Achieving Semantic Interoperability of Cancer Registries NAACCR 2007

Achieving Semantic Interoperability of Cancer Registries NAACCR 2007 Annual Meeting June 7, 2007 Peter Winkelstein, MD Chief, Division of General Pediatrics CMIO, UPA and UBA University at Buffalo, New York pwink@buffalo. edu

Informatics Goals for Cancer Registries • Improve efficiency and accuracy of data entry • Automated data extraction from EHRs and other systems • Automated data checking • Improve data extraction • Facilitate reporting (querying) • Facilitate secondary use of registry data • Translational research • Data mining Peter Winkelstein, MD Biomedical Informatics Consulting

Computable Health Information • Registry data must be: • Stored in such a way as to be amenable to computational processes (e. g. database elements) • Have carefully defined meaning (e. g. ontologies) • Required for interoperability • Required for robust querying Peter Winkelstein, MD Biomedical Informatics Consulting

What is Ontology (philosophically)? A representation of some pre-existing domain of reality which: • reflects the properties of the objects within its domain in such a way that there is a systematic correlation between reality and the representation itself • is intelligible to a domain expert • is formalized in a way that allows it to support automatic information processing (Source: Werner Ceusters, 12/23/2005) Peter Winkelstein, MD Biomedical Informatics Consulting

What is an ontology (practically)? A formal model of a domain which consists of: • Classes (aka concepts) • Instances (particular members of classes) • Instances are not generally part of published ontologies • Relationships between classes (aka roles) In contrast to controlled vocabularies, which are word lists with no (or limited) relationships Peter Winkelstein, MD Biomedical Informatics Consulting
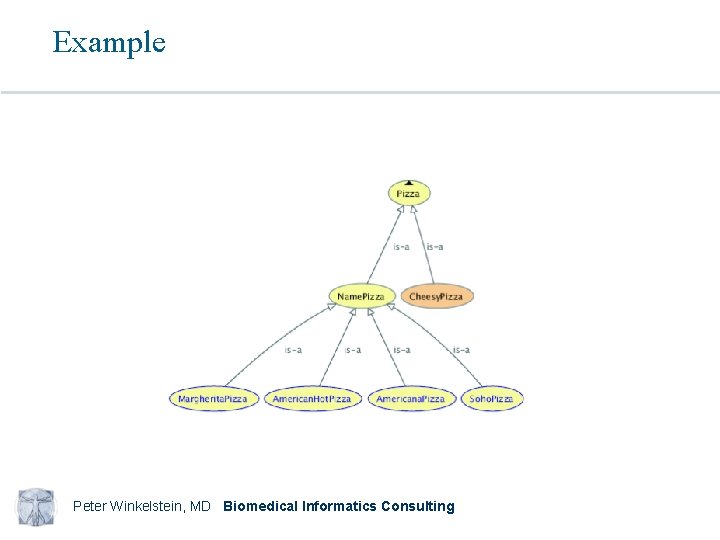
Example Peter Winkelstein, MD Biomedical Informatics Consulting

Syntactic vs. Semantic Interoperability • The ability to exchange information among systems and software, especially EHRs, is called interoperability • The ability to exchange data is called syntactic interoperability • Interfaces/Messaging • The ability to exchange data and meaning is called semantic interoperability • Interfaces/Messaging plus Ontologies Peter Winkelstein, MD Biomedical Informatics Consulting

Syntactic Interoperability Peter Winkelstein, MD Biomedical Informatics Consulting
Semantic Interoperability Peter Winkelstein, MD Biomedical Informatics Consulting

Syntactic Interoperability Alone Is Not Enough • Free text data has little (if any) machine-interpretable meaning • Data elements are not guaranteed interoperable • Multiple encodings • Inflexible encoding • Incompatible lists/Granularity • Data abstraction • Impression vs. criteria • Qualitative qualifiers • Lack of standards Peter Winkelstein, MD Biomedical Informatics Consulting

Semantic Interoperability • Semantic interoperability allows the exchange of both structure and meaning among systems • Requires data in structured form (database elements) plus encoding data with machine-interpretable meaning • Ontologies are the tools for this encoding Peter Winkelstein, MD Biomedical Informatics Consulting

State-of-the-art in Ontology • There are few, if any, well-formed medical ontologies (or even controlled vocabularies) • Notably for cancer informatics, the current state of the NCI Thesaurus is suboptimal Peter Winkelstein, MD Biomedical Informatics Consulting

Some Existing Vocabularies and Ontologies Controlled Vocabularies “Ontologies” • DICOM • HL 7 RIM • LOINC • UMLS • HCPCS • NCIT • ICD-9 • FMA • CPT • GO • ICD-O • SNOMED-CT • Med. DRA Peter Winkelstein, MD Biomedical Informatics Consulting

Examples NAACCR Standards for Cancer Registries Volume II Data Standards and Data Dictionary Eleventh Edition Record Layout Version 11. 1 Peter Winkelstein, MD Biomedical Informatics Consulting
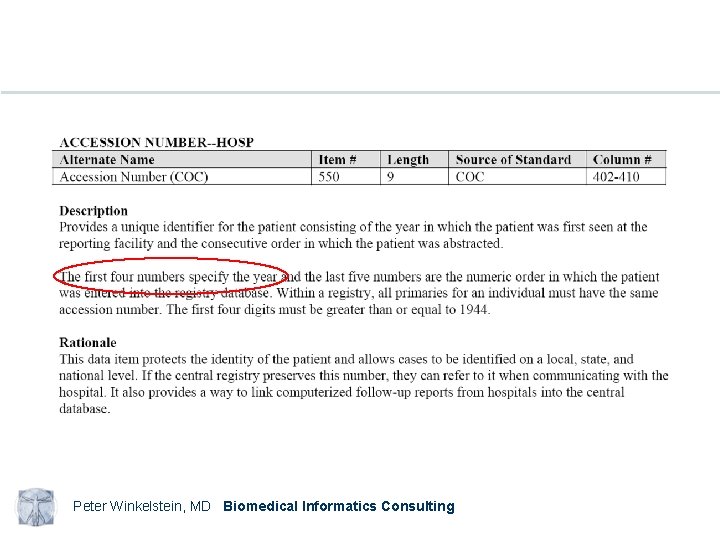
Peter Winkelstein, MD Biomedical Informatics Consulting

Peter Winkelstein, MD Biomedical Informatics Consulting
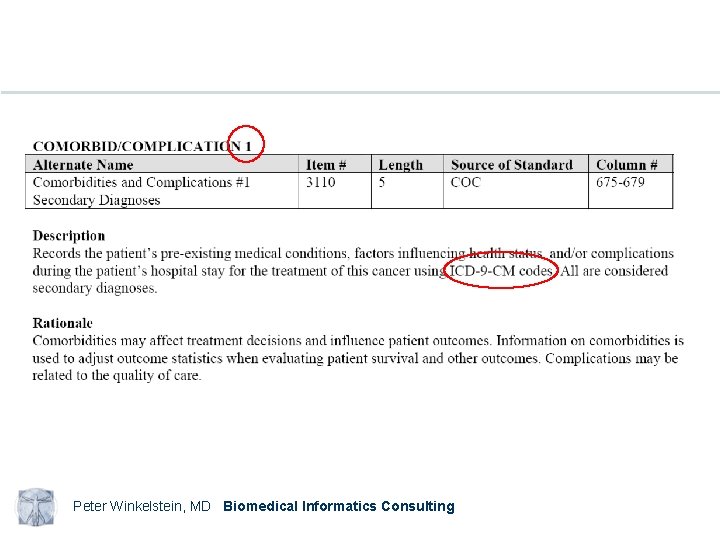
Peter Winkelstein, MD Biomedical Informatics Consulting
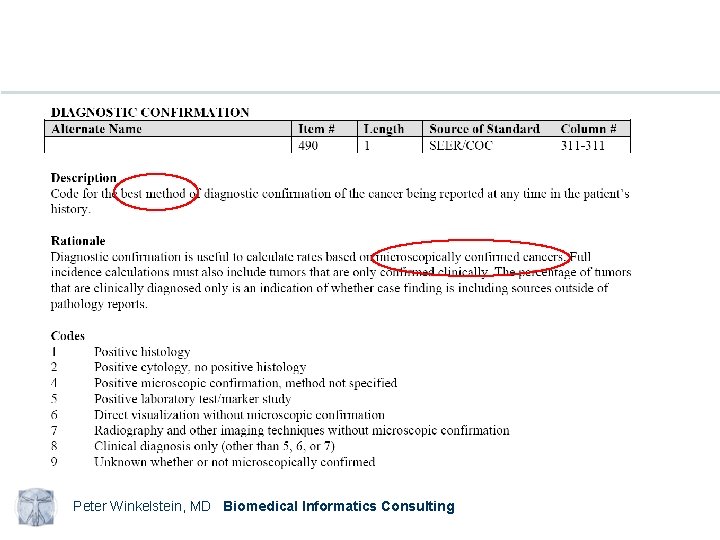
Peter Winkelstein, MD Biomedical Informatics Consulting

Peter Winkelstein, MD Biomedical Informatics Consulting

Peter Winkelstein, MD Biomedical Informatics Consulting

Peter Winkelstein, MD Biomedical Informatics Consulting
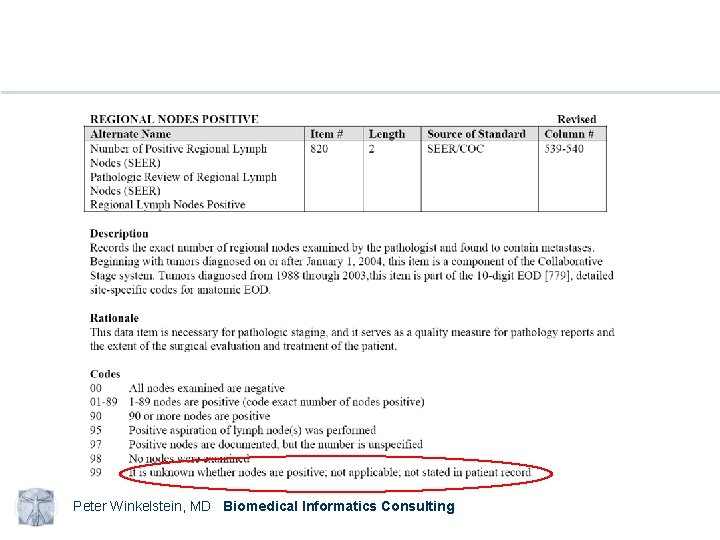
Peter Winkelstein, MD Biomedical Informatics Consulting
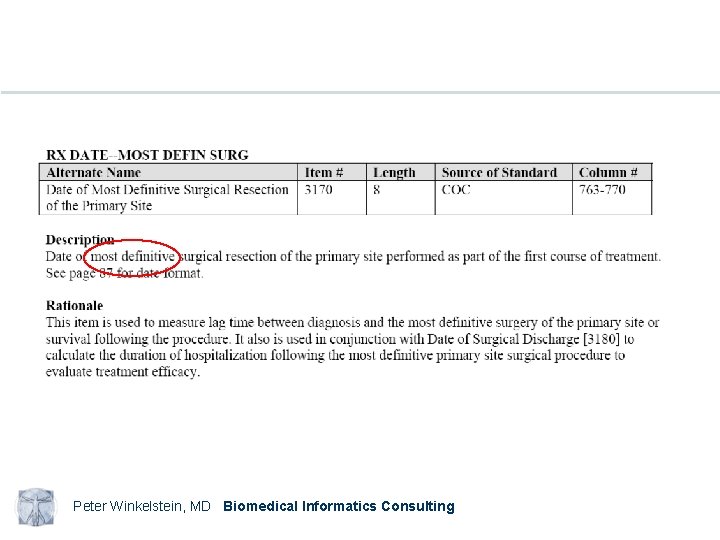
Peter Winkelstein, MD Biomedical Informatics Consulting

Peter Winkelstein, MD Biomedical Informatics Consulting
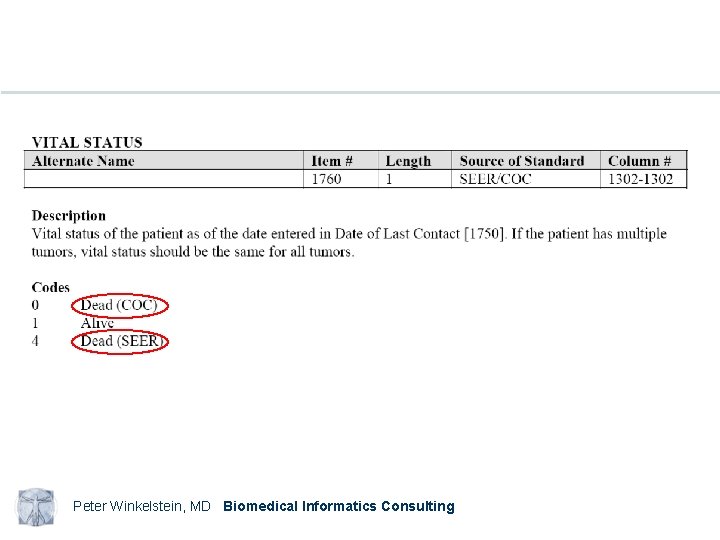
Peter Winkelstein, MD Biomedical Informatics Consulting

Keys to Achieving Semantic Interoperability • Structured data collection (no free text) • Highest granularity possible • Careful definition of meaning for each data element • Consistent use of each data element • Assignment of standard code to each data element if possible • Vocabulary creation and maintenance for meanings not available among current standards • Publication and versioning of all definitions Peter Winkelstein, MD Biomedical Informatics Consulting

Recommendations • Cancer registries should move towards semantic interoperability • This would facilitate improvements in both data collection and extraction • A strong link to EHR data is particularly desirable • Registry organizations should actively work with vendors towards semantic interoperability with oncology EHRs Peter Winkelstein, MD Biomedical Informatics Consulting
- Slides: 27