Ability to genotype differing variants with arrays vs

Ability to genotype differing variants with arrays vs. whole genome sequencing P. M. Van. Raden 1, G. L. Spangler 1, C. P. Van Tassell 1, J. Jiang 2, L. Ma 2, J. R. O’Connell 3, S. Smith 4, and S. K. De. Nise 4 1 USDA-ARS-AGIL, Beltsville, MD, 2 U. Maryland. College Park, 3 U. Maryland-Baltimore, 4 Zoetis, Inc. , Kalamazoo, MI paul. vanraden@ars. usda. gov American Society of Animal Science annual meeting, Baltimore, MD; July 9 -12, 2017 (1) Van. Raden

Questions l Can all sequence SNPs be genotyped using chips? l Can all chip SNPs be genotyped from sequence data? l What properties help predict success or failure? w Illumina design scores, SNP heritability, repetitive DNA location, gene location, allele pattern, MAF l How do these properties affect optimal chip design? (SNPs you want to use vs. SNPs that genotype well) American Society of Animal Science annual meeting, Baltimore, MD; July 9 -12, 2017 (2) Van. Raden

Motivation l Chip design has usually selected the highest quality SNPs to use as markers (50 K, HD, LD) l Newer chips began adding preselected QTLs, not just markers, to better track biological effects l SNP effects were estimated directly from sequence l Largest effects were then added to arrays, with no pre-screening for SNP quality l Hypothesis: Works in sequence, should work on chip American Society of Animal Science annual meeting, Baltimore, MD; July 9 -12, 2017 (3) Van. Raden

Sequence vs. array genotype chemistry l Arrays (SNP chips) w Alleles attach to beads, indicating the 3 genotypes w Each allele should have none, half, or all attached l Sequence w Physically read ~150 bases at both ends of a DNA segment ~1000 bases in length w Multiple reads are needed to detect both alleles American Society of Animal Science annual meeting, Baltimore, MD; July 9 -12, 2017 (4) Van. Raden

SNP selection from sequence data l Run 5 genotypes from 1000 Bulls Project l 39 million SNPs for 440 sequenced Holsteins l 1 million used after edits for minor allele frequency, gene location, and linkage disequilibrium l 26, 970 bulls with 50 K or HD imputed to sequence were used in SNP selection American Society of Animal Science annual meeting, Baltimore, MD; July 9 -12, 2017 (5) Van. Raden
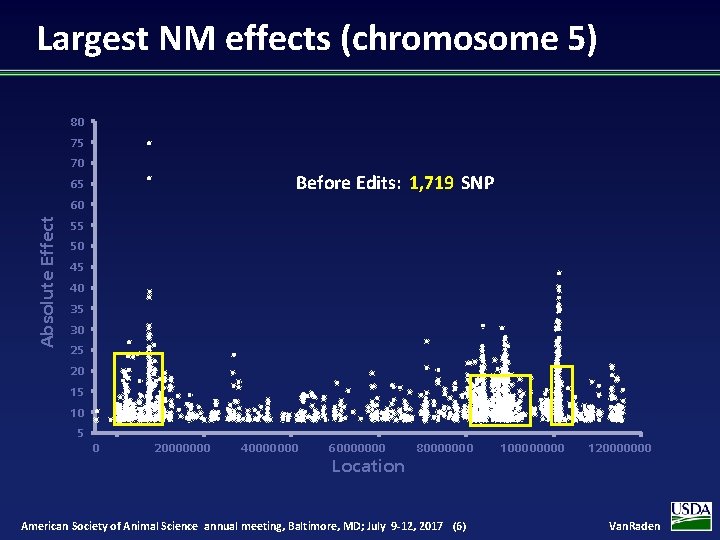
Largest NM effects (chromosome 5) 80 75 70 Before Edits: 1, 719 SNP 65 Absolute Effect 60 55 50 45 40 35 30 25 20 15 10 5 0 20000000 40000000 60000000 Location 80000000 American Society of Animal Science annual meeting, Baltimore, MD; July 9 -12, 2017 (6) 10000 120000000 Van. Raden
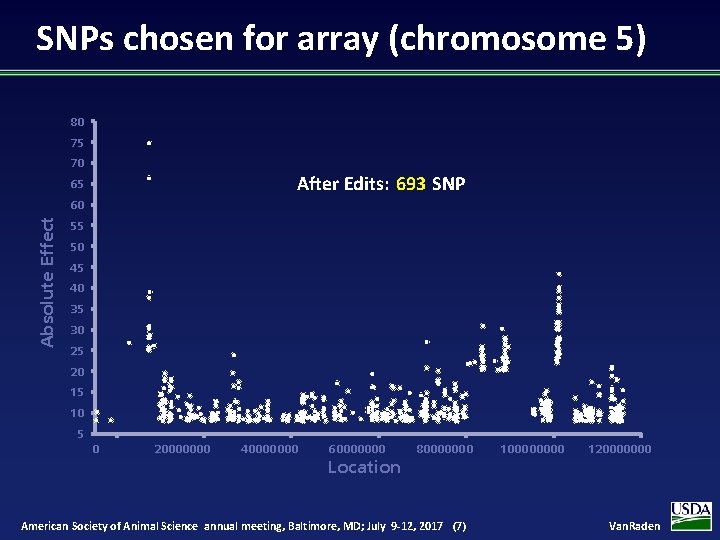
SNPs chosen for array (chromosome 5) 80 75 70 After Edits: 693 SNP 65 Absolute Effect 60 55 50 45 40 35 30 25 20 15 10 5 0 20000000 40000000 60000000 Location 80000000 American Society of Animal Science annual meeting, Baltimore, MD; July 9 -12, 2017 (7) 10000 120000000 Van. Raden

SNPs attempted to place on array l SNP selection described in 2017 GSE 49: 32 w 4, 821 SNPs with largest effects added to Zoetis low density chip, version 5 ( ZL 5) w 1, 601 new SNPs from sequence w 3, 220 from HD American Society of Animal Science annual meeting, Baltimore, MD; July 9 -12, 2017 (8) Van. Raden

Results: SNPs passing quality control l l Success rates for selected SNPs added to Zoetis ZL 5 w 96% for SNPs selected from Bovine HD chip w 64% for new SNPs selected from sequence data What causes new SNPs to fail QC? w Examine correlations of pass/fail (0, 1) status with several SNP properties American Society of Animal Science annual meeting, Baltimore, MD; July 9 -12, 2017 (9) Van. Raden

SNP properties tested l Illumina design scores (official prediction of success) l Estimated heritability of SNP genotype from sequence l Distance inside a repeated DNA section (using Repeat. Masker) l Location within gene (exon, intron, intergenic, etc. ) l Reference / alternate allele ( transitions, transversions ) l Minor allele frequency in Holstein breed American Society of Animal Science annual meeting, Baltimore, MD; July 9 -12, 2017 (10) Van. Raden

Illumina design scores l Some DNA patterns are easier to read than others l Single strand may loop on itself or with other strands l Example of hairpin loop l Occurs in RNA or DNA l Santa. Lucia, J. , and D. Hicks. 2004. The thermodynamics of DNA structural motifs. Ann. Rev. Biophys. Biomol. Struct. 33: 415– 40. American Society of Animal Science annual meeting, Baltimore, MD; July 9 -12, 2017 (11) Van. Raden
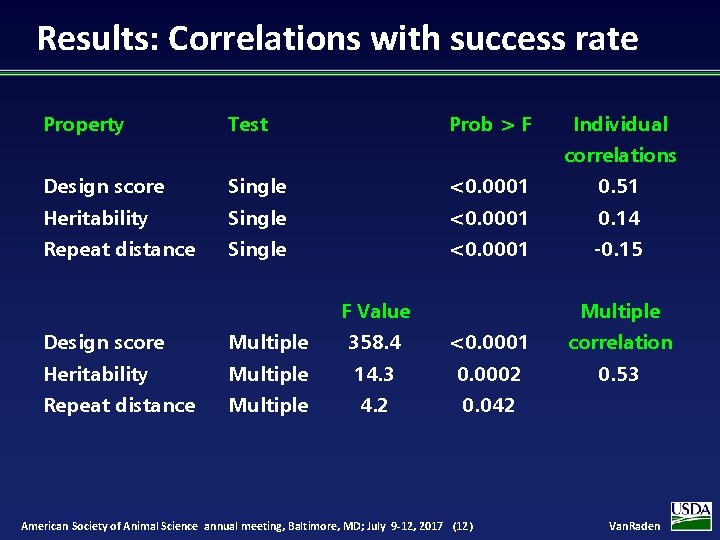
Results: Correlations with success rate Property Test Prob > F Individual correlations Design score Single <0. 0001 0. 51 Heritability Single <0. 0001 0. 14 Repeat distance Single <0. 0001 -0. 15 F Value Multiple Design score Multiple 358. 4 <0. 0001 correlation Heritability Multiple 14. 3 0. 0002 0. 53 Repeat distance Multiple 4. 2 0. 042 American Society of Animal Science annual meeting, Baltimore, MD; July 9 -12, 2017 (12) Van. Raden
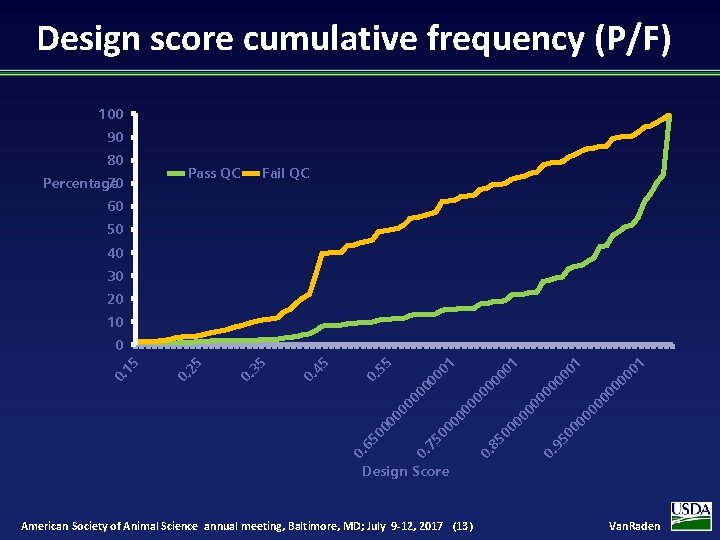
Design score cumulative frequency (P/F) 100 90 80 70 Percentage Pass QC Fail QC 60 50 40 30 20 10 00 00 0. 95 00 00 85 0. 1 00 00 75 0. 1 1 00 00 00 65 0. 00 00 1 55 0. 45 0. 35 0. 25 0. 15 0 Design Score American Society of Animal Science annual meeting, Baltimore, MD; July 9 -12, 2017 (13) Van. Raden

Heritability tests for sequence and array l 1) Estimate h 2 of imputed sequence SNPs selected w Included 3, 000 random bulls and pedigree A matrix w Mean h 2 = failed l 98. 5% for SNPs that passed, 96% for 2) Estimate h 2 of array genotypes from the ZL 5 chip w Included 5, 000 random animals genotyped with ZL 5 w Mean h 2 = 95. 1% for new SNPs, 95. 8% for previous American Society of Animal Science annual meeting, Baltimore, MD; July 9 -12, 2017 (14) Van. Raden

Reverse test: 50 K to sequence l 56, 815 SNPs from Bovine 50 K version 1 w 15, 772 SNPs previously declared not usable − 87% also not identified in 1000 Bulls sequence w 43, 053 currently used SNPs from 50 K − 9% were not identified in 1000 Bulls sequence l Missing SNPs were not associated with MAF or reference / alternate allele pattern American Society of Animal Science annual meeting, Baltimore, MD; July 9 -12, 2017 (15) Van. Raden

Strategies to design chips and use SNPs l Discovery of true QTLs from sequence does not guarantee quality genotypes from chips. l If two SNPs are highly correlated with similar effect sizes, choose the SNP with best design score (and heritability). l Design scores can be obtained online by uploading groups of SNPs plus flanking sequence American Society of Animal Science annual meeting, Baltimore, MD; July 9 -12, 2017 (16) Van. Raden

Summary l About 35% of sequence SNPs do not convert to chips w Design scores were very helpful to predict success, whereas SNP heritability & repetitive DNA location somewhat helpful. Gene location, allele pattern, and MAF were not helpful. l About 9% of usable chip SNPs not in sequence data l Arrays are excellent for tracking marker SNPs, but some true QTLs may require targeted sequencing American Society of Animal Science annual meeting, Baltimore, MD; July 9 -12, 2017 (17) Van. Raden

Acknowledgements l 1000 Bull Genomes Project for sequence genotypes l Council on Dairy Cattle Breeding for array genotypes USDA-ARS project 1265 -31000 -101 -00 , “Improving Genetic Predictions in Dairy Animals Using Phenotypic and Genomic Information. ” American Society of Animal Science annual meeting, Baltimore, MD; July 9 -12, 2017 (18) Van. Raden
- Slides: 18