Abstractions for DataIntensive Computing on Condor Christopher Moretti

Abstractions for Data-Intensive Computing on Condor Christopher Moretti University of Notre Dame 4/21/2009 Christopher Moretti – University of Notre Dame

I want to complete big workloads 3. 6 B Hamming distance computations, . 02 seconds each on a 2 GHz dual-core desktop computer, each creating 1 real number output: 2. 3 CPUYrs and 29 GB of output. 85 M 1000 x 1000 dynamic programming tables, . 04 seconds each on a 3 GHz quad-core Xeon server: 39 CPUDays requiring a total of 77 GB of input data. 500 x 500 recurrence matrix, recurrence functions 7 seconds each on the desktop: 22 CPUDays, not completely independently parallelizable. … can these be run this week? How about this afternoon? Can we even turn these into “lunchtime” or “coffee refill” problems? 4/21/2009 Christopher Moretti – University of Notre Dame

How? 4 On my workstation. < Write my program, make sure to make it partitionable, because it takes a really long time and might crash, debug it. Now run it for 39 days – 2. 3 years. 4 On my department’s 128 -node research cluster < Learn MPI, determine how I want to move many GBs of data around, re-write my program and re-debug, wait until the cluster can give me 8 -128 homogeneous nodes at once, or go buy my own. Now run it. 4 Blue. Gene < Get $$$ or access, learn custom MPI-like computation and communication working language, determine how I want to handle communication and data movement, re-write my program, wait for configuration or access, re-debug my program, re-run. 4/21/2009 Christopher Moretti – University of Notre Dame

So? 4 Serially 4 Cluster 4 Supercomputer 4 So I can either take my program as -is and it’ll take forever, or I can do a new custom implementation to a certain particular architecture and re-write and re-debug it every time we upgrade (assuming I’m lucky enough to have a Blue. Gene in the first place)? 4 Well what about Condor? 4/21/2009 Christopher Moretti – University of Notre Dame

Yes, what about Condor? Which resources ? How do I fit my workload into jobs? What about job input data? How can I measure job stats? 4/21/2009 What is Condor? How Many? What happens when things fail? How long will it take? What do I do with the results? Christopher Moretti – University of Notre Dame
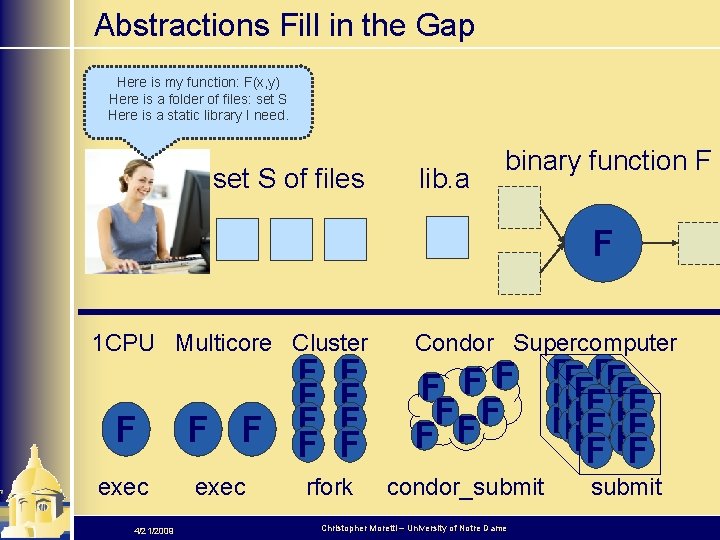
Abstractions Fill in the Gap Here is my function: F(x, y) Here is a folder of files: set S Here is a static library I need. set S of files lib. a binary function F F 1 CPU Multicore Cluster F exec 4/21/2009 F F F exec F F rfork Condor Supercomputer F F F F FF F FFFF FF condor_submit Christopher Moretti – University of Notre Dame submit

What is an abstraction? 4 Abstraction: a declarative specification of the computation and data of a workload. 4 A restricted pattern, not meant to be a general purpose programming language. 4 Uses data structures instead of files. 4 Regular structure makes it tractable to model and predict performance. 4 Allows a user to repeat the same pattern of work many times, making slight changes to the data and algorithms 4/21/2009 Christopher Moretti – University of Notre Dame

Abstractions Example Approaches 4 Data management: distribute data only to nodes where it is necessary for computation. Distribute broadcasted data via efficient algorithms. Access data in a memoryefficient order. Use data structures instead of flat files. 4 Task management: assign appropriate amounts of data per discrete task. Adapt to the environment by choosing nodes showing good performance. Submit/manage tasks that do not overwhelm the batch system. 4/21/2009 Christopher Moretti – University of Notre Dame
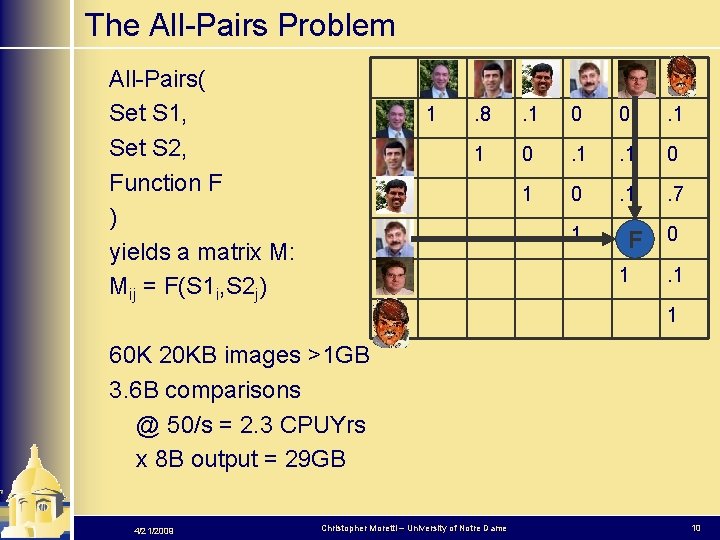
The All-Pairs Problem All-Pairs( Set S 1, Set S 2, Function F ) yields a matrix M: Mij = F(S 1 i, S 2 j) 1 . 8 . 1 0 0 . 1 1 0 1 0 . 1 . 7 1 F 1 0. 1 1 60 K 20 KB images >1 GB 3. 6 B comparisons @ 50/s = 2. 3 CPUYrs x 8 B output = 29 GB 4/21/2009 Christopher Moretti – University of Notre Dame 10

All Pairs Abstraction set S of files binary function F F invocation M = All. Pairs(F, S) 4/21/2009 Christopher Moretti – University of Notre Dame

Biometrics All-Pairs at Notre Dame 4/21/2009 Christopher Moretti – University of Notre Dame
![Wavefront Recurrence ( R[x, 0], R[0, y], F(x, y, d) ) R[0, 4] x Wavefront Recurrence ( R[x, 0], R[0, y], F(x, y, d) ) R[0, 4] x](http://slidetodoc.com/presentation_image_h2/d791726e944c350e1641b73604aef491/image-13.jpg)
Wavefront Recurrence ( R[x, 0], R[0, y], F(x, y, d) ) R[0, 4] x F d R[0, 3] x d R[0, 2] R[0, 0] 4/21/2009 R[3, 2] R[4, 3] F R[4, 2] d y F d F x R[4, 4] x y d R[0, 1] R[3, 4] y F x R[2, 4] y R[1, 0] y F d x d y F d x y R[2, 0] x y F d y R[3, 0] x F d y R[4, 0] Christopher Moretti – University of Notre Dame 14
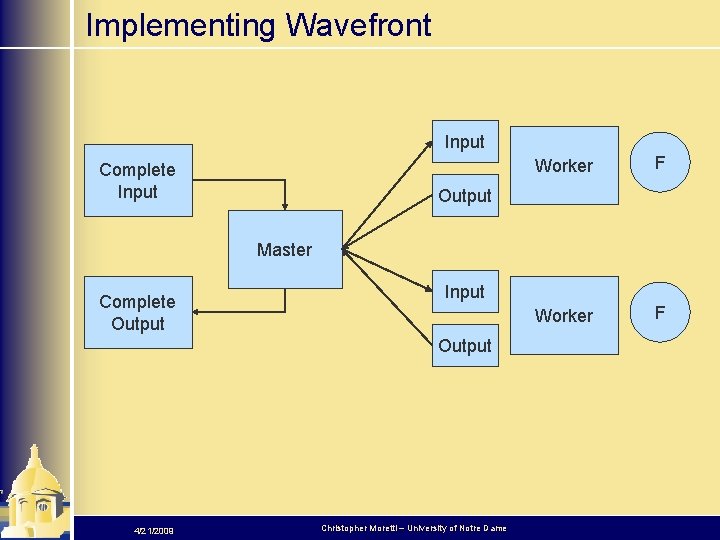
Implementing Wavefront Input Complete Input Worker F Output Master Complete Output Input Output 4/21/2009 Christopher Moretti – University of Notre Dame
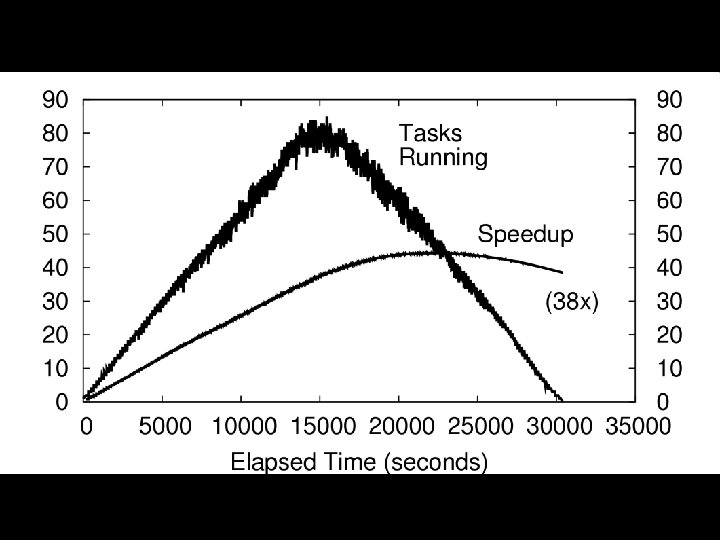

Genome Assembly 4 Bioinformatics sequencers can only extract DNA from samples 50 -1000 basepairs (A, C, G, T) at a time. 4 Biologists need the DNA together in genome profiles of millions of contiguous basepairs. 4 Genome assembly is the process of putting the pieces of the puzzle back together again in the right configuration. A principal step is “overlapping”. 4 One way would be a huge All-Pairs problem, but this isn’t necessary. Algorithms exist to extract a sparse matrix of possible candidate pairs (two sequences that might overlap in the right answer). So we must only compute the overlaps for the candidates. 4/21/2009 Christopher Moretti – University of Notre Dame
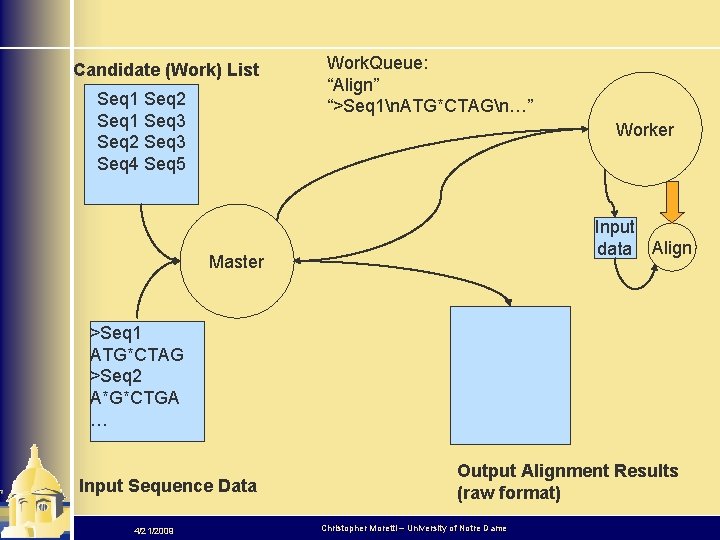
Candidate (Work) List Seq 1 Seq 2 Seq 1 Seq 3 Seq 2 Seq 3 Seq 4 Seq 5 Work. Queue: “Align” “>Seq 1n. ATG*CTAGn…” Worker Input data Align Master >Seq 1 ATG*CTAG >Seq 2 A*G*CTGA … Input Sequence Data 4/21/2009 Output Alignment Results (raw format) Christopher Moretti – University of Notre Dame
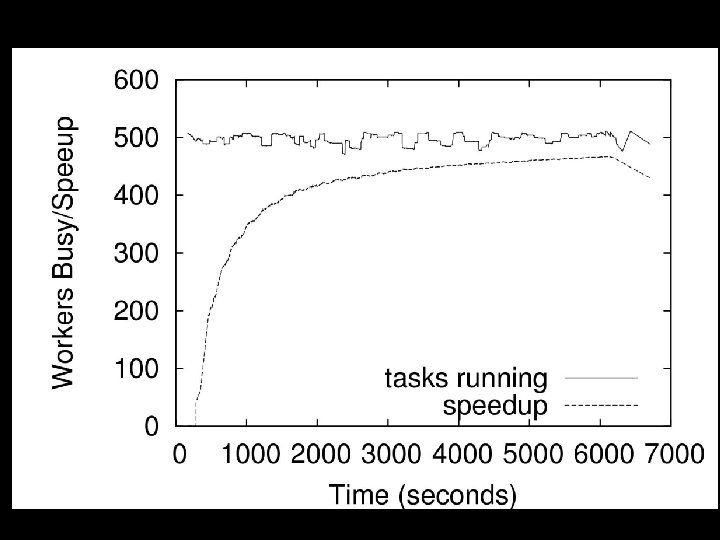

Purpose of a Suite of Abstractions 4 Engineering: Big systems to solve cool problems. 4 Science: < What are common elements of abstractions? < What is an intuitive interface for users to adapt their existing serial solutions to larger problems? data Set S computation invocation F M = All. Pairs(F, S) F R = Wavefront(F, R) F O = Overlap(F, C, S) Initial State Seqs 4/21/2009 Cands Christopher Moretti – University of Notre Dame

Challenges Remaining with Abstractions 4 Exploiting a regular pattern doesn’t mean that nothing can go wrong … < It’s not just domain scientists who can make mistakes that lead to disastrous consequences. < Sometimes good solutions to problems beget new and interesting problems. 4 A motivating example to finish … 4/21/2009 Christopher Moretti – University of Notre Dame
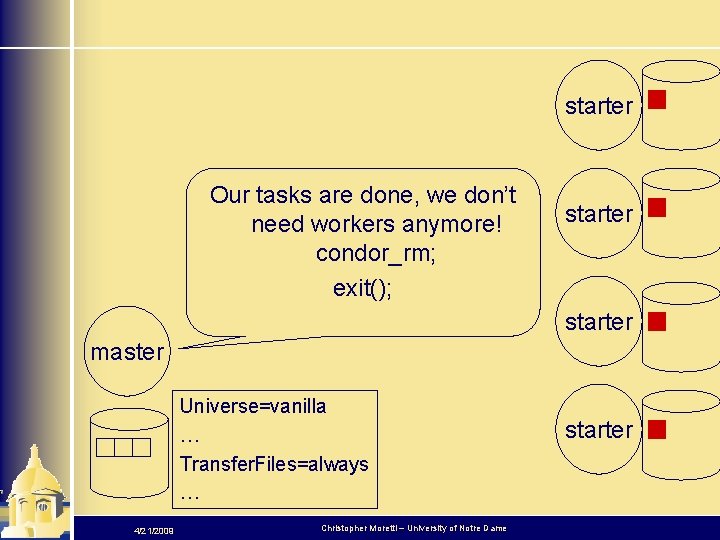
starter Our tasks are done, we don’t need workers anymore! condor_rm; exit(); starter master Universe=vanilla … Transfer. Files=always … 4/21/2009 Christopher Moretti – University of Notre Dame starter
starter Okay, I’ll send back the data the workers generated. shadow schedd . . . Okay, I’ll send back the data the workers generated. 4/21/2009 Christopher Moretti – University of Notre Dame starter
starter Here’s the data the worker generated! starter shadow starter schedd starter 4/21/2009 Christopher Moretti – University of Notre Dame
starter shadow schedd Here’s the data the worker generated! starter 4/21/2009 Christopher Moretti – University of Notre Dame
starter shadow starter schedd Here’s the data the worker generated! 4/21/2009 Christopher Moretti – University of Notre Dame starter
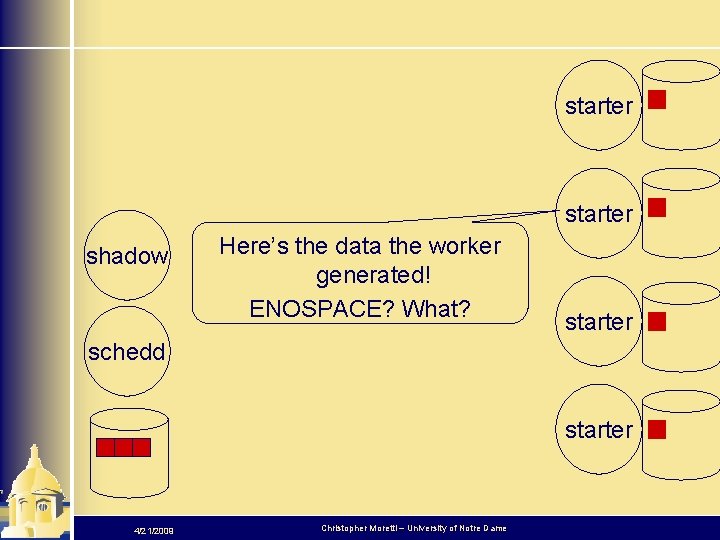
starter shadow Here’s the data the worker generated! ENOSPACE? What? starter schedd starter 4/21/2009 Christopher Moretti – University of Notre Dame

starter shadow schedd Ack! We didn’t want those files back, they were temporary. “Transfer. Files=Always” was a mistake! Now we’re out of space! starter 4/21/2009 Christopher Moretti – University of Notre Dame

Hrm, I can’t transfer back their files. I guess I’ll hang out and won’t remove the jobs until I can. starter shadow schedd 4/21/2009 Christopher Moretti – University of Notre Dame
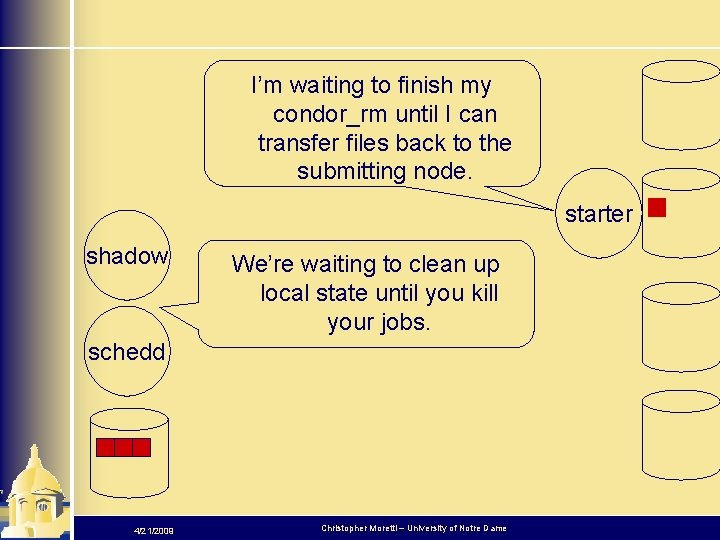
I’m waiting to finish my condor_rm until I can transfer files back to the submitting node. starter shadow We’re waiting to clean up local state until you kill your jobs. schedd 4/21/2009 Christopher Moretti – University of Notre Dame

I’m waiting to finish my condor_rm until I can transfer files back to the submitting node. starter shadow We’re waiting to clean up local state until you kill your jobs. ARGH! schedd 4/21/2009 Christopher Moretti – University of Notre Dame

Moral of the Story 4 Abstractions are a way to give domain scientists tools that don’t require drastically changing their alreadycomplete solutions, but still allow for efficient HPC/HTC. 4 Exploiting a regular pattern doesn’t mean that nothing can go wrong. 4 Interesting challenges remain, among these: < Predicting performance on an unpredictable system < Monitoring and adapting to fit a changing system < Dealing with entangling relationships = e. g. Remote state requires local state < Tying together the lessons learned from these abstractions = What do they have in common? = Why is one solution right for one problem and wrong for another? 4/21/2009 Christopher Moretti – University of Notre Dame

For More Information 4 Christopher Moretti < cmoretti@cse. nd. edu 4 Douglas Thain < dthain@cse. nd. edu 4 Cooperative Computing Lab < http: //cse. nd. edu/~ccl 4/21/2009 Christopher Moretti – University of Notre Dame
- Slides: 32