Abinitio Density Functional Theory from quantum dots to

Ab-initio Density Functional Theory: from quantum dots to solar cell Saifful Kamaluddin Muzakir Staf ID: 01009 019 276 3844 skmuzakir@yahoo. com www. facebook. com/Saifful Kamaluddin
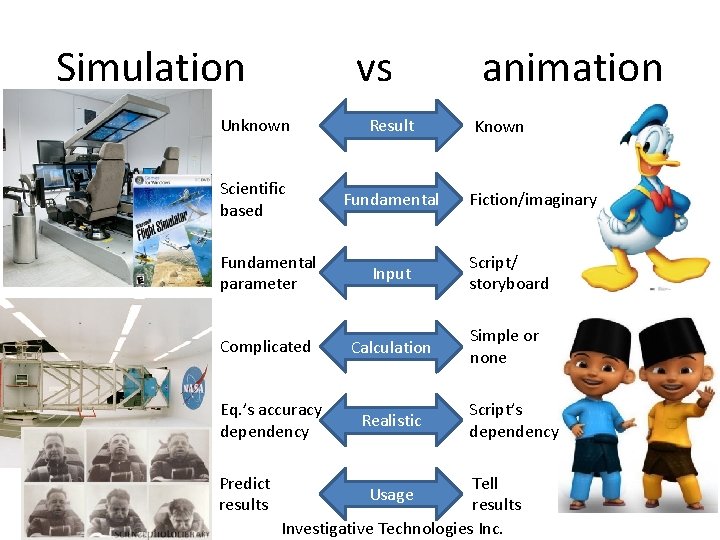
Simulation vs Unknown Result Scientific based Fundamental animation Known Fiction/imaginary Fundamental parameter Input Script/ storyboard Complicated Calculation Simple or none Eq. ’s accuracy dependency Realistic Predict results Script’s dependency Tell results Investigative Technologies Inc. Usage
The advantages of simulation wet lab Excited state Ground state Absorption spectroscopy Cluster size Electron density IR spectroscopy
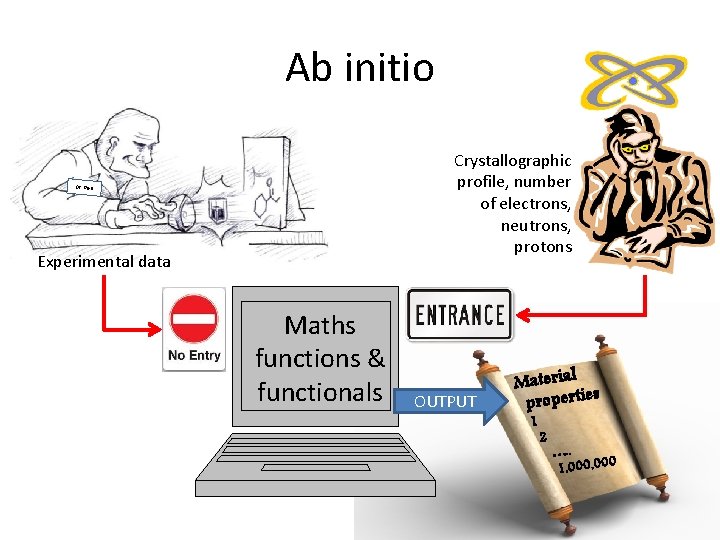
Ab initio Crystallographic profile, number of electrons, neutrons, protons Dr. Izan Experimental data Maths functions & functionals OUTPUT Material s propertie 1 2 …. . 0 1, 000, 00
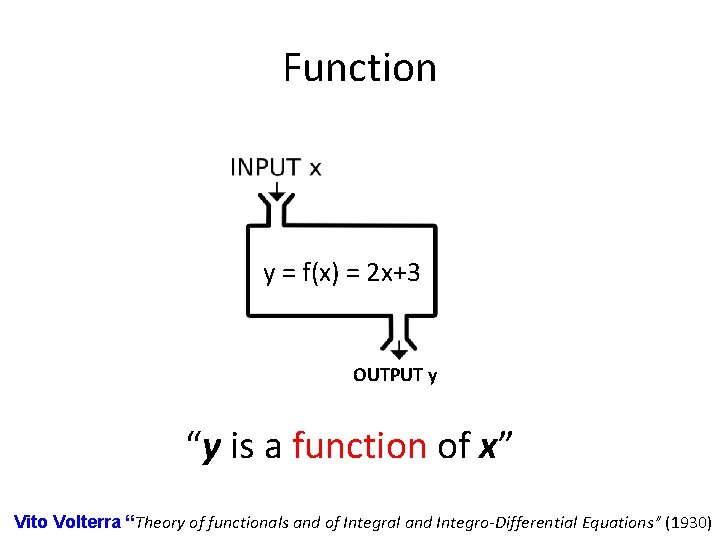
Function y = f(x) = 2 x+3 OUTPUT y “y is a function of x” Vito Volterra “Theory of functionals and of Integral and Integro-Differential Equations” (1930)
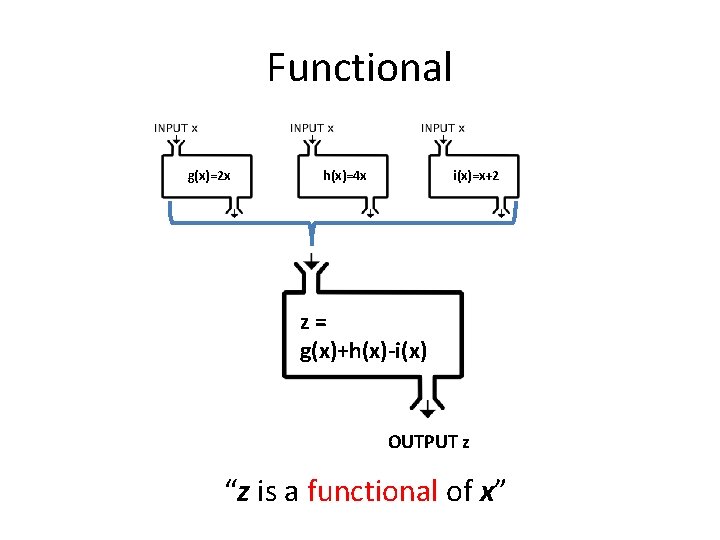
Functional g(x)=2 x h(x)=4 x i(x)=x+2 z= g(x)+h(x)-i(x) OUTPUT z “z is a functional of x”
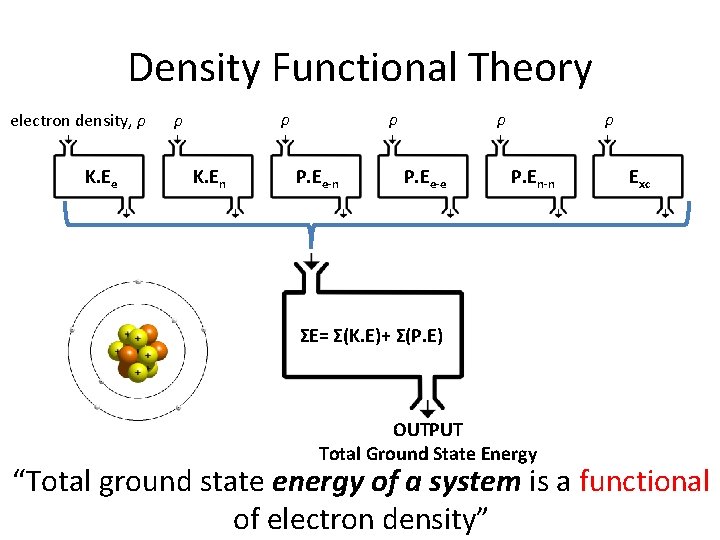
Density Functional Theory electron density, ρ K. Ee ρ ρ K. En ρ P. Ee-e ρ P. En-n Exc ΣE= Σ(K. E)+ Σ(P. E) OUTPUT Total Ground State Energy “Total ground state energy of a system is a functional of electron density”
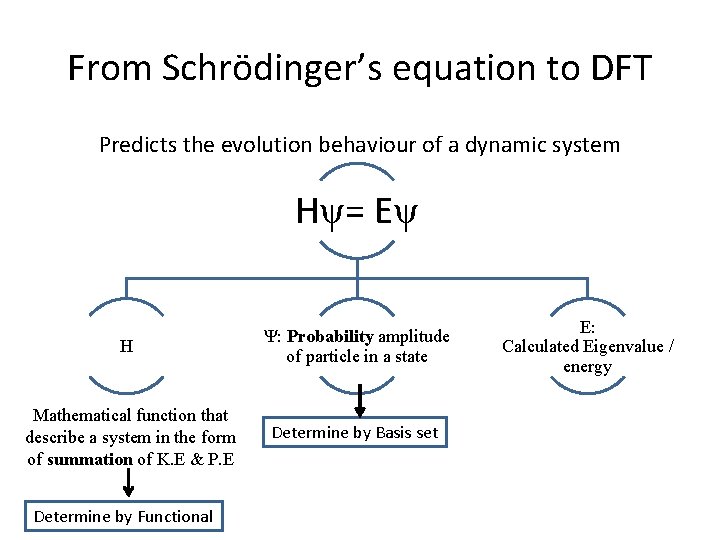
From Schrödinger’s equation to DFT Predicts the evolution behaviour of a dynamic system Hψ= Eψ H Ψ: Probability amplitude of particle in a state Mathematical function that describe a system in the form of summation of K. E & P. E Determine by Basis set Determine by Functional E: Calculated Eigenvalue / energy
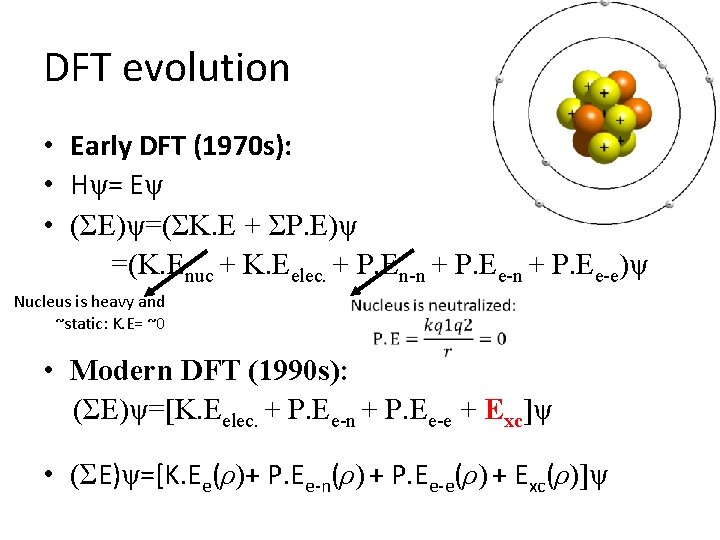
DFT evolution • Early DFT (1970 s): • Hψ= Eψ • (ΣE)ψ=(ΣK. E + ΣP. E)ψ =(K. Enuc + K. Eelec. + P. En-n + P. Ee-e)ψ Nucleus is heavy and ~static: K. E= ~0 • Modern DFT (1990 s): (ΣE)ψ=[K. Eelec. + P. Ee-n + P. Ee-e + Exc]ψ • (ΣE)ψ=[K. Ee(ρ)+ P. Ee-n(ρ) + P. Ee-e(ρ) + Exc(ρ)]ψ
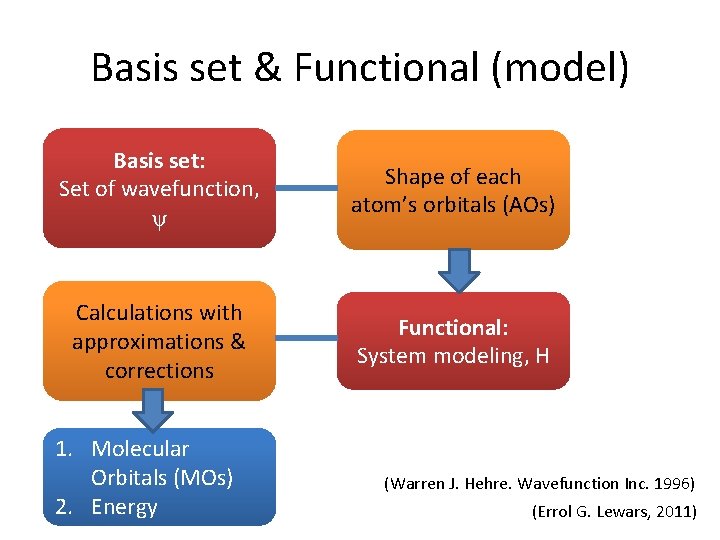
Basis set & Functional (model) Basis set: Set of wavefunction, ψ Shape of each atom’s orbitals (AOs) Calculations with approximations & corrections Functional: System modeling, H 1. Molecular Orbitals (MOs) 2. Energy (Warren J. Hehre. Wavefunction Inc. 1996) (Errol G. Lewars, 2011)
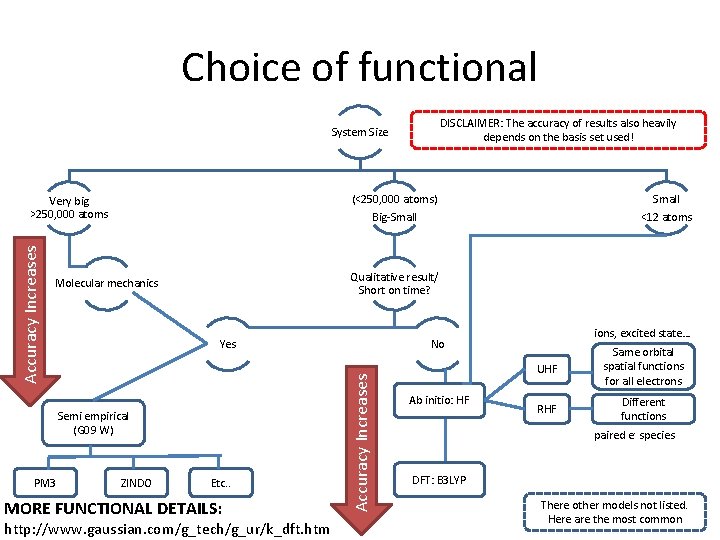
Choice of functional DISCLAIMER: The accuracy of results also heavily depends on the basis set used! System Size (<250, 000 atoms) Big-Small Yes Semi empirical (G 09 W) PM 3 ZINDO Small <12 atoms Qualitative result/ Short on time? Molecular mechanics Etc. . MORE FUNCTIONAL DETAILS: http: //www. gaussian. com/g_tech/g_ur/k_dft. htm ions, excited state… No Accuracy Increases Very big >250, 000 atoms Ab initio: HF UHF Same orbital spatial functions for all electrons RHF Different functions paired e- species DFT: B 3 LYP There other models not listed. Here are the most common
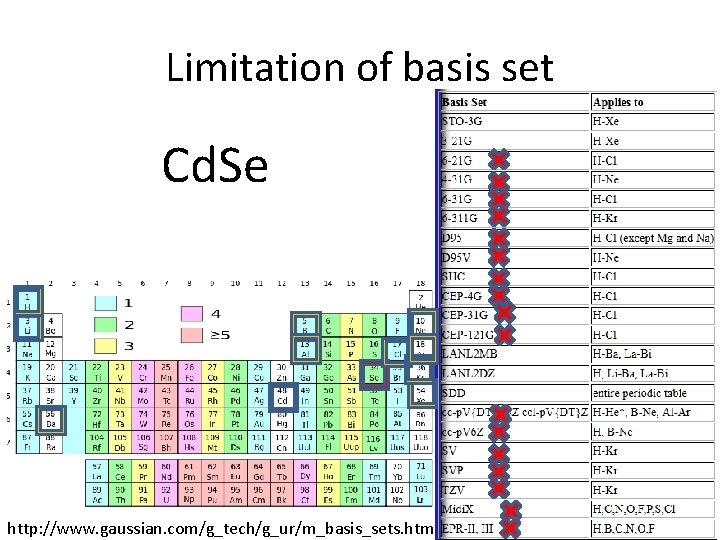
Limitation of basis set Cd. Se http: //www. gaussian. com/g_tech/g_ur/m_basis_sets. htm
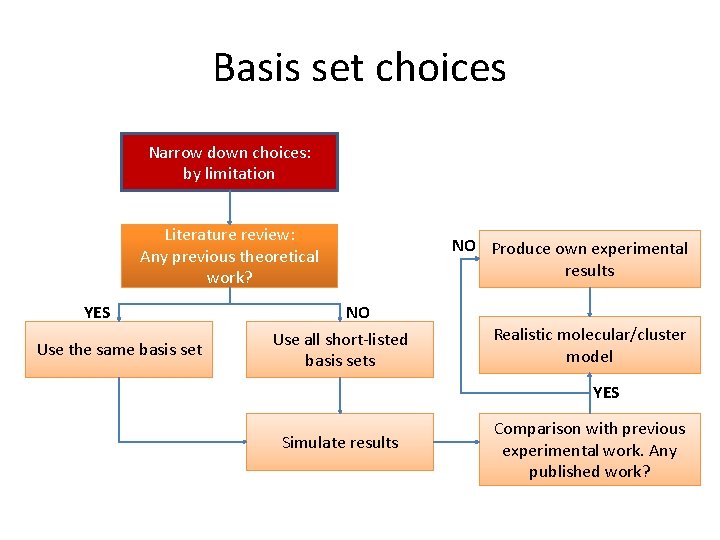
Basis set choices Narrow down choices: by limitation Literature review: Any previous theoretical work? YES Use the same basis set NO Use all short-listed basis sets NO Produce own experimental results Realistic molecular/cluster model YES Simulate results Comparison with previous experimental work. Any published work?

Basis set accuracy: comparison with experimental data Dye 1 Ligand-QD Literature Dye 2 Cd. Se QD List of basis set: http: //www. gaussian. com/g_tech/g_ur/m_basis_sets. htm
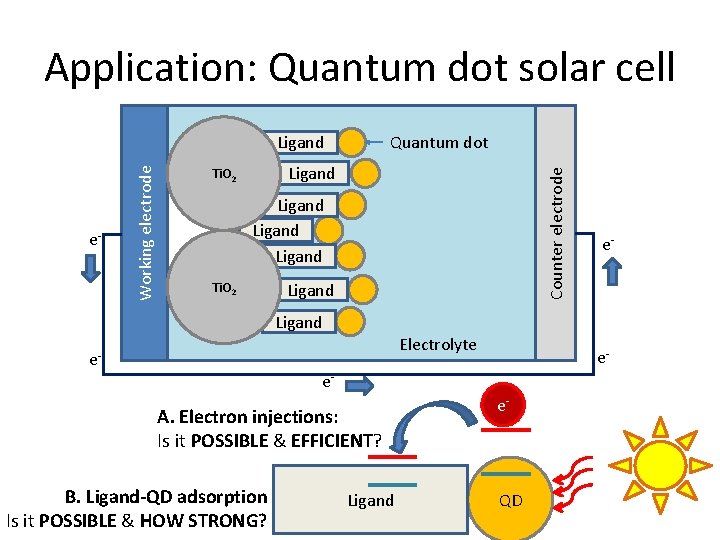
Application: Quantum dot solar cell Quantum dot Ligand Ti. O 2 Counter electrode e- Working electrode Ligand Ti. O 2 Ligand e- Ligand Electrolyte e- e- e- A. Electron injections: Is it POSSIBLE & EFFICIENT? B. Ligand-QD adsorption Is it POSSIBLE & HOW STRONG? Ligand e- QD
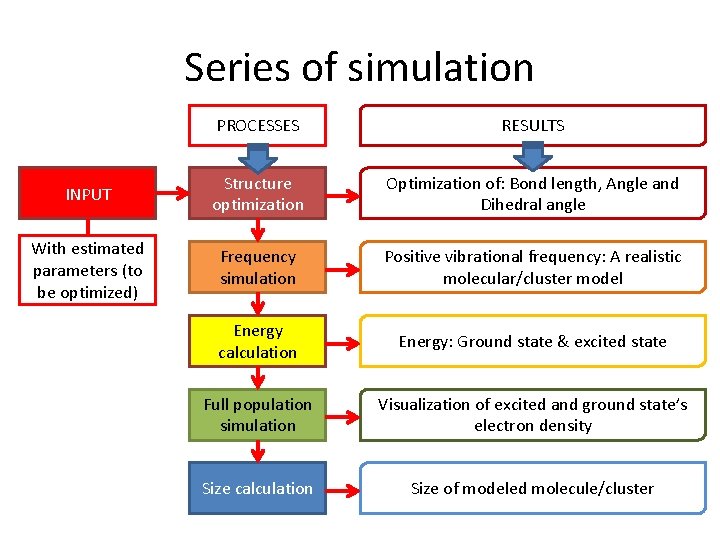
Series of simulation PROCESSES RESULTS INPUT Structure optimization Optimization of: Bond length, Angle and Dihedral angle With estimated parameters (to be optimized) Frequency simulation Positive vibrational frequency: A realistic molecular/cluster model Energy calculation Energy: Ground state & excited state Full population simulation Visualization of excited and ground state’s electron density Size calculation Size of modeled molecule/cluster

Input preparations • Input: – Molecules (i. e. , ligand molecules) – Quasi-Crystals (Quantum dots semiconductor) • Drawing Tools: – Chem. Draw – Chem 3 D – Gaussview • Ab initio DFT Tool: Gaussian 09 W

Input: Ligand molecule What is a molecule? A group of atoms bonded together, representing the smallest fundamental unit of a chemical compound that can take part in a chemical reaction Merriam-Webster Dictionary A single molecule can represents a system consists of ten/hundreds/thousands/millions/billions of them
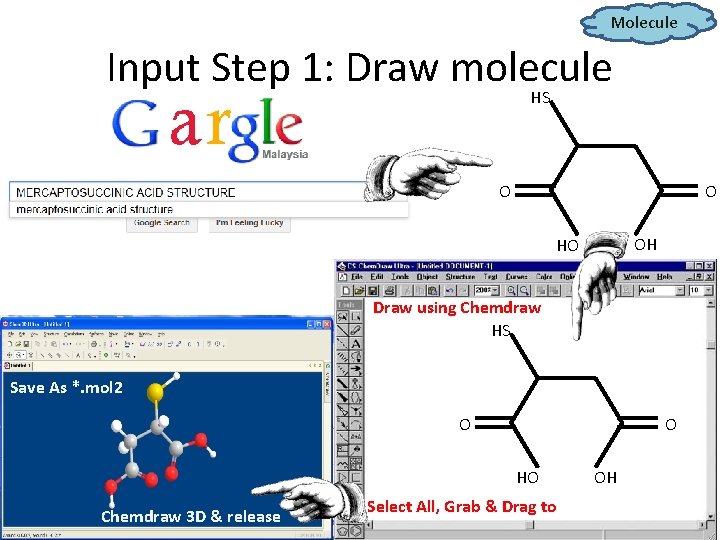
Molecule Input Step 1: Draw molecule HS O O OH HO Draw using Chemdraw HS Save As *. mol 2 O O HO Chemdraw 3 D & release Select All, Grab & Drag to OH
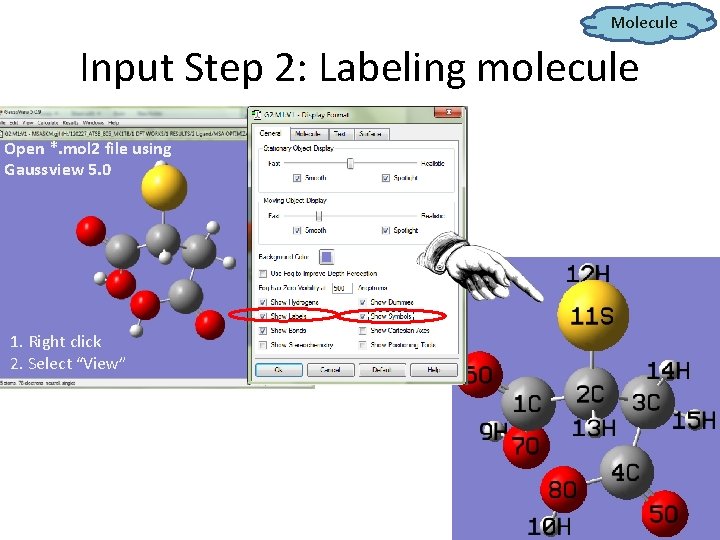
Molecule Input Step 2: Labeling molecule Open *. mol 2 file using Gaussview 5. 0 1. Right click 2. Select “View”

Molecule Input Step 3: Rearrange numbers • Click “edit” – “Atom list” • A new interface appears: showing labels. Rearrange the numbers in a nice flow. Computer generated Z-Matrix: Pro: Can be straight away use as simulation input. Con: Tested & may cause longer simulation time. Solution: Spend time building our own Z-Matrix Before & After
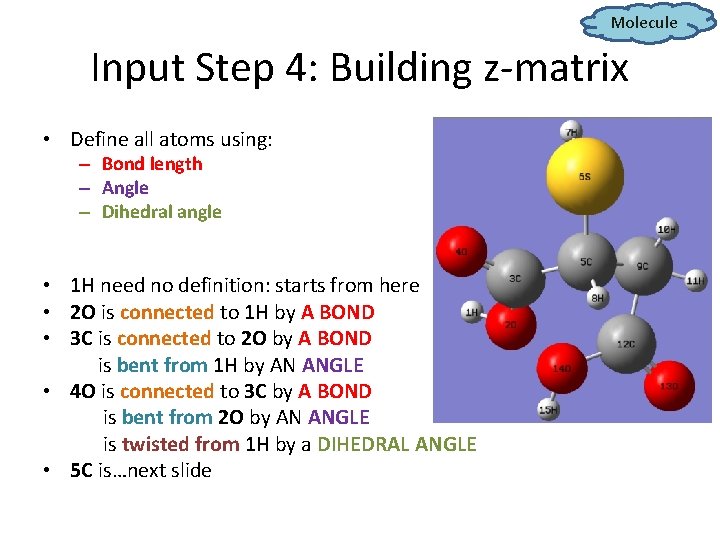
Molecule Input Step 4: Building z-matrix • Define all atoms using: – Bond length – Angle – Dihedral angle • 1 H need no definition: starts from here • 2 O is connected to 1 H by A BOND • 3 C is connected to 2 O by A BOND is bent from 1 H by AN ANGLE • 4 O is connected to 3 C by A BOND is bent from 2 O by AN ANGLE is twisted from 1 H by a DIHEDRAL ANGLE • 5 C is…next slide
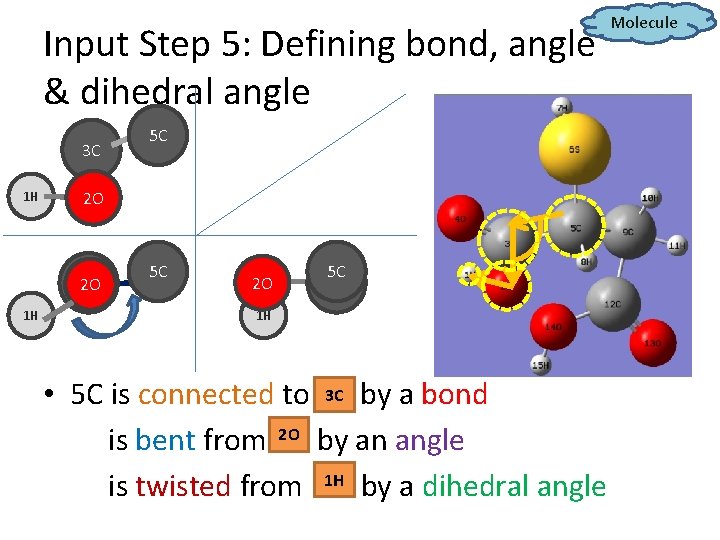
Input Step 5: Defining bond, angle & dihedral angle 3 C 1 H 2 O 3 C 2 O 1 H 5 C 5 C 2 O 5 C 3 C 1 H • 5 C is connected to 3 C? by a bond is bent from 2 O? by an angle is twisted from 1 H? by a dihedral angle Molecule

Molecule Input Step 6: Z-matrix, the format 1 H 2 O is connected to 1 H by A BOND 3 C is connected to 2 O by A BOND is bent from 1 H by AN ANGLE 4 O is connected to 3 C by A BOND is bent from 2 O by AN ANGLE is twisted from 1 H by a DIHEDRAL ANGLE H O 1 B 1 C 2 B 2 1 A 1 O 3 B 3 2 A 2 1 D 1 A description of an atom must be: 1. In 1 LINE 2. Each line is meant for 1 atom’s description ONLY 3. May use any symbol for bond, angle and dihedral 4. B 1, B 2, A 1…D 1…. must be stated as simulation parameters and the value will be optimized OR as CONSTANT i. stated in DECIMAL POINT as INDICATION of CONSTANT, i. e. , 1. 546 ii. stated in the form of 1. ONLY as INDICATION of TO BE OPTIMIZED (Will show the full format later)
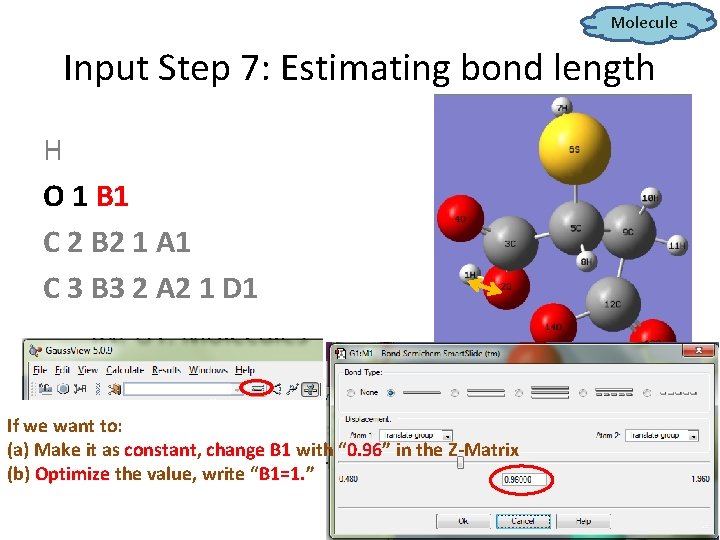
Molecule Input Step 7: Estimating bond length H O 1 B 1 C 2 B 2 1 A 1 C 3 B 3 2 A 2 1 D 1 If we want to: (a) Make it as constant, change B 1 with “ 0. 96” in the Z-Matrix (b) Optimize the value, write “B 1=1. ”
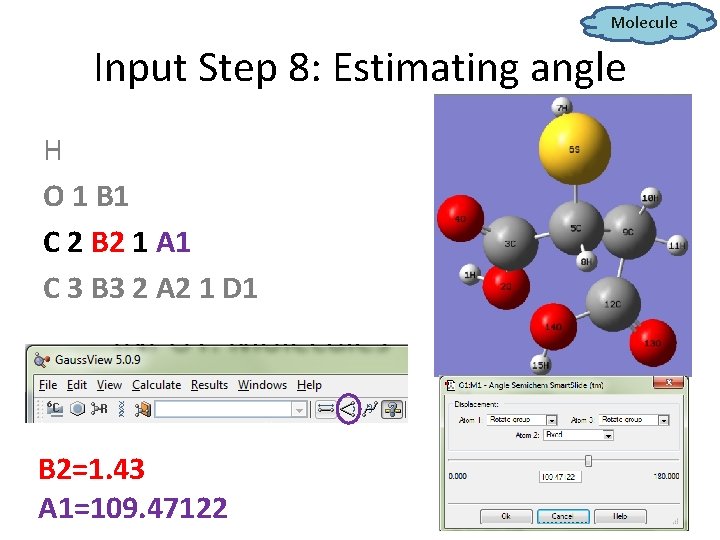
Molecule Input Step 8: Estimating angle H O 1 B 1 C 2 B 2 1 A 1 C 3 B 3 2 A 2 1 D 1 B 2=1. 43 A 1=109. 47122

Molecule Input Step 9: Estimating dihedral angle H O 1 B 1 C 2 B 2 1 A 1 C 3 B 3 2 A 2 1 D 1 B 3=1. 2584 A 2=120. 0 D 1=30. 0
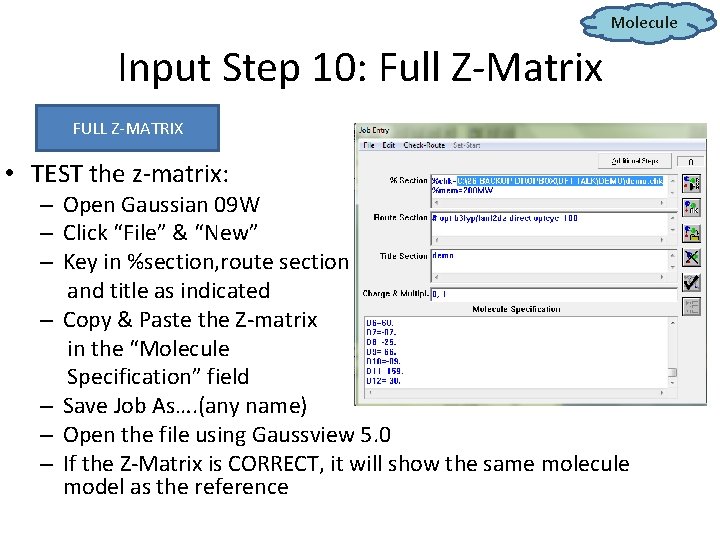
Molecule Input Step 10: Full Z-Matrix FULL Z-MATRIX • TEST the z-matrix: – Open Gaussian 09 W – Click “File” & “New” – Key in %section, route section and title as indicated – Copy & Paste the Z-matrix in the “Molecule Specification” field – Save Job As…. (any name) – Open the file using Gaussview 5. 0 – If the Z-Matrix is CORRECT, it will show the same molecule model as the reference
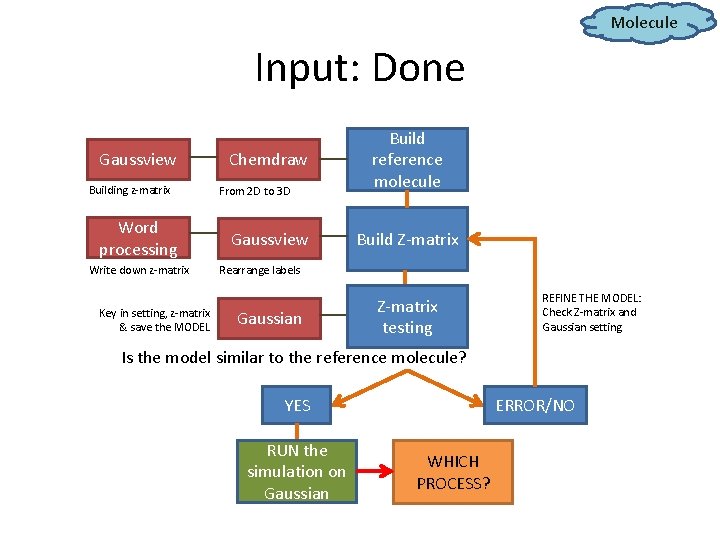
Molecule Input: Done Gaussview Building z-matrix Word processing Write down z-matrix Key in setting, z-matrix & save the MODEL Chemdraw From 2 D to 3 D Gaussview Build reference molecule Build Z-matrix Rearrange labels Gaussian Z-matrix testing REFINE THE MODEL: Check Z-matrix and Gaussian setting Is the model similar to the reference molecule? YES RUN the simulation on Gaussian ERROR/NO WHICH PROCESS?
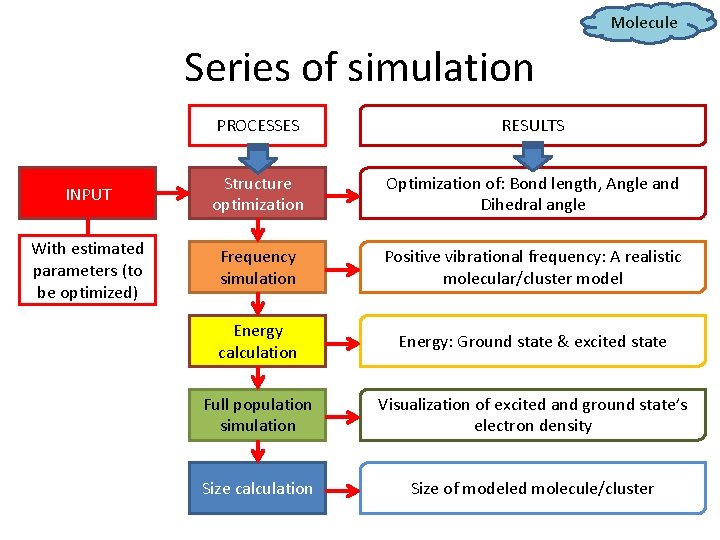
Molecule Series of simulation PROCESSES RESULTS INPUT Structure optimization Optimization of: Bond length, Angle and Dihedral angle With estimated parameters (to be optimized) Frequency simulation Positive vibrational frequency: A realistic molecular/cluster model Energy calculation Energy: Ground state & excited state Full population simulation Visualization of excited and ground state’s electron density Size calculation Size of modeled molecule/cluster
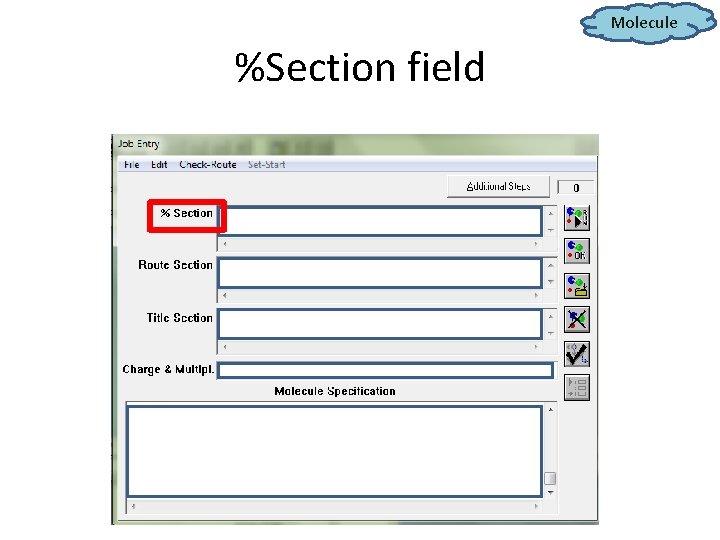
Molecule %Section field

Molecule %Section field • The directory to save “. chk” file (a file that records all calculations, achievable at any simulation process by using “check” command in “Route Section”): %chk= C: g 09 wPb. Te. chk • Stating the amount of memory usage in MW (megawords). 1 MW=3. 81 MBytes %mem=200 MW
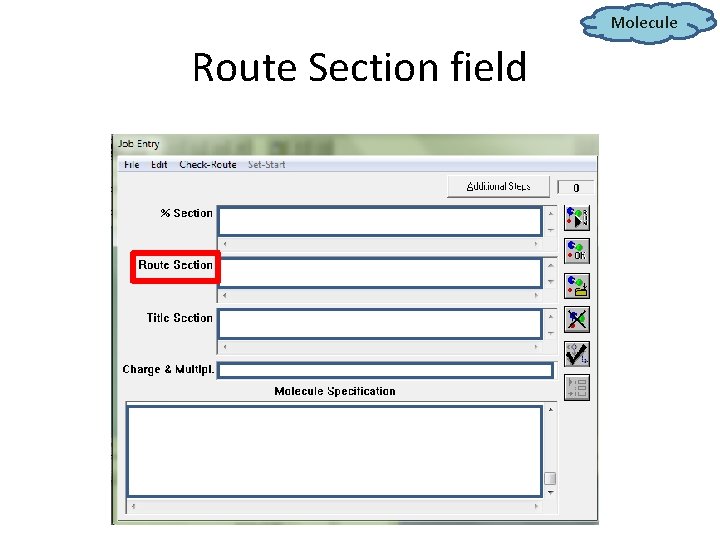
Molecule Route Section field
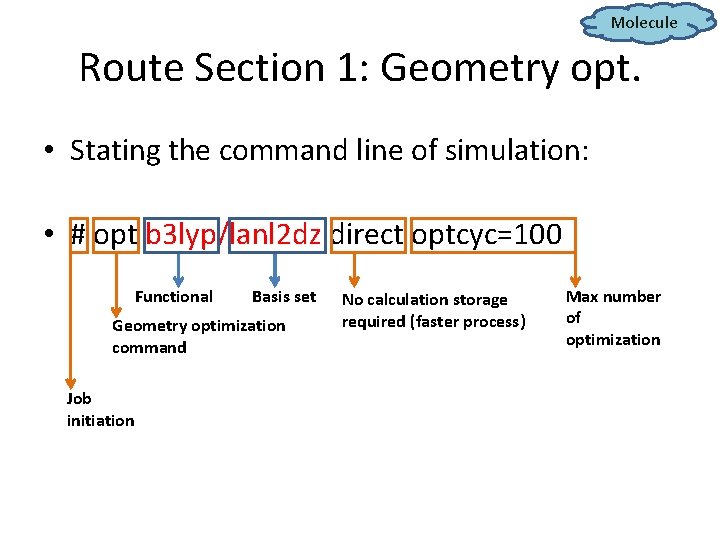
Molecule Route Section 1: Geometry opt. • Stating the command line of simulation: • # opt b 3 lyp/lanl 2 dz direct optcyc=100 Functional Basis set Geometry optimization command Job initiation No calculation storage required (faster process) Max number of optimization

Molecule Route Section 2: Frequency • Stating the command line of simulation: • #n b 3 lyp/lanl 2 dz direct freq geom=check guess=check Default: Normal print level of output Frequency command Retrieve molecular orbitals data from “. chk” file (previous geometry opt. ) Retrieve internal coordinate from “. chk” file (previous geometry opt. ) Without “geom=check” and “guess=check” command: Have to state “Optimized Z-matrix” in the “Molecule Spec” field
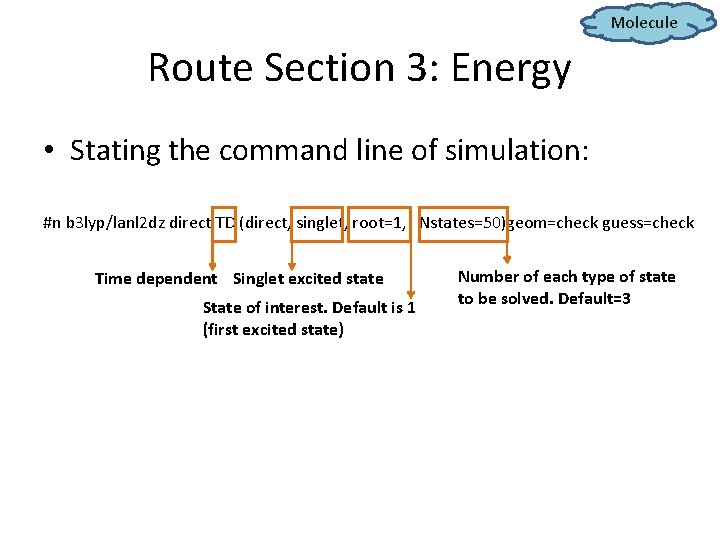
Molecule Route Section 3: Energy • Stating the command line of simulation: #n b 3 lyp/lanl 2 dz direct TD (direct, singlet, root=1, Nstates=50)geom=check guess=check Time dependent Singlet excited state State of interest. Default is 1 (first excited state) Number of each type of state to be solved. Default=3

Molecule Route Section 4: Full population • Stating the command line of simulation: #n b 3 lyp/lanl 2 dz pop=full geom=check guess=check Molecular orbitals visualization, total atomic charge and orbital energies

Route Section 5: Cluster size estimation Molecule • Stating the command line of simulation: Calculate volume & surface area of cluster in solution. Default: Water #p b 3 lyp/lanl 2 dz scrf=pcm geom=check Additional output (execution timing, messages etc) Use with energy, geometry opt, freq calc to model systems in solution • Choices of solvent (to be specified in command line): http: //www. gaussian. com/g_tech/g_ur/k_scrf. htm
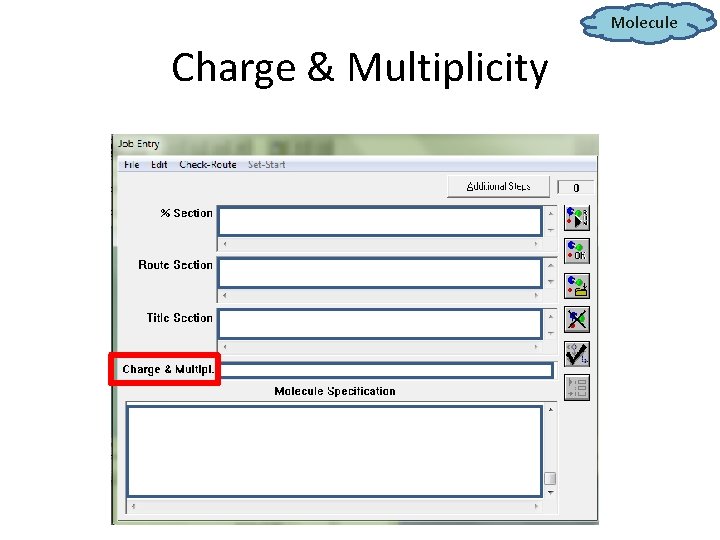
Molecule Charge & Multiplicity
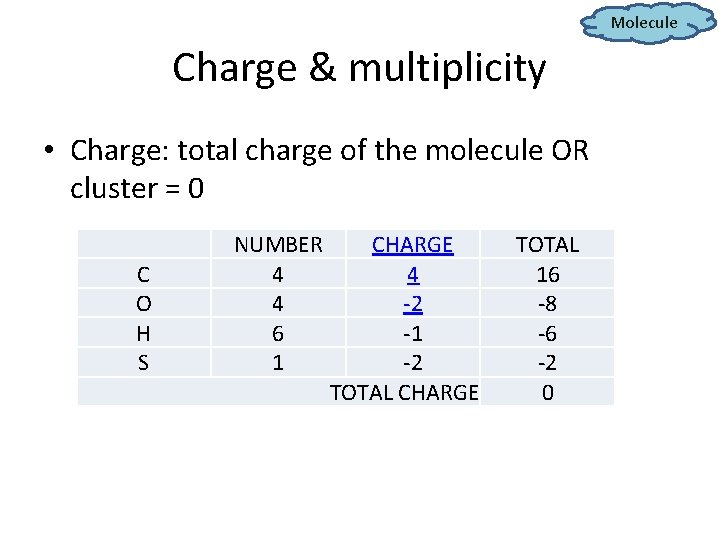
Molecule Charge & multiplicity • Charge: total charge of the molecule OR cluster = 0 C O H S NUMBER 4 4 6 1 CHARGE 4 -2 -1 -2 TOTAL CHARGE TOTAL 16 -8 -6 -2 0

Molecule Charge & multiplicity • Multiplicity/spin multiplicity: describes how the electrons of the system exist • 2 S+1=spin multiplicity, where S is total spin quantum number • S=n(1/2) where n=unpaired electron H H H S H O Inorganic Chemistry J. E. House, Academic Press 2008 O H O O mercaptosuccinic acid molecule H
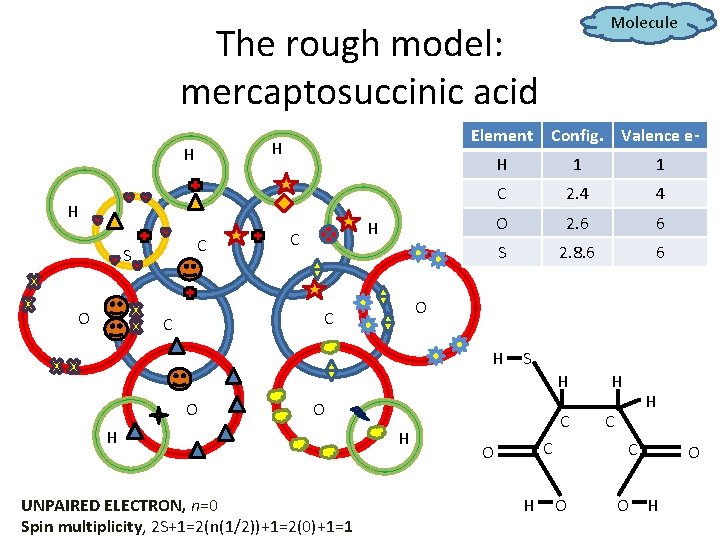
Molecule The rough model: mercaptosuccinic acid Element H H H C S O H C H 1 1 C 2. 4 4 O 2. 6 6 S 2. 8. 6 6 O C C Config. Valence e- H S H O O H UNPAIRED ELECTRON, n=0 Spin multiplicity, 2 S+1=2(n(1/2))+1=2(0)+1=1 H C C O H H H C C O O O H

Molecule Running the simulation Specify *. chk directory & memory Command line Your desired title Molecule/cluster charge & spin multiplicity Molecule/cluster’s Z-Matrix 1. Click RUN 2. Pop up message: specifying output directory
Extracting result: Structure optimization Molecule Optimization complete Double click the output Work doesn’t kill, But worry does -unknown post graduate student Frequency simulation Open in Gaussview Energy calculation Save as Gaussian Input Files (. gjf) Full population simulation Input for NEXT SIMULATION Size calculation
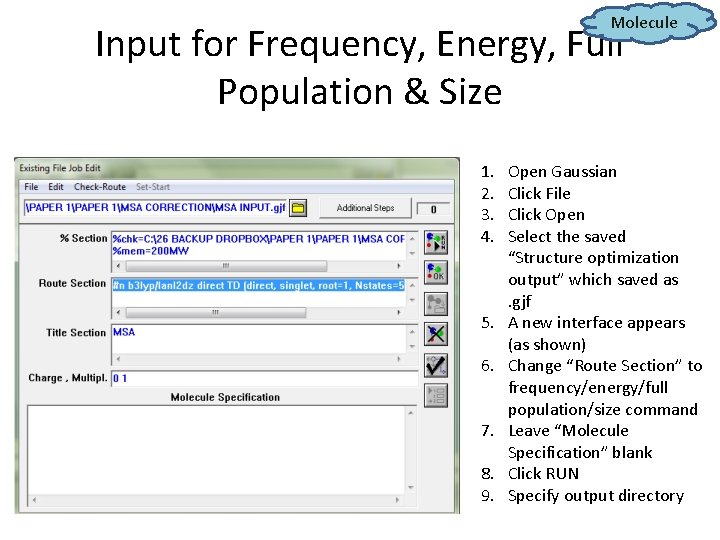
Molecule Input for Frequency, Energy, Full Population & Size 1. 2. 3. 4. 5. 6. 7. 8. 9. Open Gaussian Click File Click Open Select the saved “Structure optimization output” which saved as. gjf A new interface appears (as shown) Change “Route Section” to frequency/energy/full population/size command Leave “Molecule Specification” blank Click RUN Specify output directory
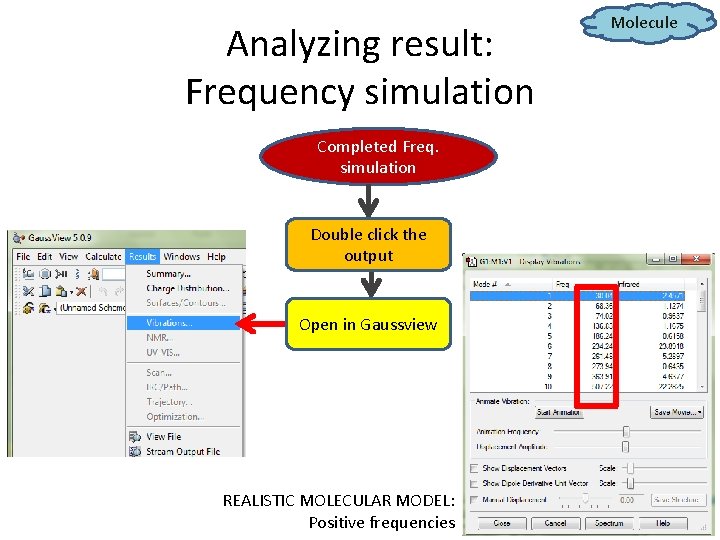
Analyzing result: Frequency simulation Completed Freq. simulation Double click the output Open in Gaussview REALISTIC MOLECULAR MODEL: Positive frequencies Molecule
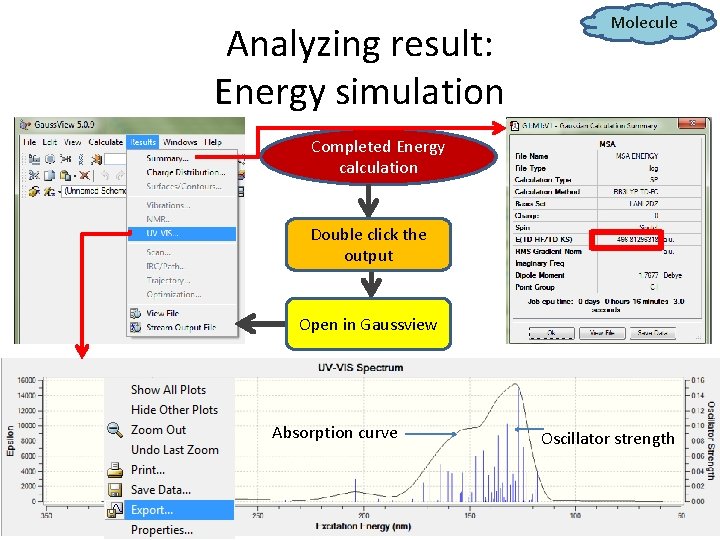
Analyzing result: Energy simulation Molecule Completed Energy calculation Double click the output Open in Gaussview Absorption curve Oscillator strength
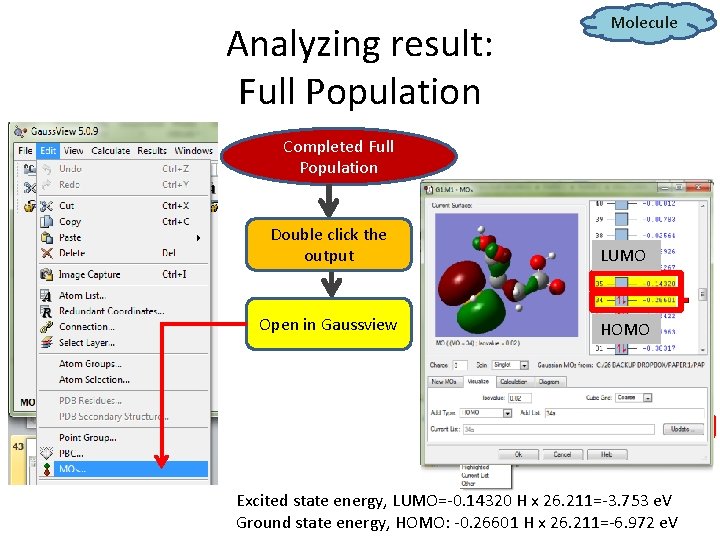
Analyzing result: Full Population Molecule Completed Full Population Double click the output LUMO Open in Gaussview HOMO Excited state energy, LUMO=-0. 14320 H x 26. 211=-3. 753 e. V Ground state energy, HOMO: -0. 26601 H x 26. 211=-6. 972 e. V
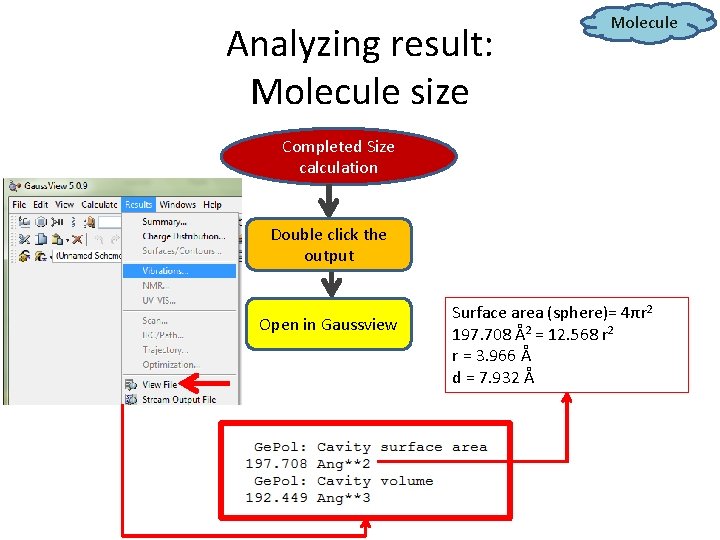
Analyzing result: Molecule size Molecule Completed Size calculation Double click the output Open in Gaussview Surface area (sphere)= 4πr 2 197. 708 Å2 = 12. 568 r 2 r = 3. 966 Å d = 7. 932 Å

Crystal Input: QD semiconductor • BULK semiconductor: – Crystal – Bond length, angle and dihedral angle are constant • QUANTUM DOT semiconductor: – Quasi-crystal – Bond length, angle & dihedral angle change due to surface relaxations – FULL OPTIMIZATION (bond length, angle & dihedral) Puzder et al Phys. Rev. Lett. 92, 217401 (2004) Self-Healing of Cd. Se Nanocrystals: First. Principles Calculations

Crystal Input Step 1: Download CIF • Download (example: Cd. Se) CIF file from: www. crystallography. net • Based on the CIF, build our own cluster
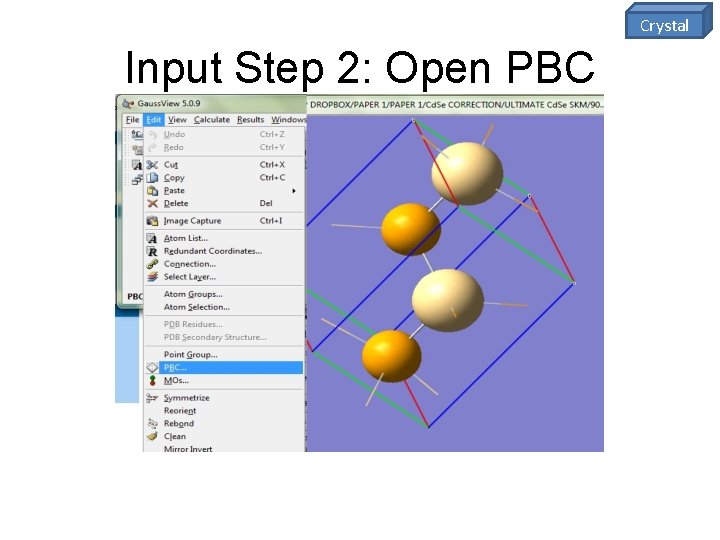
Crystal Input Step 2: Open PBC
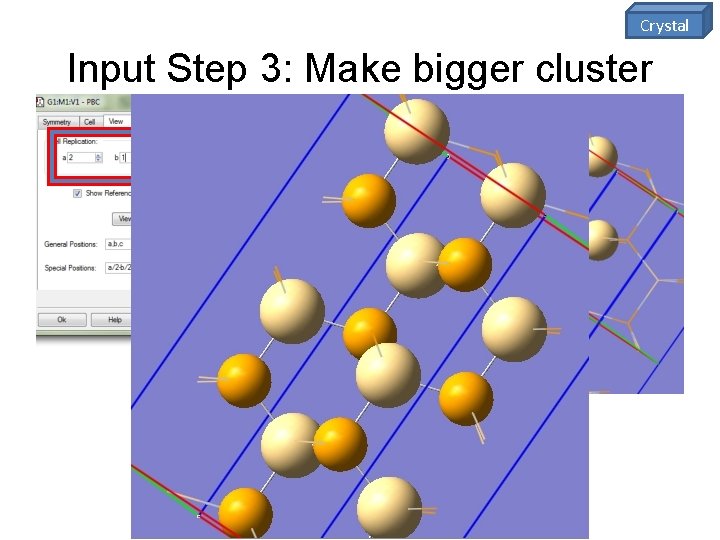
Crystal Input Step 3: Make bigger cluster
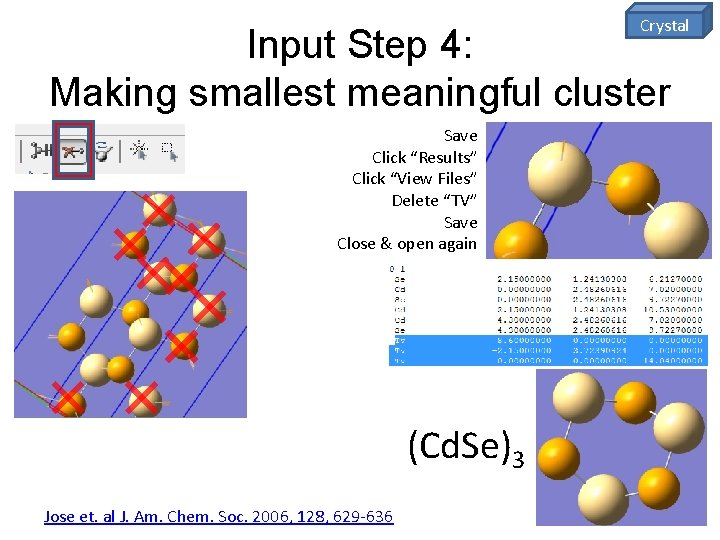
Crystal Input Step 4: Making smallest meaningful cluster Save Click “Results” Click “View Files” Delete “TV” Save Close & open again (Cd. Se)3 Jose et. al J. Am. Chem. Soc. 2006, 128, 629 -636

Crystal Spin multiplicity • Singlet State: 2 S+1=1 – 2(nx 1/2)+1=1 – n=0 (no unpaired electrons/all electrons are paired) – No resultant magnetic moment – Magnetic moment produced by electron spin +1/2 and -1/2 cancel out each other – The materials becomes diamagnetic – Produce slight magnetic field opposing external magnetic field – Bulk Cd. Se is a diamagnetic material Neeleshwar et. al. Physical Review B, 2005
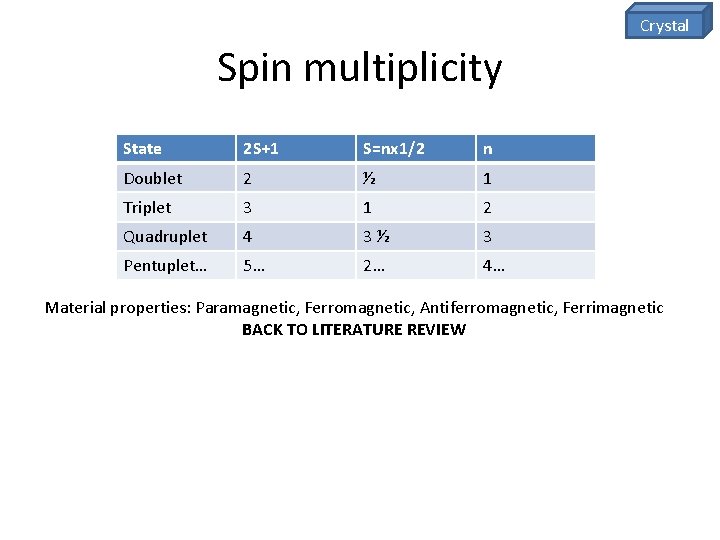
Crystal Spin multiplicity State 2 S+1 S=nx 1/2 n Doublet 2 ½ 1 Triplet 3 1 2 Quadruplet 4 3½ 3 Pentuplet… 5… 2… 4… Material properties: Paramagnetic, Ferromagnetic, Antiferromagnetic, Ferrimagnetic BACK TO LITERATURE REVIEW
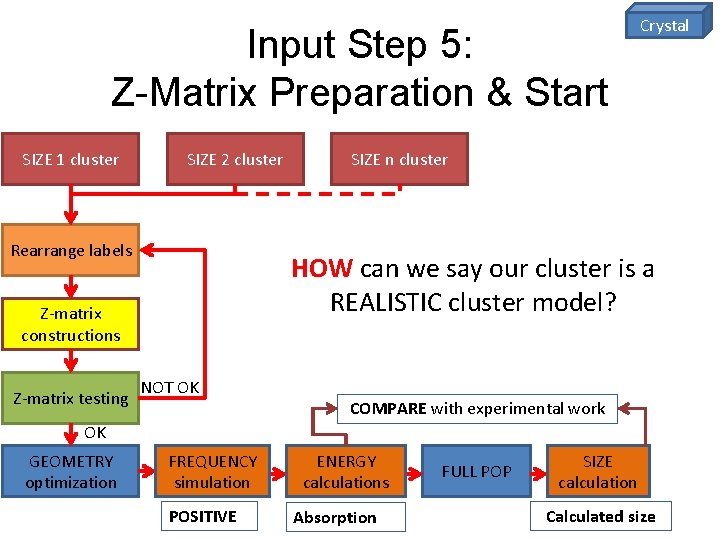
Input Step 5: Z-Matrix Preparation & Start SIZE 1 cluster SIZE 2 cluster Rearrange labels SIZE n cluster HOW can we say our cluster is a REALISTIC cluster model? Z-matrix constructions Z-matrix testing Crystal NOT OK COMPARE with experimental work OK GEOMETRY optimization FREQUENCY simulation POSITIVE ENERGY calculations Absorption FULL POP SIZE calculation Calculated size
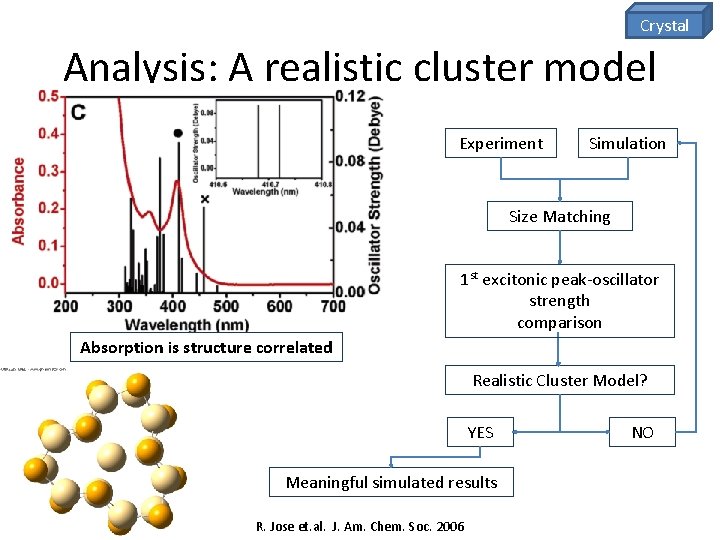
Crystal Analysis: A realistic cluster model Experiment Simulation Size Matching 1 st excitonic peak-oscillator strength comparison Absorption is structure correlated Realistic Cluster Model? YES Meaningful simulated results R. Jose et. al. J. Am. Chem. Soc. 2006 NO
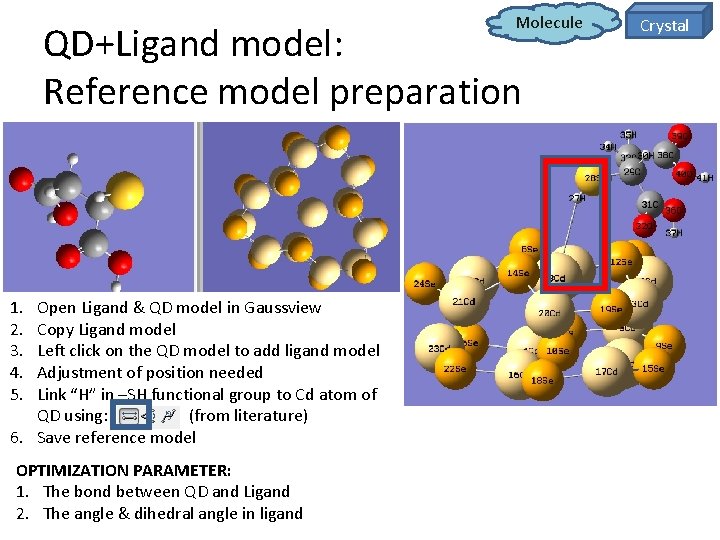
Molecule QD+Ligand model: Reference model preparation 1. 2. 3. 4. 5. Open Ligand & QD model in Gaussview Copy Ligand model Left click on the QD model to add ligand model Adjustment of position needed Link “H” in –SH functional group to Cd atom of QD using: (from literature) 6. Save reference model OPTIMIZATION PARAMETER: 1. The bond between QD and Ligand 2. The angle & dihedral angle in ligand Crystal

Molecule QD+Ligand model: z-matrix preparations QD’s Z-matrix: With pre-optimized bond length, angle & dihedral angle Ligand’s Z-matrix: With fixed bond length. Angle & dihedral angle are to be optimized 1. Change ligand’s Zmatrix numbering by adding 26 to the current reference number 2. Ligand’s atom number 1 will be = 1+26=27 3. 2 will be 2+26=28…so on & so forth This Se is the 26 th defined atom Crystal
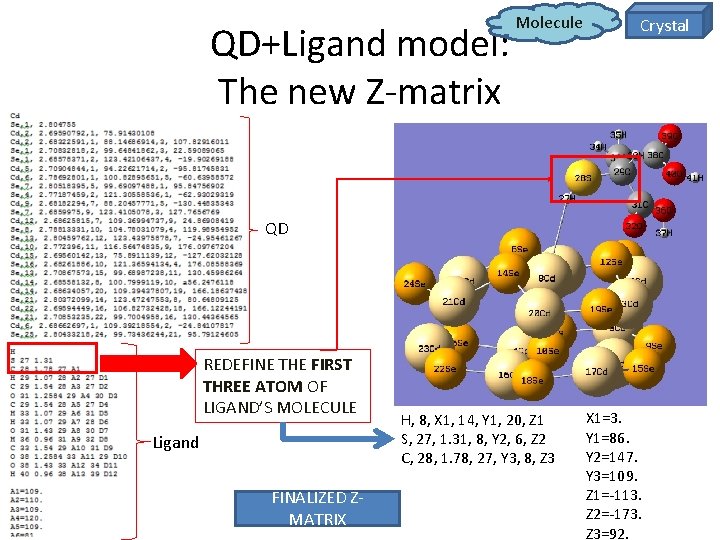
QD+Ligand model: The new Z-matrix Molecule Crystal QD REDEFINE THE FIRST THREE ATOM OF LIGAND’S MOLECULE Ligand FINALIZED ZMATRIX H, 8, X 1, 14, Y 1, 20, Z 1 S, 27, 1. 31, 8, Y 2, 6, Z 2 C, 28, 1. 78, 27, Y 3, 8, Z 3 X 1=3. Y 1=86. Y 2=147. Y 3=109. Z 1=-113. Z 2=-173. Z 3=92.

Result analyses: Magic size cluster S C • Determination of electronically stable QD structure for synthesis. (1. 5 nm in Cd. Se): – Largest quantities with identical properties – Efficient electron injections (Cd. Se)6 (Cd. Se)13 WHICH ONE OF THESE CLUSTERS, ELECTRONICALLY STABLE? R. Jose et al & Saifful et al (Cd. Se)16
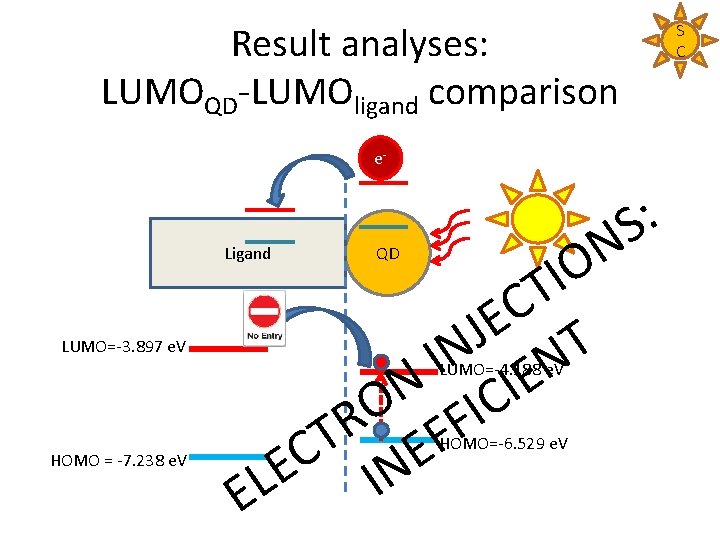
Result analyses: LUMOQD-LUMOligand comparison e- : S N O I T C E J T N I N E N I C O I R F T F E C E IN L E Ligand LUMO=-3. 897 e. V QD LUMO=-4. 188 e. V HOMO = -7. 238 e. V HOMO=-6. 529 e. V S C
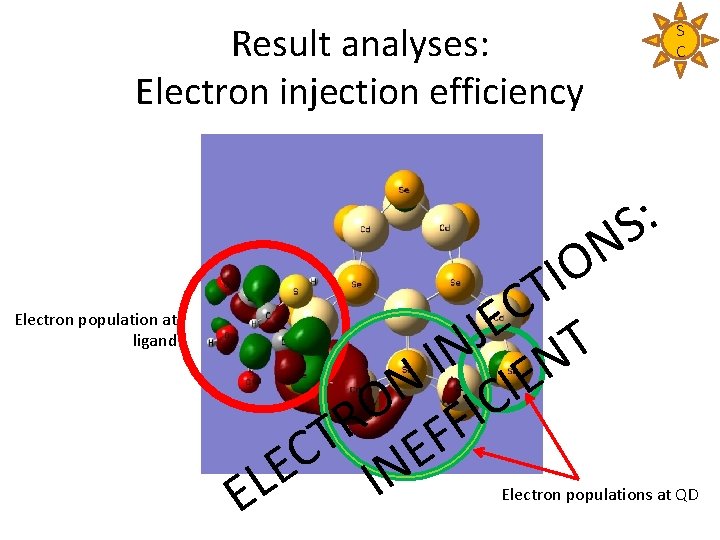
Result analyses: Electron injection efficiency Electron population at ligand : S N O I T C E J T N I N E N I C O I R F T F E C E IN L E S C Electron populations at QD
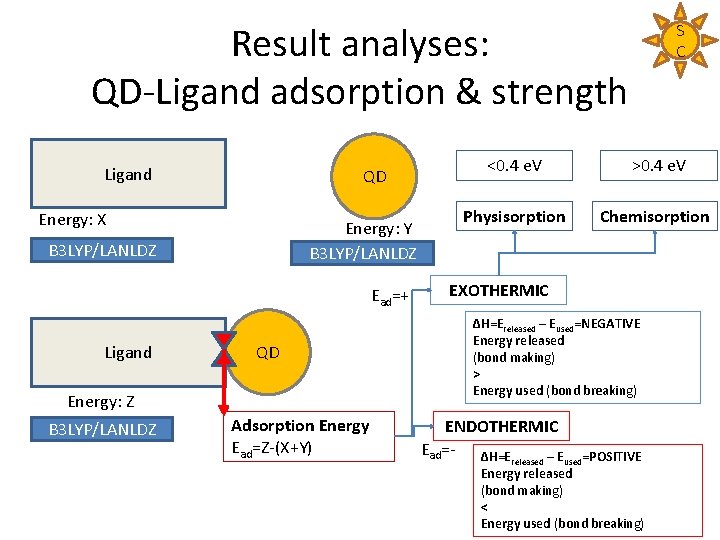
Result analyses: QD-Ligand adsorption & strength Ligand QD Energy: X Energy: Y B 3 LYP/LANLDZ Ead=+ Ligand QD Energy: Z B 3 LYP/LANLDZ Adsorption Energy Ead=Z-(X+Y) S C <0. 4 e. V >0. 4 e. V Physisorption Chemisorption EXOTHERMIC ΔH=Ereleased – Eused=NEGATIVE Energy released (bond making) > Energy used (bond breaking) ENDOTHERMIC Ead=- ΔH=E –E released used=POSITIVE Energy released (bond making) < Energy used (bond breaking)
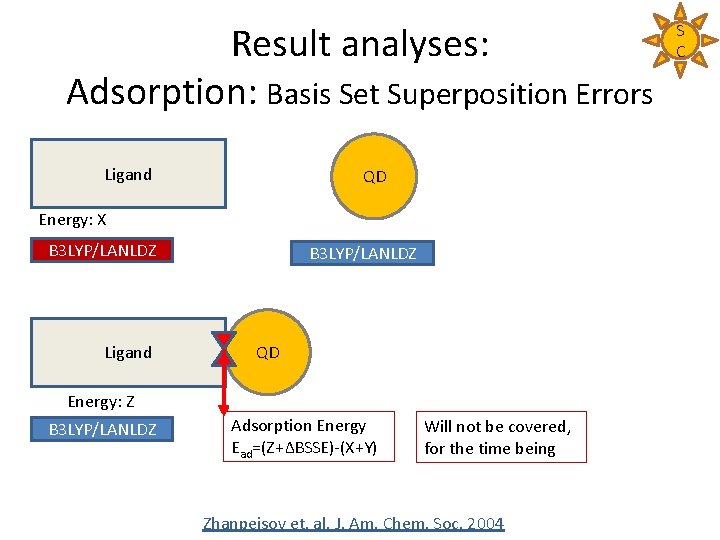
Result analyses: Adsorption: Basis Set Superposition Errors Ligand QD Energy: X B 3 LYP/LANLDZ Ligand B 3 LYP/LANLDZ QD Energy: Z B 3 LYP/LANLDZ Adsorption Energy Ead=(Z+ΔBSSE)-(X+Y) Will not be covered, for the time being Zhanpeisov et. al. J. Am. Chem. Soc. 2004 S C

DFT Kick Start (David S. Sholl et. al. 2009)
- Slides: 67