p Rb E 2 F ON
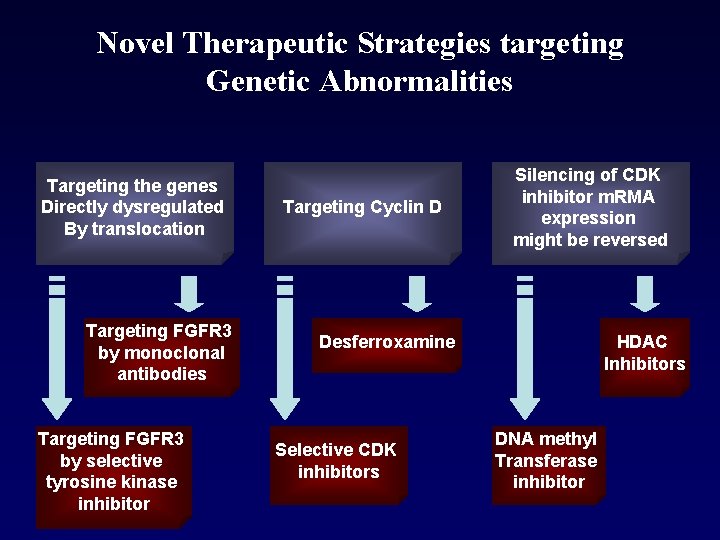
Novel Therapeutic Strategies targeting Genetic Abnormalities Targeting the genes Directly dysregulated By translocation Targeting FGFR 3 by monoclonal antibodies Targeting FGFR 3 by selective tyrosine kinase inhibitor Targeting Cyclin D Silencing of CDK inhibitor m. RMA expression might be reversed HDAC Inhibitors Desferroxamine Selective CDK inhibitors DNA methyl Transferase inhibitor
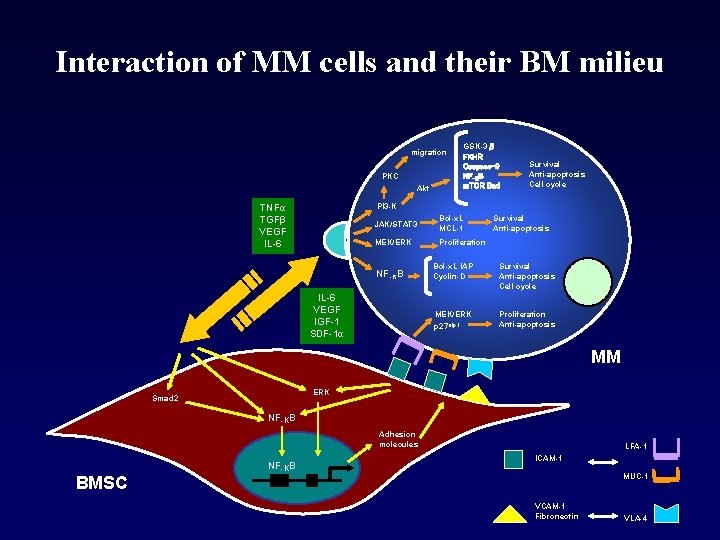
Interaction of MM cells and their BM milieu migration PKC Akt GSK-3β FKHR Caspase-9 NF-KB m. TOR Bad Survival Anti-apoptosis Cell cycle PI 3 -K TNFα TGFβ VEGF IL-6 JAK/STAT 3 Bcl-x. L MCL-1 MEK/ERK Proliferation NF-KB IL-6 VEGF IGF-1 SDF-1α Survival Anti-apoptosis Bcl-x. L IAP Cyclin-D Survival Anti-apoptosis Cell cycle MEK/ERK p 27 Kip 1 Proliferation Anti-apoptosis MM ERK Smad 2 NF-KB Adhesion molecules NF-KB LFA-1 ICAM-1 MUC-1 BMSC VCAM-1 Fibronectin VLA-4

Myeloma Cells and BM Microenvironment Bruno et al, The Lancet Oncology, July 2004, 430 -442
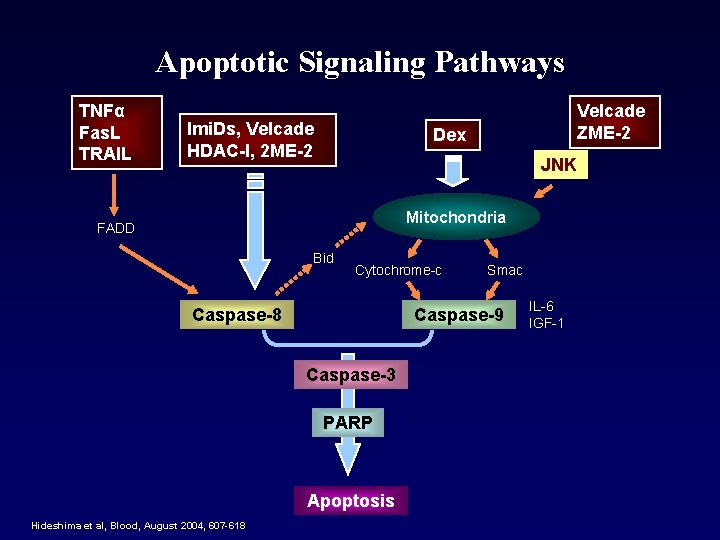
Apoptotic Signaling Pathways TNFα Fas. L TRAIL Imi. Ds, Velcade HDAC-I, 2 ME-2 Velcade ZME-2 Dex JNK Mitochondria FADD Bid Cytochrome-c Caspase-8 Caspase-9 Caspase-3 PARP Apoptosis Hideshima et al, Blood, August 2004, 607 -618 Smac IL-6 IGF-1
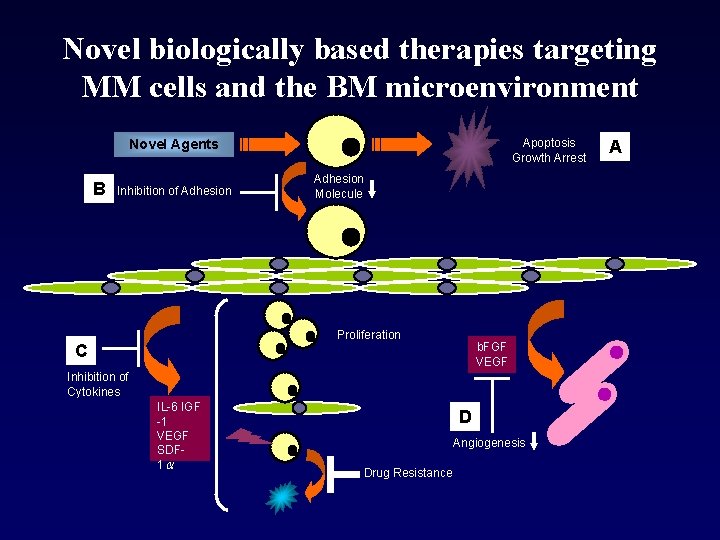
Novel biologically based therapies targeting MM cells and the BM microenvironment Apoptosis Growth Arrest Novel Agents B Inhibition of Adhesion Molecule Proliferation C b. FGF VEGF Inhibition of Cytokines IL-6 IGF -1 VEGF SDF 1α D Angiogenesis Drug Resistance A

Novel Agents for Myeloma Ø Targeting both MM cells and interaction of MM cells with the BM microenvironment Ø Targeting circuits mediating MM cell growth and survival Ø Targeting the BM microenvironment Ø Targeting cell surface receptors

Novel Agents for Myeloma Targeting both MM cells and their interaction with BM microenvironment Thalidomide and its analogs (Revlimid) Proteasome inhibitor (Bortezomib) Arsenic trioxide 2 -Methoxyestradiol (2 -ME 2) Lysophosphatidic acid acyltransferase-β inhibitor Triterpinoid 2 -cyano-3, 12 -dioxoolean-1, 9 -dien 28 - oic acid (CDDO) N-N-Diethl-8, 8 -dipropyl-2 -azaspiro [4. 5] decane-2 -propanamine (Atiprimod) Targeting circuits mediating MM cell growth and survival VEGF receptor tyrosine kinase inhibitor (PTK 787/ZK 222584, GW 654652) Farnesyltransferase inhibitor Histone deacetylase inhibitor (SAHA, LAQ 824) Heat shock protein-90 inhibitor (Geldanamycin, 17 -AAG) Telomerase inhibitor (Telomestatin) bcl-2 antisense oligonucleotide (Genasense) Inosine monophophate dehydrogenase (VX-944) Rapamycin Targeting the bone marrow microenvironment IĸB kinase (IKK) inhibitor (PS-1145) p 38 MAPK inhibitor (VX-745, SCIO-469) TFG-β inhibitor (SD-208) Targeting cell surface receptors TNF related apoptosis-inducing ligand (TRAIL) / Apo 2 ligand IGF-1 receptor inhibitor ( ADW) HMG-Co. A reductase inhibitor (statins) Anti-CD 20 antibody (Rituximab)
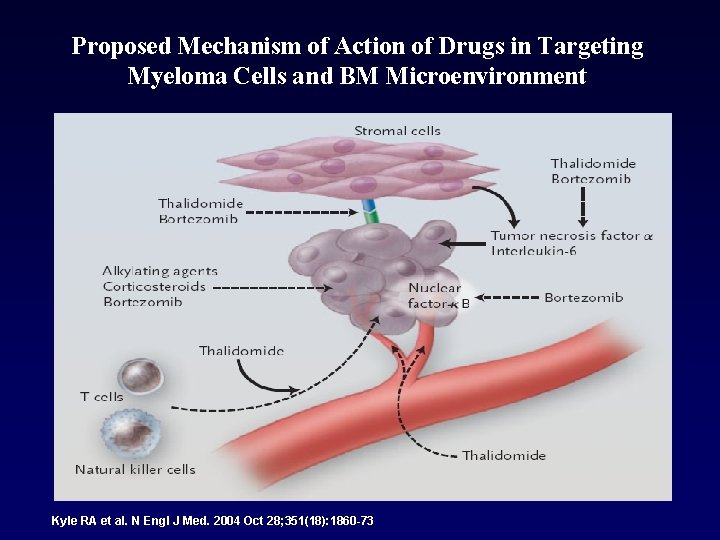
Proposed Mechanism of Action of Drugs in Targeting Myeloma Cells and BM Microenvironment Kyle RA et al. N Engl J Med. 2004 Oct 28; 351(18): 1860 -73
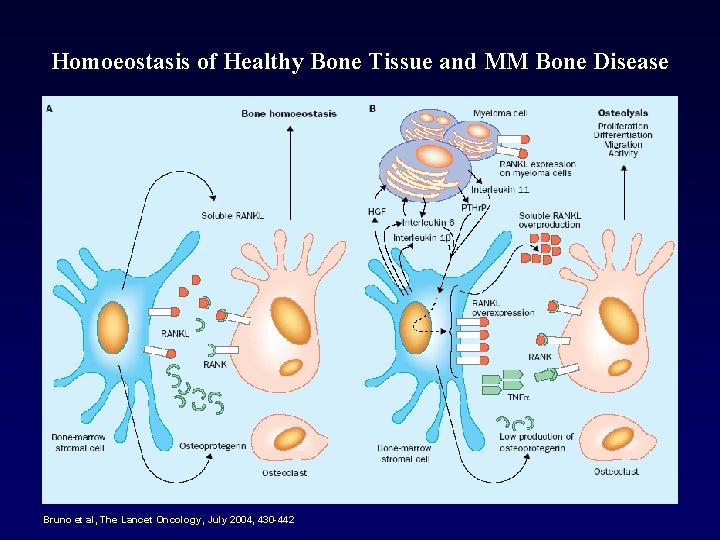
Homoeostasis of Healthy Bone Tissue and MM Bone Disease Bruno et al, The Lancet Oncology, July 2004, 430 -442
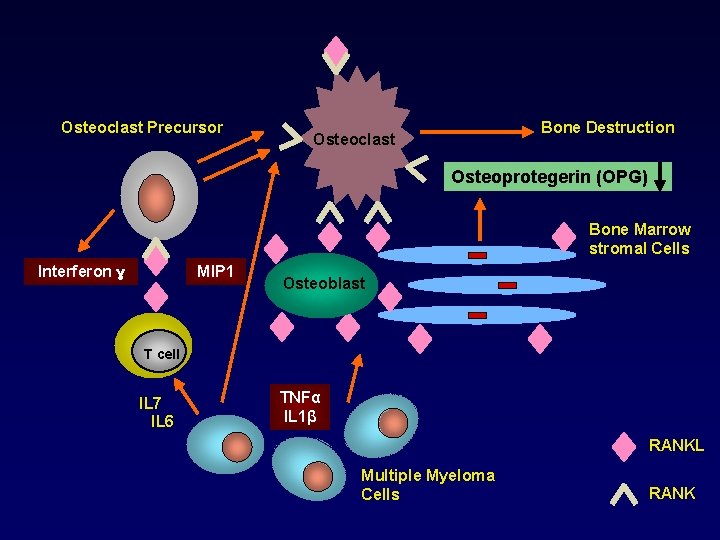
Osteoclast Precursor Bone Destruction Osteoclast Osteoprotegerin (OPG) Bone Marrow stromal Cells Interferon ɣ MIP 1 Osteoblast T cell IL 7 IL 6 TNFα IL 1β RANKL Multiple Myeloma Cells RANK

Effects of Thalidomide on the Myeloma Microenvironment Bruno et al, The Lancet Oncology, July 2004, 430 -442
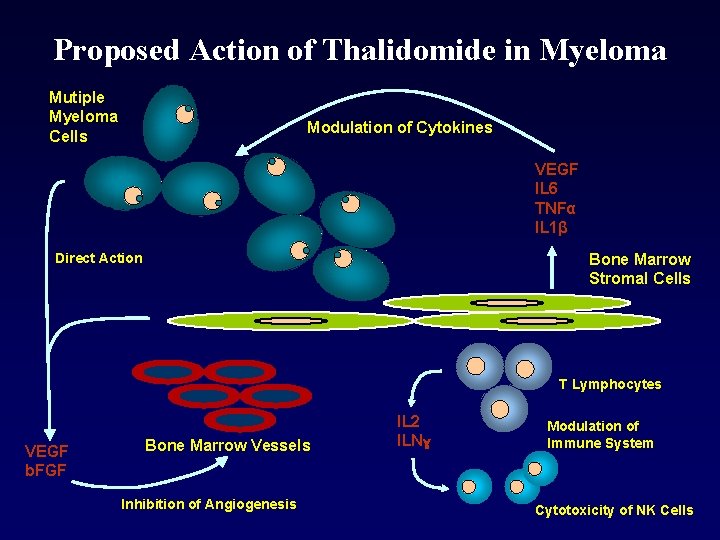
Proposed Action of Thalidomide in Myeloma Mutiple Myeloma Cells Modulation of Cytokines VEGF IL 6 TNFα IL 1β Direct Action Bone Marrow Stromal Cells T Lymphocytes VEGF b. FGF Bone Marrow Vessels Inhibition of Angiogenesis IL 2 ILNɣ Modulation of Immune System Cytotoxicity of NK Cells

Mechanism of Action of Bortezomib Phosphorylation of NFKB inhibitory partner protein IKB leads to degradation of IKB by the proteosome and release of NFKB migrates into the nucleus to induce arrest of apoptosis and expression of adhesion molecule Affinity of Bortezomib for the proteosome inhibits protein degradation, and prevents nuclear translocation of NFKB Bruno et al, The Lancet Oncology, July 2004, 430 -442
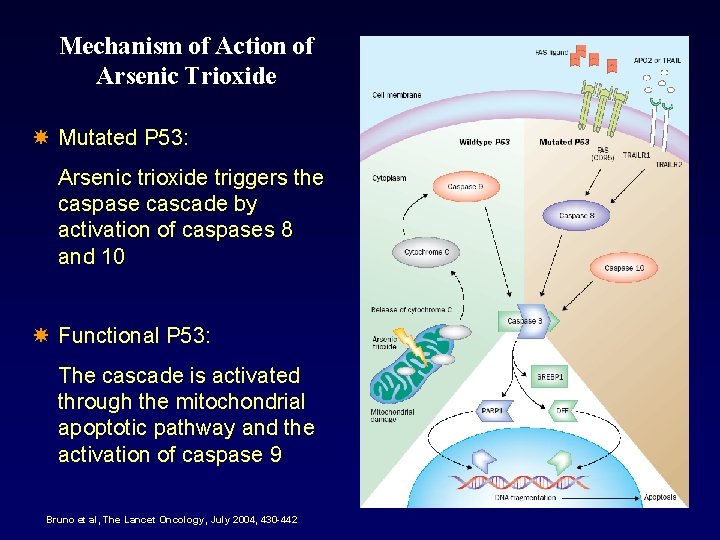
Mechanism of Action of Arsenic Trioxide Mutated P 53: Arsenic trioxide triggers the caspase cascade by activation of caspases 8 and 10 Functional P 53: The cascade is activated through the mitochondrial apoptotic pathway and the activation of caspase 9 Bruno et al, The Lancet Oncology, July 2004, 430 -442


- Slides: 54