AAAI 05 Tutorial on Bioinformatics Machine Learning Jinyan

AAAI 05 Tutorial on Bioinformatics & Machine Learning Jinyan Li & Limsoon Wong Institute for Infocomm Research 21 Heng Mui Keng Terrace Singapore Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong
Tutorial Plan 1. Intro to Biology & Bioinformatics 2. TIS Prediction feature generation & feature selection 3. Tx Optimization for Childhood ALL emerging patterns & PCL 6. Subgraphs in Protein Interactions closed patterns 7. Mining Errors from Bio Databases association rules 4. Analysis of Proteomics Data 5. Prognosis based on Gene Expression decision tree ensembles & CS 4 extreme sample selection & ERCOF Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong

Coverage • Data: – DNA sequences, gene expression profiles, proteomics mass-spectra, protein sequences, protein structures, & biomedical text/annotations • Feature selection: – 2, t-statistics, signal-to-noise, entropy, CFS • Machine learning and data mining: – Decision trees, SVMs, emerging patterns, closed patterns, graph theories Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong
Body and Cell • Our body consists of a number of organs • Each organ composes of a number of tissues • Each tissue composes of cells of the same type • Cells perform two types of function – Chemical reactions needed to maintain our life – Pass info for maintaining life to next generation • In particular – Protein performs chemical reactions – DNA stores & passes info – RNA is intermediate between DNA & proteins Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong

DNA • Stores instructions needed by the cell to perform daily life function • Consists of two strands interwoven together and form a double helix • Each strand is a chain of some small molecules called nucleotides Francis Crick shows James Watson the model of DNA in their room number 103 of the Austin Wing at the Cavendish Laboratories, Cambridge Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong
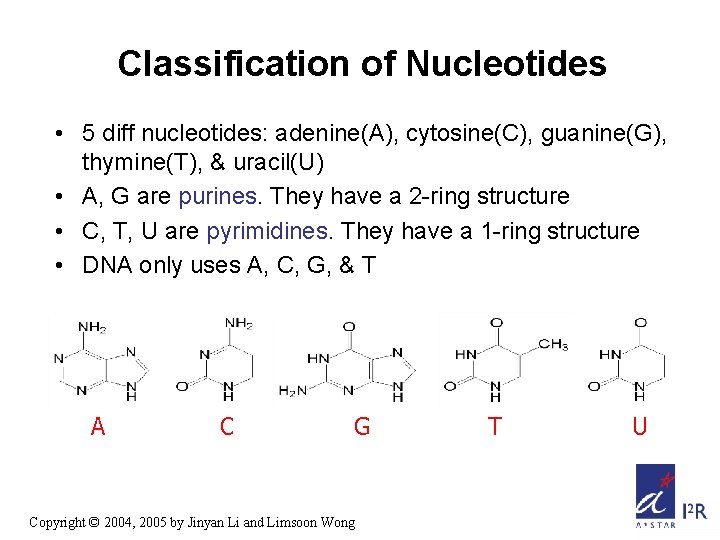
Classification of Nucleotides • 5 diff nucleotides: adenine(A), cytosine(C), guanine(G), thymine(T), & uracil(U) • A, G are purines. They have a 2 -ring structure • C, T, U are pyrimidines. They have a 1 -ring structure • DNA only uses A, C, G, & T A C G Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong T U
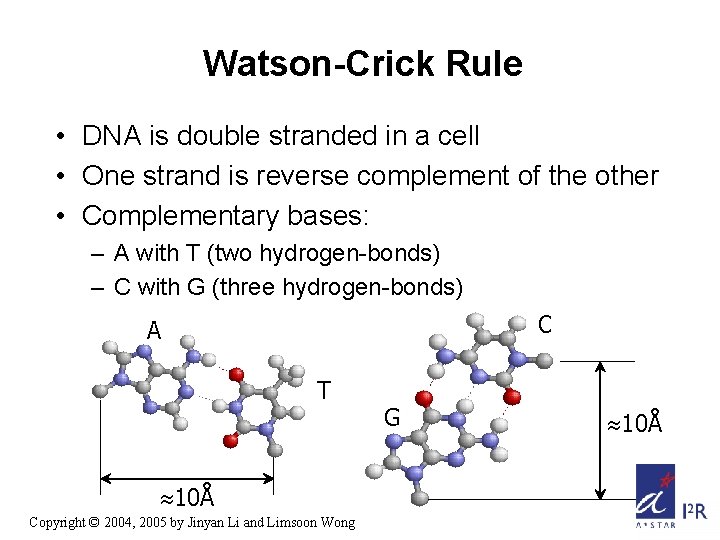
Watson-Crick Rule • DNA is double stranded in a cell • One strand is reverse complement of the other • Complementary bases: – A with T (two hydrogen-bonds) – C with G (three hydrogen-bonds) C A T 10Å Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong G 10Å

Chromosome • DNA is usually tightly wound around histone proteins and forms a chromosome • The total info stored in all chromosomes constitutes a genome • In most multi-cell organisms, every cell contains the same complete set of chromosomes – May have some small diff due to mutation • Human genome has 3 G bases, organized in 23 pairs of chromosomes Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong

Gene • A gene is a sequence of DNA that encodes a protein or an RNA molecule • About 30, 000 – 35, 000 (protein-coding) genes in human genome • For gene that encodes protein – In Prokaryotic genome, one gene corresponds to one protein – In Eukaryotic genome, one gene may correspond to more than one protein because of the process “alternative splicing” Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong
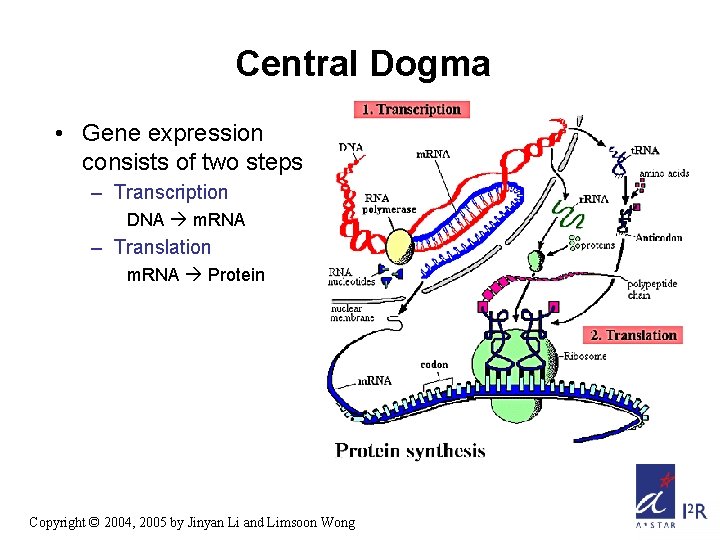
Central Dogma • Gene expression consists of two steps – Transcription DNA m. RNA – Translation m. RNA Protein Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong

Genetic Code • Start codon: ATG (code for M) • Stop codon: TAA, TAG, TGA Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong
Protein • A sequence composed from an alphabet of 20 amino acids – Length is usually 20 to 5000 amino acids – Average around 350 amino acids • Folds into 3 D shape, forming the building block & performing most of the chemical reactions within a cell Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong

Classification of Amino Acids • Amino acids can be classified into 4 types • Positively charged (basic) – Arginine (Arg, R) – Histidine (His, H) – Lysine (Lys, K) • Negatively charged (acidic) – Aspartic acid (Asp, D) – Glutamic acid (Glu, E) Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong

Classification of Amino Acids • Polar (overall uncharged, • Nonpolar (overall but uneven charge uncharged and uniform distribution. can form charge distribution. cant hydrogen bonds with form hydrogen bonds water. they are called with water. they are hydrophilic) called hydrophobic) – – – – Asparagine (Asn, N) Cysteine (Cys, C) Glutamine (Gln, Q) Glycine (Gly, G) Serine (Ser, S) Threonine (Thr, T) Tyrosine (Tyr, Y) Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong – – – – Alanine (Ala, A) Isoleucine (Ile, I) Leucine (Leu, L) Methionine (Met, M) Phenylalanine (Phe, F) Proline (Pro, P) Tryptophan (Trp, W) Valine (Val, V)
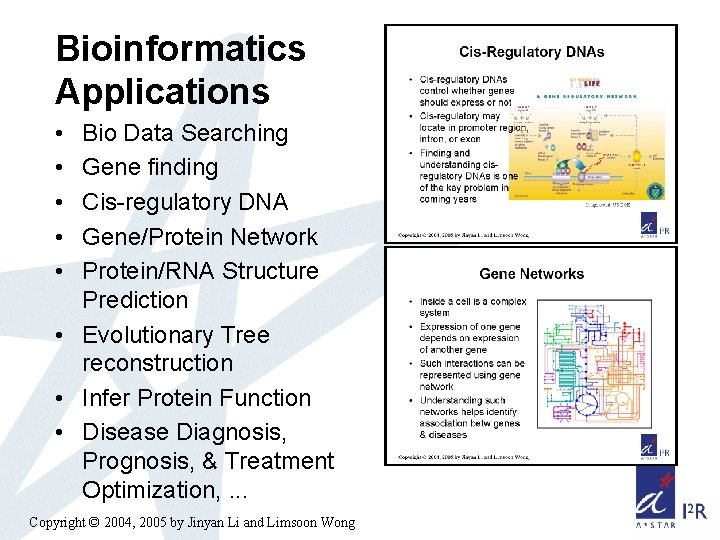
Bioinformatics Applications • • • Bio Data Searching Gene finding Cis-regulatory DNA Gene/Protein Network Protein/RNA Structure Prediction • Evolutionary Tree reconstruction • Infer Protein Function • Disease Diagnosis, Prognosis, & Treatment Optimization, . . . Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong
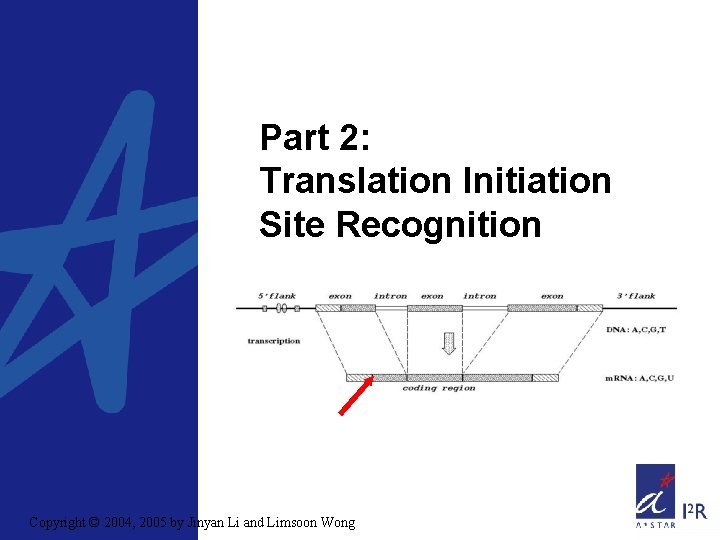
Part 2: Translation Initiation Site Recognition Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong

A Sample c. DNA 299 HSU 27655. 1 CAT U 27655 Homo sapiens CGTGTGTGCAGCAGCCTGCAGCTGCCCCAAGCCATGGCTGAACACTGACTCCCAGCTGTG CCCAGGGCTTCAAAGACTTCTCAGCTTCGAGCATGGCTTTTGGCTGTCAGGGCAGCTGTA GGAGGCAGATGAGAAGAGGGAGATGGCCTTGGAGGAAGGGGCCTGGTGCCGAGGA CCTCTCCTGGCCAGGAGCTTCCTCCAGGACAAGACCTTCCACCCAACAAGGACTCCCCT. . . . . . i. EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE • What makes the second ATG the TIS? Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong 80 160 240

Approach • Training data gathering • Signal generation – k-grams, distance, domain know-how, . . . • Signal selection – Entropy, 2, CFS, t-test, domain know-how. . . • Signal integration – SVM, ANN, PCL, CART, C 4. 5, k. NN, . . . Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong

Training & Testing Data • • • Vertebrate dataset of Pedersen & Nielsen [ISMB’ 97] 3312 sequences 13503 ATG sites 3312 (24. 5%) are TIS 10191 (75. 5%) are non-TIS Use for 3 -fold x-validation expts Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong

Signal Generation • K-grams (ie. , k consecutive letters) – – K = 1, 2, 3, 4, 5, … Window size vs. fixed position Up-stream, downstream vs. any where in window In-frame vs. any frame Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong

Too Many Signals • For each k, there are 4 k * 3 * 2 k-grams • If we use k = 1, 2, 3, 4, 5, we have 24 + 96 + 384 + 1536 + 6144 = 8184 features! • This is too many for most machine learning algorithms Þ Need to do signal selection – t-stats, 2, CFS, signal-to-noise, entropy, gini index, info gain ratio, . . . Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong
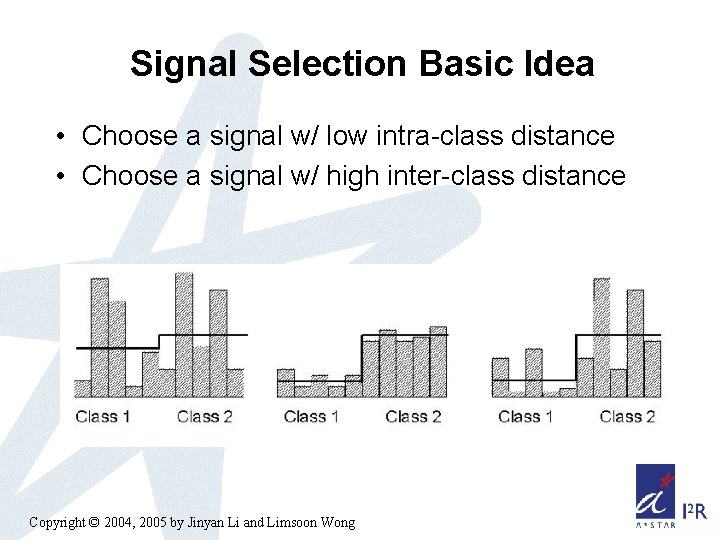
Signal Selection Basic Idea • Choose a signal w/ low intra-class distance • Choose a signal w/ high inter-class distance Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong

Signal Selection by T-Statistics Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong

Signal Selection by 2 Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong

Sample K-Grams Selected by CFS Leaky scanning Kozak consensus • Position – 3 • in-frame upstream ATG • in-frame downstream Stop codon – TAA, TAG, TGA, – CTG, GAC, GAG, and GCC Codon bias? Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong

Signal Integration • k. NN – Given a test sample, find the k training samples that are most similar to it. Let the majority class win • SVM – Given a group of training samples from two classes, determine a separating plane that maximises the margin of error • Naïve Bayes, ANN, C 4. 5, CS 4, . . . Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong
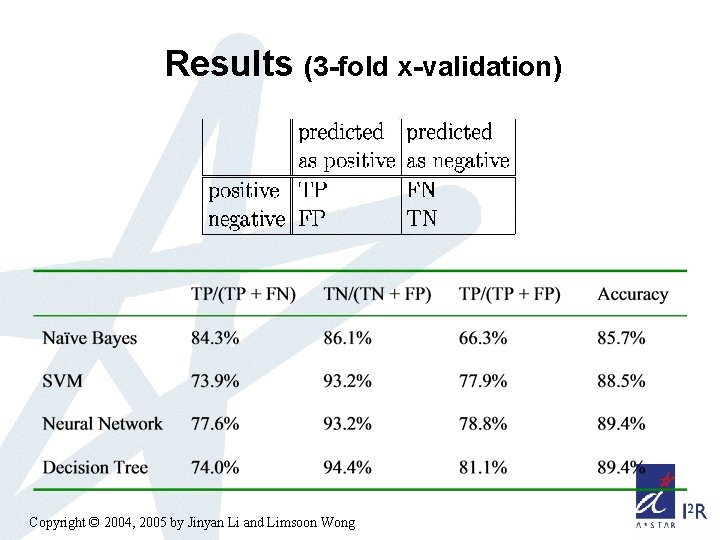
Results (3 -fold x-validation) Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong
![Technique Comparisons • Pedersen&Nielsen [ISMB’ 97] – Neural network – No explicit features • Technique Comparisons • Pedersen&Nielsen [ISMB’ 97] – Neural network – No explicit features •](http://slidetodoc.com/presentation_image_h2/df8891edbba6a76eca8be3d8fa3f918b/image-28.jpg)
Technique Comparisons • Pedersen&Nielsen [ISMB’ 97] – Neural network – No explicit features • Zien [Bioinformatics’ 00] – SVM+kernel engineering – No explicit features • Hatzigeorgiou [Bioinformatics’ 02] – Multiple neural networks – Scanning rule – No explicit features Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong • This approach – Explicit feature generation – Explicit feature selection – Use any machine learning method w/o any form of complicated tuning – Scanning rule is optional
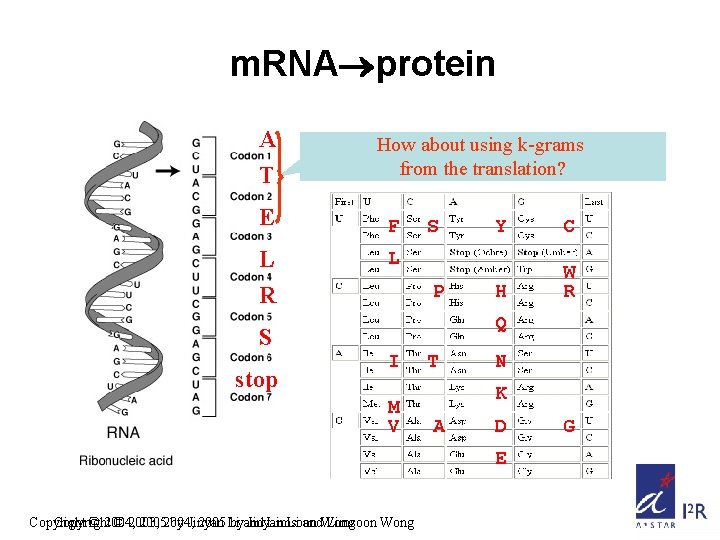
m. RNA protein A T E L R How about using k-grams from the translation? F S C H W R L P Q S stop Y I M V T N K A D E Copyright © 2004, © 2003, 20052004, by Jinyan 2005 Li byand Jinyan Limsoon Li and. Wong Limsoon Wong G
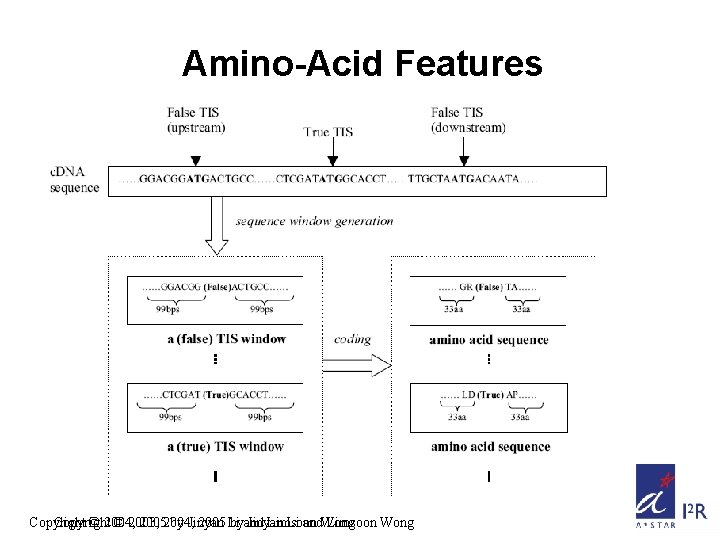
Amino-Acid Features Copyright © 2004, © 2003, 20052004, by Jinyan 2005 Li byand Jinyan Limsoon Li and. Wong Limsoon Wong
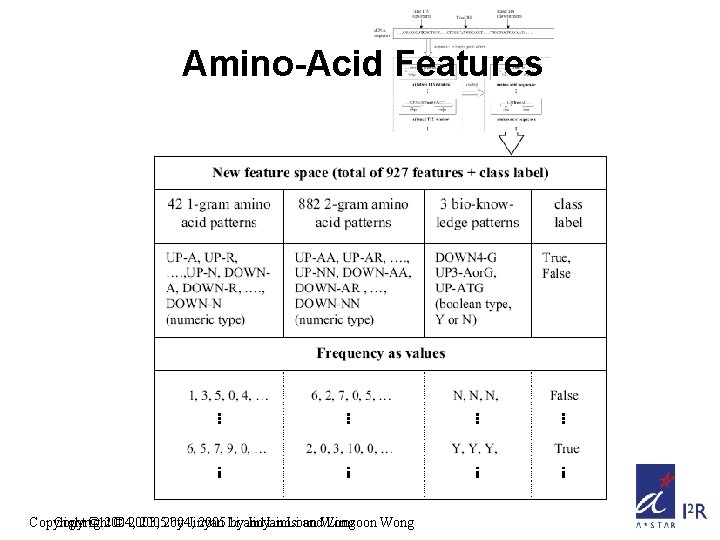
Amino-Acid Features Copyright © 2004, © 2003, 20052004, by Jinyan 2005 Li byand Jinyan Limsoon Li and. Wong Limsoon Wong
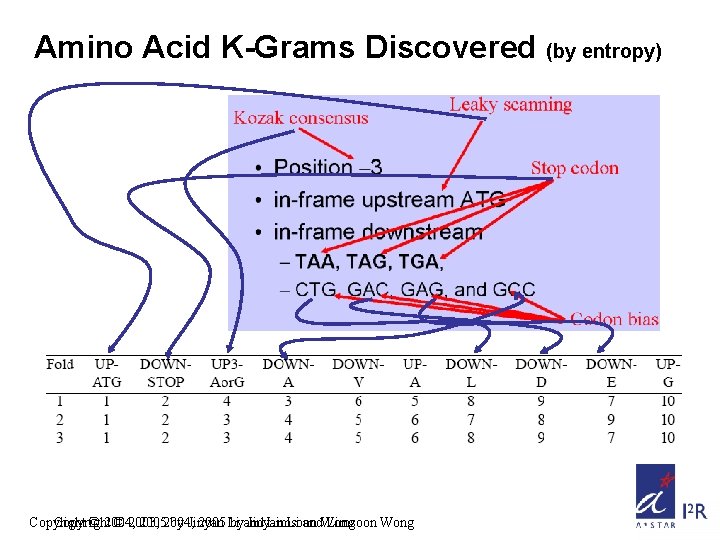
Amino Acid K-Grams Discovered (by entropy) Copyright © 2004, © 2003, 20052004, by Jinyan 2005 Li byand Jinyan Limsoon Li and. Wong Limsoon Wong
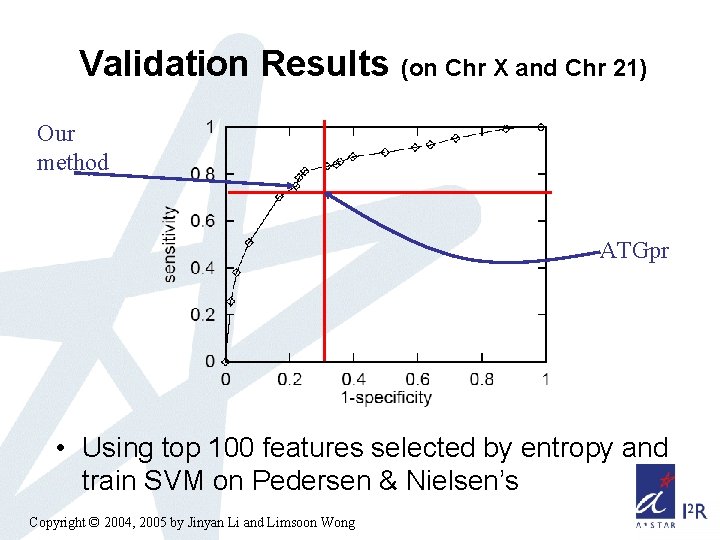
Validation Results (on Chr X and Chr 21) Our method ATGpr • Using top 100 features selected by entropy and train SVM on Pedersen & Nielsen’s Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong

Part 3: Treatment Optimization of Childhood Leukemia Image credit: FEER Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong
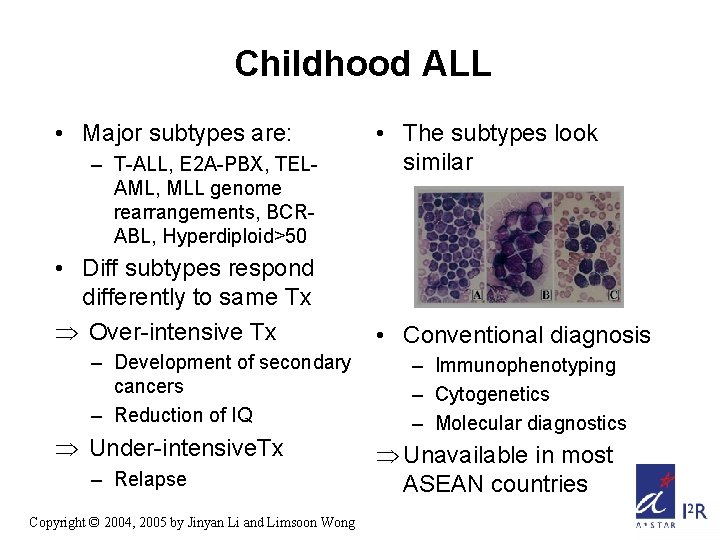
Childhood ALL • Major subtypes are: – T-ALL, E 2 A-PBX, TELAML, MLL genome rearrangements, BCRABL, Hyperdiploid>50 • Diff subtypes respond differently to same Tx Þ Over-intensive Tx – Development of secondary cancers – Reduction of IQ Þ Under-intensive. Tx – Relapse Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong • The subtypes look similar • Conventional diagnosis – Immunophenotyping – Cytogenetics – Molecular diagnostics Þ Unavailable in most ASEAN countries
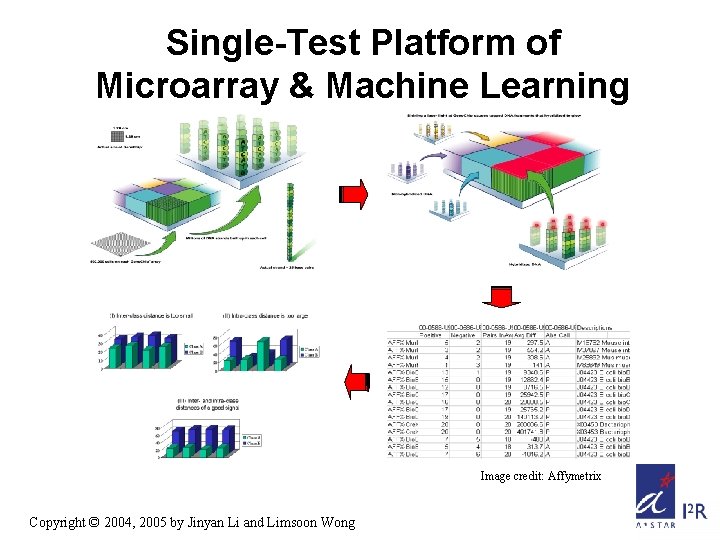
Single-Test Platform of Microarray & Machine Learning Image credit: Affymetrix Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong
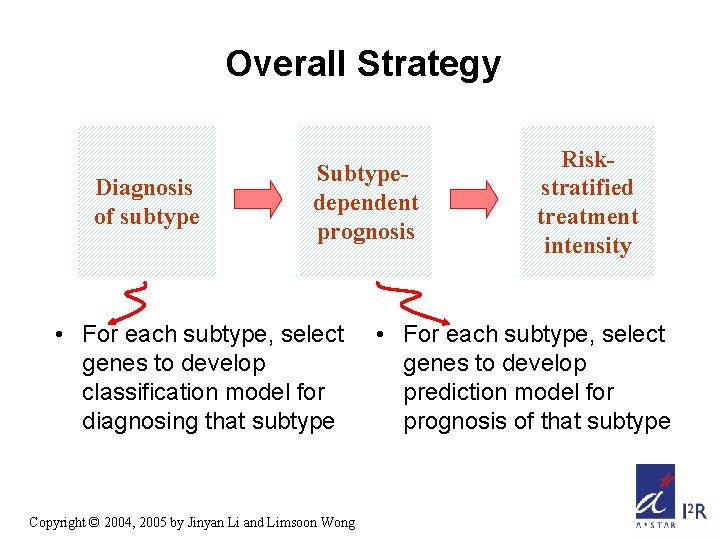
Overall Strategy Diagnosis of subtype Subtypedependent prognosis • For each subtype, select genes to develop classification model for diagnosing that subtype Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong Riskstratified treatment intensity • For each subtype, select genes to develop prediction model for prognosis of that subtype

Subtype Diagnosis by PCL • • Gene expression data collection Gene selection by 2 PCL Classifier training by emerging pattern Classifier tuning (optional for some machine learning methods) • Apply PCL for diagnosis of future cases Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong
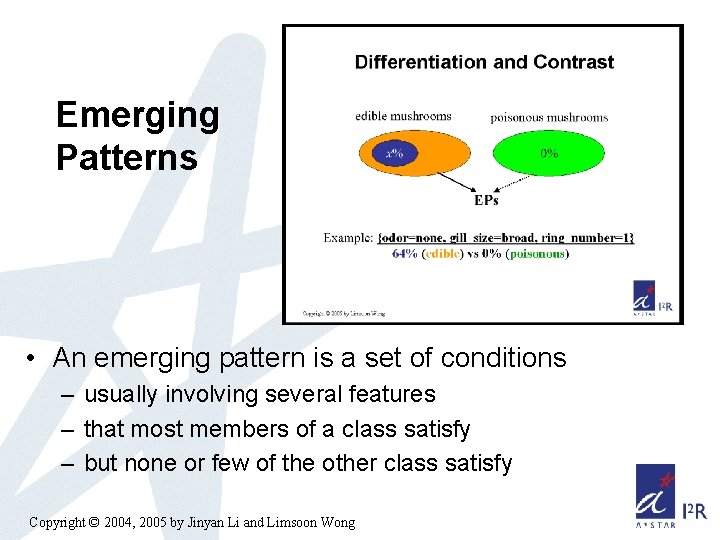
Emerging Patterns • An emerging pattern is a set of conditions – usually involving several features – that most members of a class satisfy – but none or few of the other class satisfy Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong

PCL: Prediction by Collective Likelihood Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong
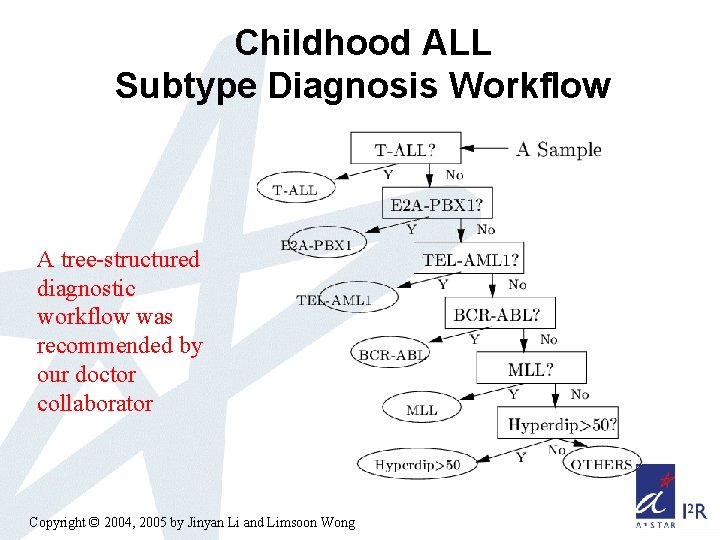
Childhood ALL Subtype Diagnosis Workflow A tree-structured diagnostic workflow was recommended by our doctor collaborator Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong
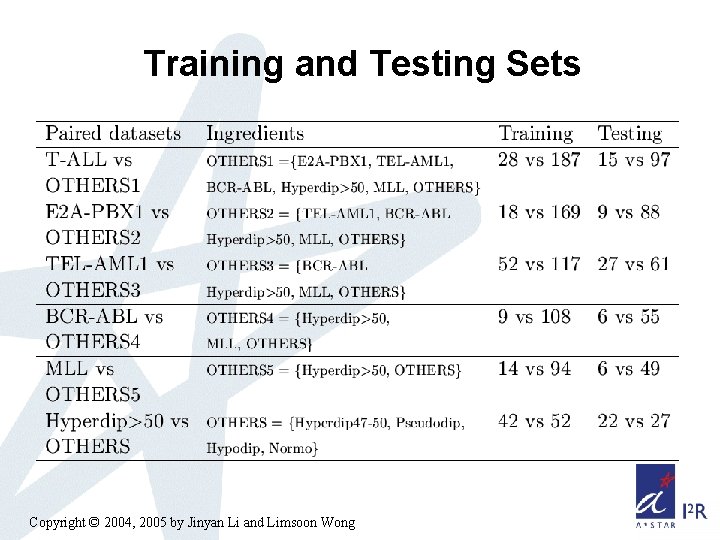
Training and Testing Sets Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong
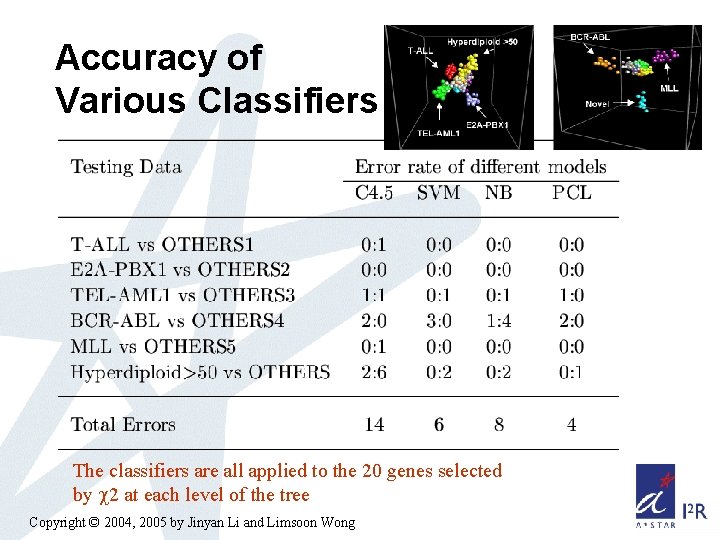
Accuracy of Various Classifiers The classifiers are all applied to the 20 genes selected by 2 at each level of the tree Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong

Understandability of EP & PCL • E. g. , for T-ALL vs. OTHERS, one ideally discriminatory gene 38319_at was found, inducing these 2 EPs • These give us the diagnostic rule Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong
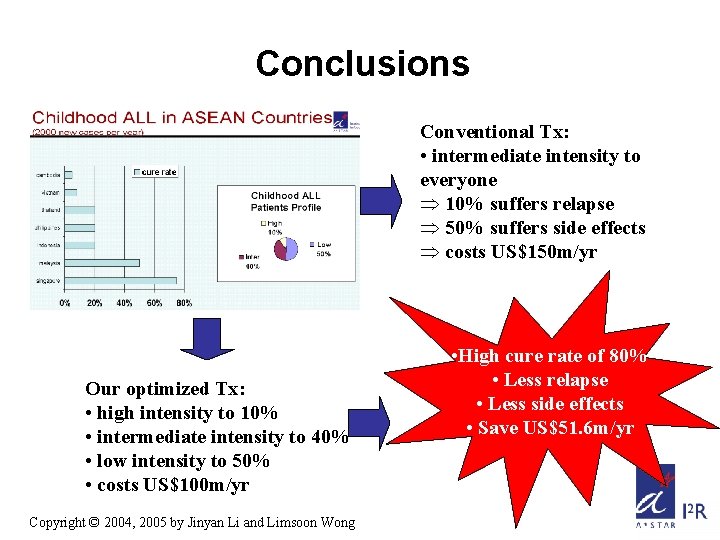
Conclusions Conventional Tx: • intermediate intensity to everyone Þ 10% suffers relapse Þ 50% suffers side effects Þ costs US$150 m/yr Our optimized Tx: • high intensity to 10% • intermediate intensity to 40% • low intensity to 50% • costs US$100 m/yr Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong • High cure rate of 80% • Less relapse • Less side effects • Save US$51. 6 m/yr

Part 4: Topology of Protein Interaction Networks: Hubs, Cores, Bipartites Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong
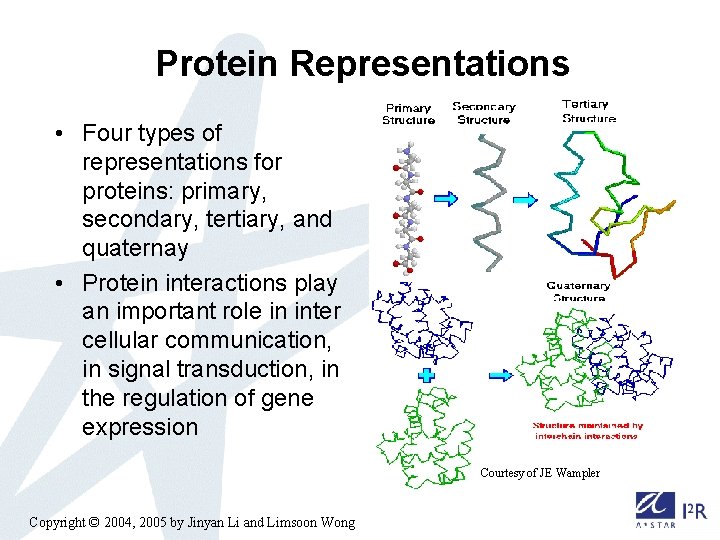
Protein Representations • Four types of representations for proteins: primary, secondary, tertiary, and quaternay • Protein interactions play an important role in inter cellular communication, in signal transduction, in the regulation of gene expression Courtesy of JE Wampler Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong
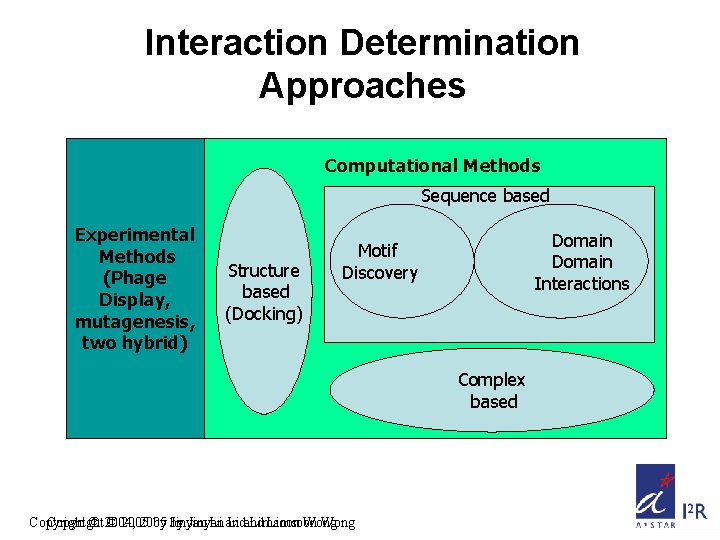
Interaction Determination Approaches Computational Methods Sequence based Experimental Methods (Phage Display, mutagenesis, two hybrid) Structure based (Docking) Domain Interactions Motif Discovery Complex based Copyright © 2004, © 2005 by by Jinyan Li and Limsoon Wong

Protein Interaction Databases • PDB---Protein Databank (http: //www. rcsb. org/pdb/; reference article: H. M. Berman, etal: The Protein Data Bank. NAR, 28 pp. 235 -242 (2000) ) structure info; complex info. • BIND---The Biomolecular Interaction Network Database (http: //binddb. org; Bader et al. NAR, 29. 2001) • The GRID: The General Repository for Interaction Datasets http: //biodata. mshri. on. ca/grid/servlet/Index • DIP---Database of Interacting Proteins (http: //dip. doe-mbi. ucla. edu/). Xml format. Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong
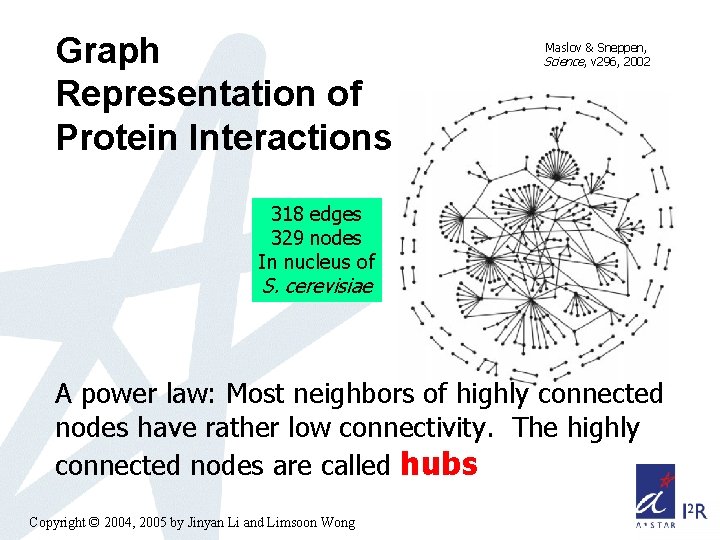
Graph Representation of Protein Interactions Maslov & Sneppen, Science, v 296, 2002 318 edges 329 nodes In nucleus of S. cerevisiae A power law: Most neighbors of highly connected nodes have rather low connectivity. The highly connected nodes are called hubs Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong
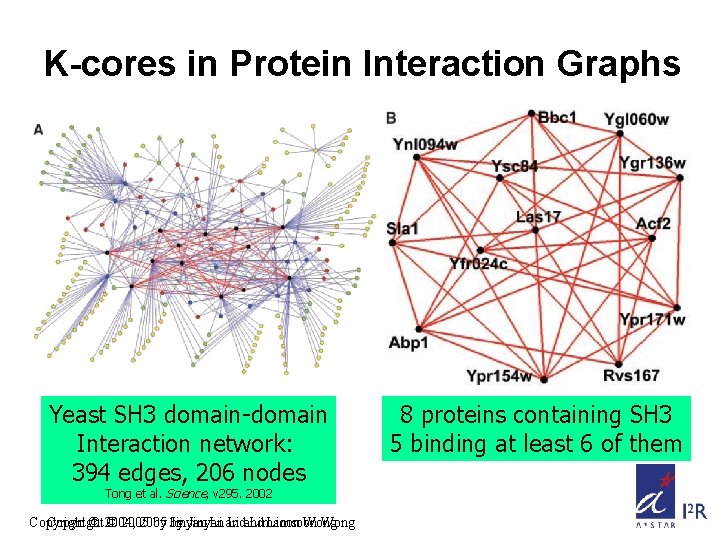
K-cores in Protein Interaction Graphs Yeast SH 3 domain-domain Interaction network: 394 edges, 206 nodes Tong et al. Science, v 295. 2002 Copyright © 2004, © 2005 by by Jinyan Li and Limsoon Wong 8 proteins containing SH 3 5 binding at least 6 of them
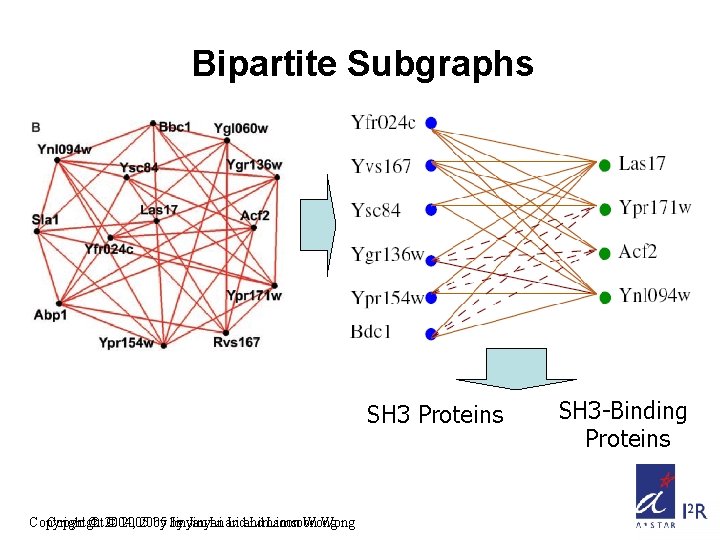
Bipartite Subgraphs SH 3 Proteins Copyright © 2004, © 2005 by by Jinyan Li and Limsoon Wong SH 3 -Binding Proteins

Some Definitions • A graph G= VG, EG , where VG is a set of vertices and EG is a set of edges. No self loops, no parallel edges, undirected • A graph H= VH, EH is a subgraph of G, if VH is contained in VG and EH is contained in EG • H = V 1 V 2, EH is bipartite subgraph of G if – H is a subgraph of G, and – V 1 V 2 = {}, and – every edge in EH joins a vertex in V 1 and a vertex in V 2 Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong
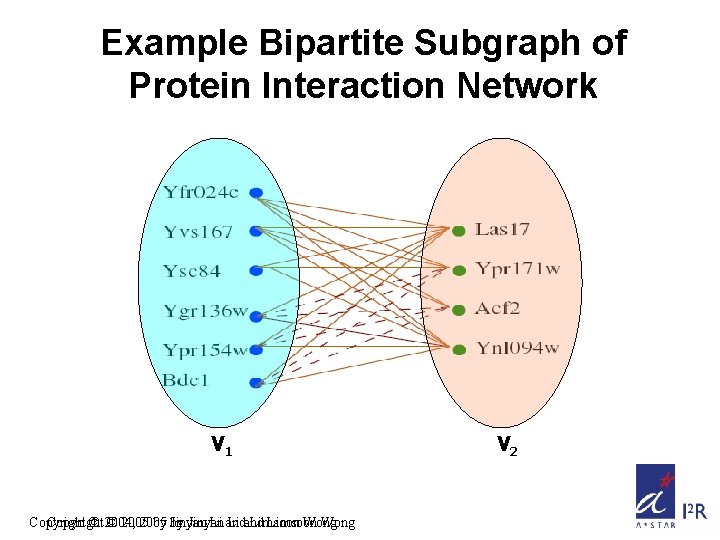
Example Bipartite Subgraph of Protein Interaction Network V 1 Copyright © 2004, © 2005 by by Jinyan Li and Limsoon Wong V 2

Maximal Complete Bipartite Subgraphs • A complete bipartite subgraph H = V 1 V 2, EH of G is said to be maximal if there is no other complete bipartite subgraph H’ = V’ 1 V’ 2, EH’ of G such that V 1 V’ 1 and V 2 V’ 2 Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong

Study of Protein Interaction Network Topology • List all maximal complete bipartite subgraphs from a protein interaction network (Step 1) • Discover motifs from these protein groups (Step 2) • Form binding motif pairs (Step 3) The problem previously studied by Epptein (1994, Info Proc Lett); Makino & Uno (2004, SWAT); Zaki & Ogihara (1998). Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong

Efficient Mining of Maximal Complete Bipartite Subgraphs • Equiv to mining of closed patterns from adjacency matrix of G • Why: – the number of closed patterns is even, – is precisely double of the number of the maximal complete bipartite subgraphs, and – there is one-to-one correspondence betw a maximal complete bipartite subgraph & a closed pattern pair • Many state-of-the-art algorithms for mining closed patterns Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong
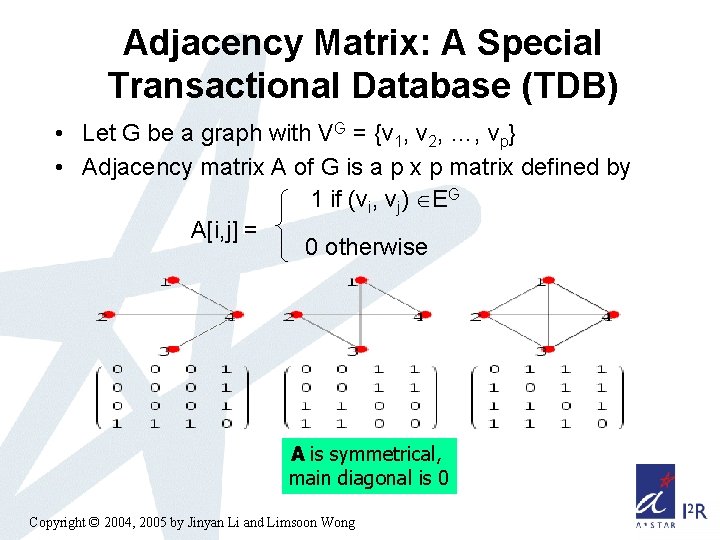
Adjacency Matrix: A Special Transactional Database (TDB) • Let G be a graph with VG = {v 1, v 2, …, vp} • Adjacency matrix A of G is a p x p matrix defined by 1 if (vi, vj) EG A[i, j] = 0 otherwise A is symmetrical, main diagonal is 0 Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong
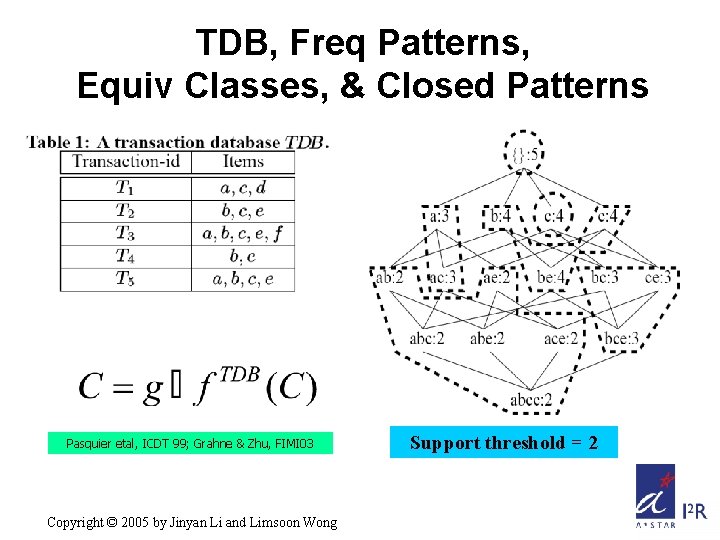
TDB, Freq Patterns, Equiv Classes, & Closed Patterns Pasquier etal, ICDT 99; Grahne & Zhu, FIMI 03 Copyright © 2005 by Jinyan Li and Limsoon Wong Support threshold = 2
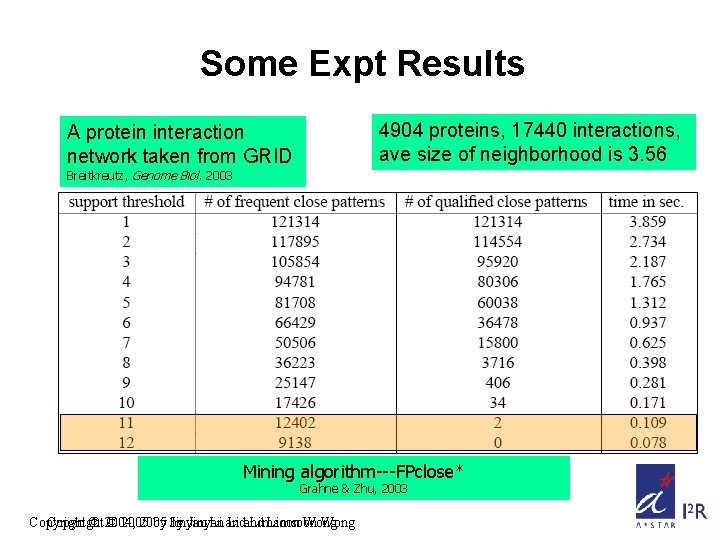
Some Expt Results 4904 proteins, 17440 interactions, ave size of neighborhood is 3. 56 A protein interaction network taken from GRID Breitkreutz, Genome Biol. 2003 Mining algorithm---FPclose* Grahne & Zhu, 2003 Copyright © 2004, © 2005 by by Jinyan Li and Limsoon Wong

Fixed Point Idea • The idea: f(x)=x • Instantiation in life science, f(DNA)=DNA, f(protein. Type) = protein. Type (Meng et al. Bioinformatics, 2004) • The variable x can be a pair of AA segment patterns, then after the transformations by f, the stable pattern is binding motif pair. (Li and Li, Bioinformatics, 2005) Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong

Part 5: Treatment Prognosis for DLBC Lymphoma Image credit: Rosenwald et al, 2002 Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong

Diffuse Large B-Cell Lymphoma • DLBC lymphoma is the most common type of lymphoma in adults • Can be cured by anthracycline-based chemotherapy in 35 to 40 percent of patients Þ DLBC lymphoma comprises several diseases that differ in responsiveness to chemotherapy Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong • Intl Prognostic Index (IPI) – age, “Eastern Cooperative Oncology Group” Performance status, tumor stage, lactate dehydrogenase level, sites of extranodal disease, . . . • Not very good for stratifying DLBC lymphoma patients for therapeutic trials Þ Use gene-expression profiles to predict outcome of chemotherapy?
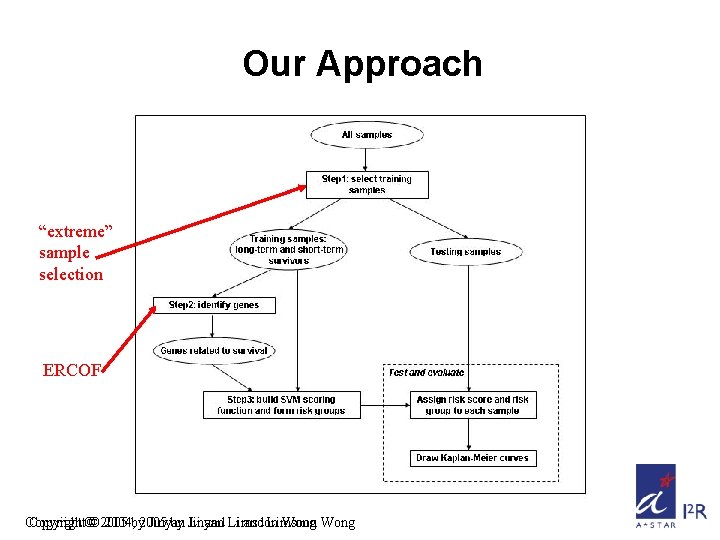
Our Approach “extreme” sample selection ERCOF Copyright©© 2005 2004, by 2005 Jinyan by Jinyan Li and Li Limsoon and Limsoon Wong

Extreme Sample Selection Short-term Survivors v. s. Long-term Survivors Short-term survivors who died within a short period F(T) < c 1 and E(T) = 1 Long-term survivors who were alive after a long follow-up time F(T) > c 2 T: sample F(T): follow-up time E(T): status (1: unfavorable; 0: favorable) c 1 and c 2: thresholds of survival time Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong
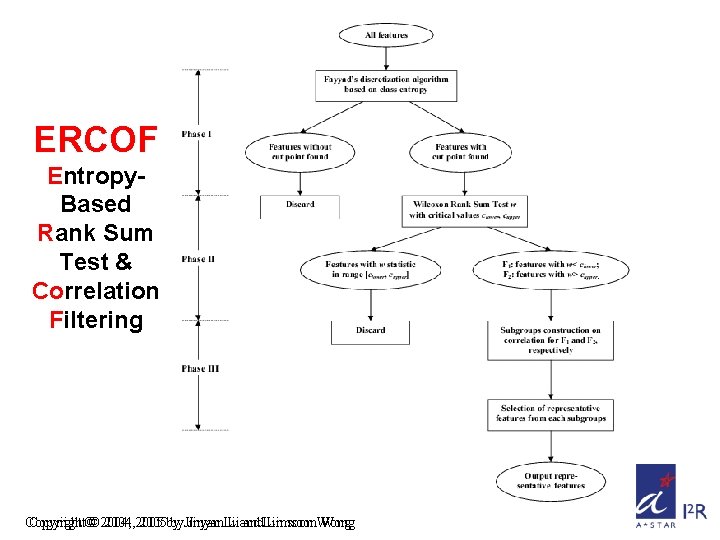
ERCOF Entropy. Based Rank Sum Test & Correlation Filtering Copyright©© 2004, 2005 by by. Jinyan. Li Liand and. Limsoon. Wong
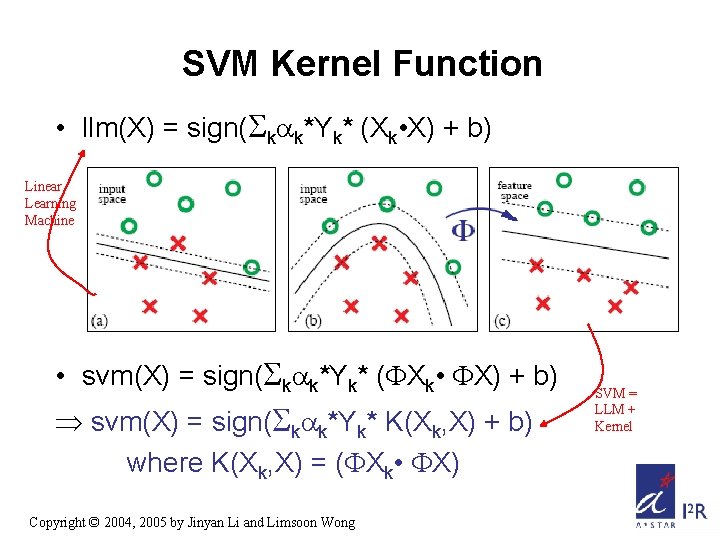
SVM Kernel Function • llm(X) = sign( k k*Yk* (Xk • X) + b) Linear Learning Machine • svm(X) = sign( k k*Yk* ( Xk • X) + b) Þ svm(X) = sign( k k*Yk* K(Xk, X) + b) where K(Xk, X) = ( Xk • X) Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong SVM = LLM + Kernel

Risk Score Construction Linear Kernel SVM regression function T: test sample, x(i): support vector, yi: class label (1: short-term survivors; -1: long-term survivors) Transformation function (posterior probability) S(T): risk score of sample T Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong
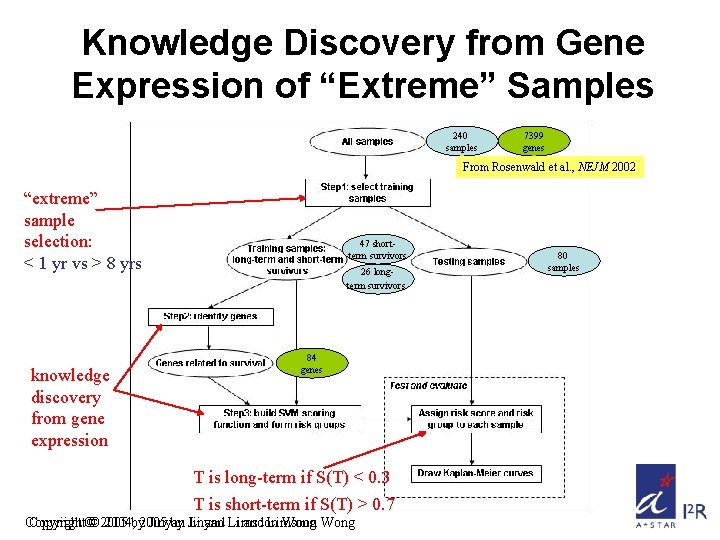
Knowledge Discovery from Gene Expression of “Extreme” Samples 240 samples 7399 genes From Rosenwald et al. , NEJM 2002 “extreme” sample selection: < 1 yr vs > 8 yrs knowledge discovery from gene expression 47 shortterm survivors 26 longterm survivors 84 genes T is long-term if S(T) < 0. 3 T is short-term if S(T) > 0. 7 Copyright©© 2005 2004, by 2005 Jinyan by Jinyan Li and Li Limsoon and Limsoon Wong 80 samples
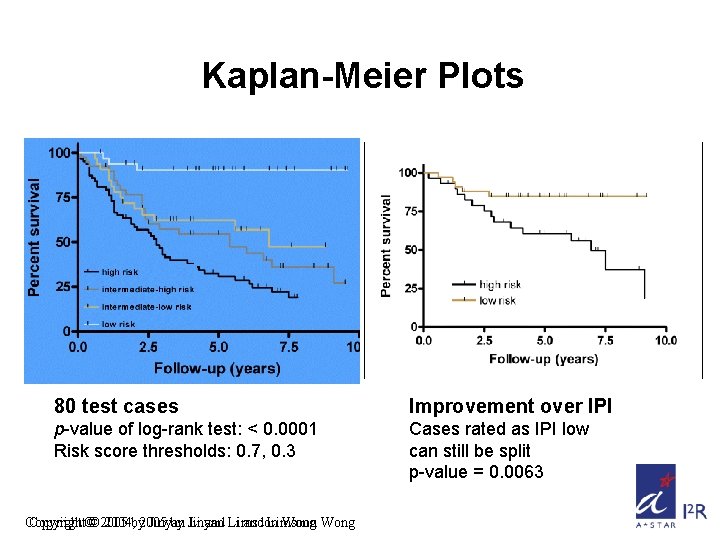
Kaplan-Meier Plots 80 test cases Improvement over IPI p-value of log-rank test: < 0. 0001 Risk score thresholds: 0. 7, 0. 3 Cases rated as IPI low can still be split p-value = 0. 0063 Copyright©© 2005 2004, by 2005 Jinyan by Jinyan Li and Li Limsoon and Limsoon Wong
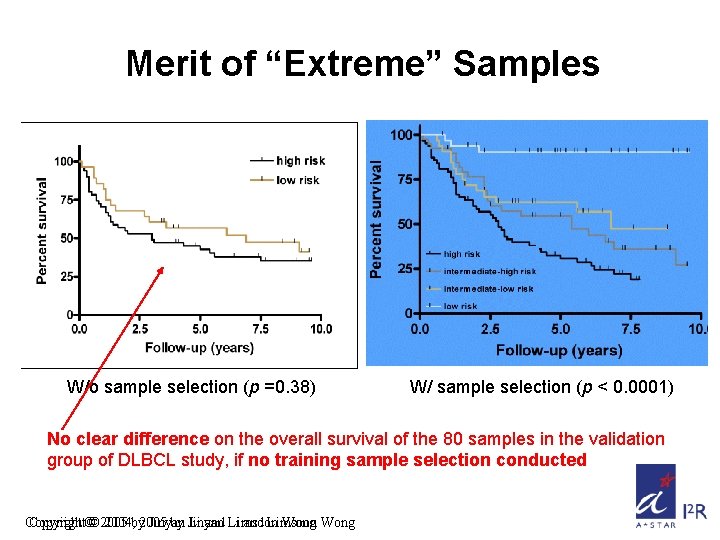
Merit of “Extreme” Samples W/o sample selection (p =0. 38) W/ sample selection (p < 0. 0001) No clear difference on the overall survival of the 80 samples in the validation group of DLBCL study, if no training sample selection conducted Copyright©© 2005 2004, by 2005 Jinyan by Jinyan Li and Li Limsoon and Limsoon Wong
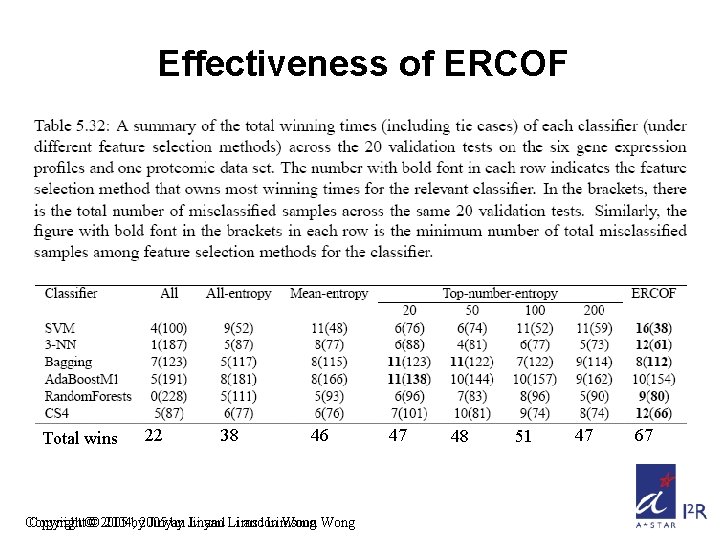
Effectiveness of ERCOF Total wins 22 38 46 Copyright©© 2005 2004, by 2005 Jinyan by Jinyan Li and Li Limsoon and Limsoon Wong 47 48 51 47 67

Conclusions • Selecting extreme cases as training samples is an effective way to improve patient outcome prediction based on gene expression profiles and clinical information • ERCOF is a systematic feature selection method that is very useful – ERCOF is very suitable for SVM, 3 -NN, CS 4, Random Forest, as it gives these learning algos highest no. of wins – ERCOF is suitable for Bagging also, as it gives this classifier the lowest no. of errors Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong

Part 6: Decision Tree Based In -Silico Cancer Diagnosis • Bagging, • Boosting, • Random Forest, • Randomization Trees, & • CS 4 Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong
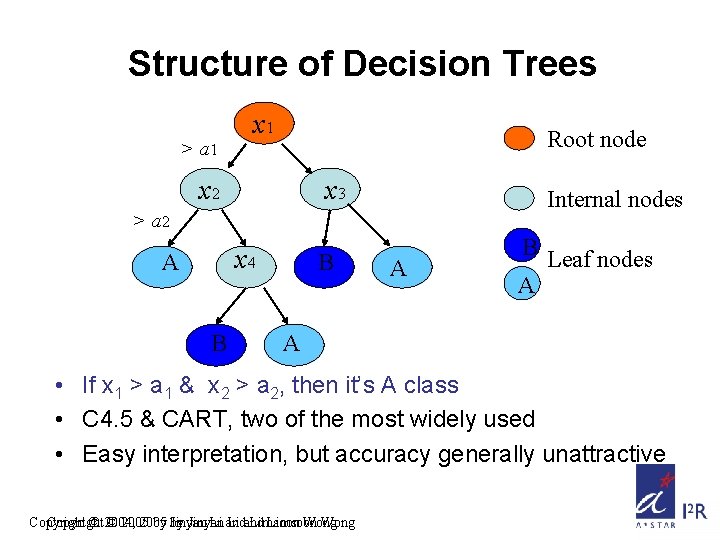
Structure of Decision Trees > a 1 x 1 Root node x 2 x 3 Internal nodes > a 2 x 4 A B B A B Leaf nodes A A • If x 1 > a 1 & x 2 > a 2, then it’s A class • C 4. 5 & CART, two of the most widely used • Easy interpretation, but accuracy generally unattractive Copyright © 2004, © 2005 by by Jinyan Li and Limsoon Wong
Elegance of Decision Trees A B B A A • Diagnosis of central nervous system in hematooncologic patients (Lossos et al, Cancer, 2000) • Prediction of neurobehavioral outcome in head-injury survivors (Temin et al, J. of Neurosurgery, 1995) • Reconstruction of molecular networks from gene expression data (Soinov et al, Genome Biology, 2003) Copyright © 2004, © 2005 by by Jinyan Li and Limsoon Wong

Decision Tree Construction • Steps – Select the “best” feature as the root node of the whole tree – After partition by this feature, select the best feature (wrt the subset of training data) as the root node of this sub-tree – Recursively, until the partitions become pure or almost pure Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong • 3 Measures to Evaluate Which Feature is Best – Gini index – Information gain ratio Single coverage & Fragmentation Problem For Decision trees

Motivating Example • • • h 1, h 2, h 3 are indep classifiers w/ accuracy = 60% C 1, C 2 are the only classes t is a test instance in C 1 h(t) = argmax. C {C 1, C 2} |{hj {h 1, h 2, h 3} | hj(t) = C}| Then prob(h(t) = C 1) = prob(h 1(t)=C 1 & h 2(t)=C 1 & h 3(t)=C 1) + prob(h 1(t)=C 1 & h 2(t)=C 1 & h 3(t)=C 2) + prob(h 1(t)=C 1 & h 2(t)=C 2 & h 3(t)=C 1) + prob(h 1(t)=C 2 & h 2(t)=C 1 & h 3(t)=C 1) = 60% * 60% + 60% * 40% * 60% + 40% * 60% = 64. 8% Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong
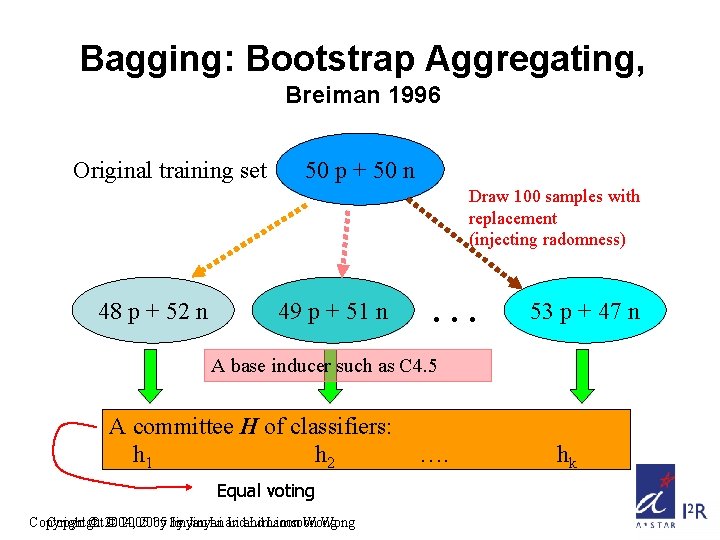
Bagging: Bootstrap Aggregating, Breiman 1996 Original training set 50 p + 50 n Draw 100 samples with replacement (injecting radomness) 48 p + 52 n 49 p + 51 n … 53 p + 47 n A base inducer such as C 4. 5 A committee H of classifiers: h 1 h 2 Equal voting Copyright © 2004, © 2005 by by Jinyan Li and Limsoon Wong …. hk
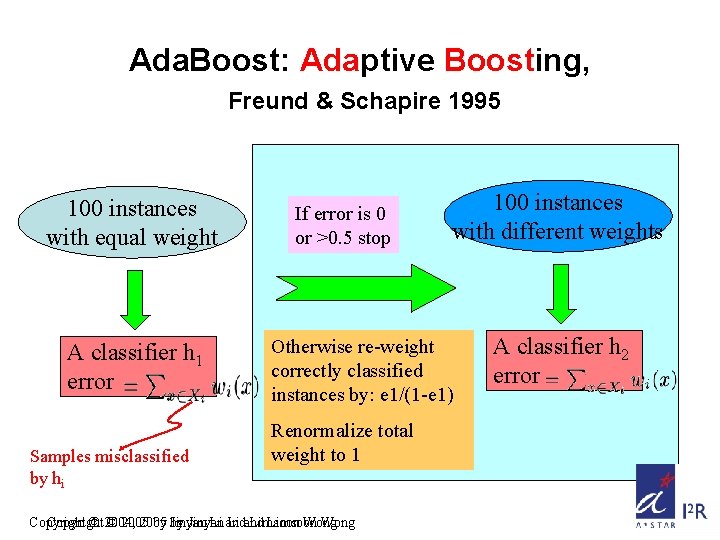
Ada. Boost: Adaptive Boosting, Freund & Schapire 1995 100 instances with equal weight A classifier h 1 error Samples misclassified by hi If error is 0 or >0. 5 stop 100 instances with different weights Otherwise re-weight correctly classified instances by: e 1/(1 -e 1) Renormalize total weight to 1 Copyright © 2004, © 2005 by by Jinyan Li and Limsoon Wong A classifier h 2 error
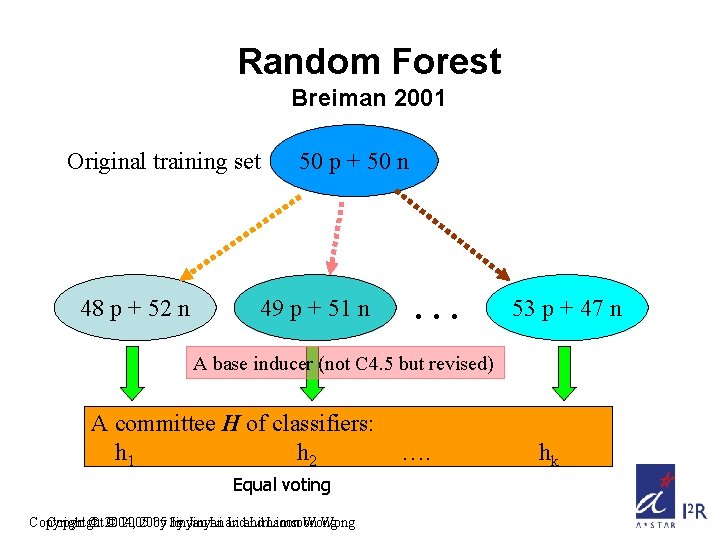
Random Forest Breiman 2001 Original training set 48 p + 52 n 50 p + 50 n 49 p + 51 n … 53 p + 47 n A base inducer (not C 4. 5 but revised) A committee H of classifiers: h 1 h 2 Equal voting Copyright © 2004, © 2005 by by Jinyan Li and Limsoon Wong …. hk
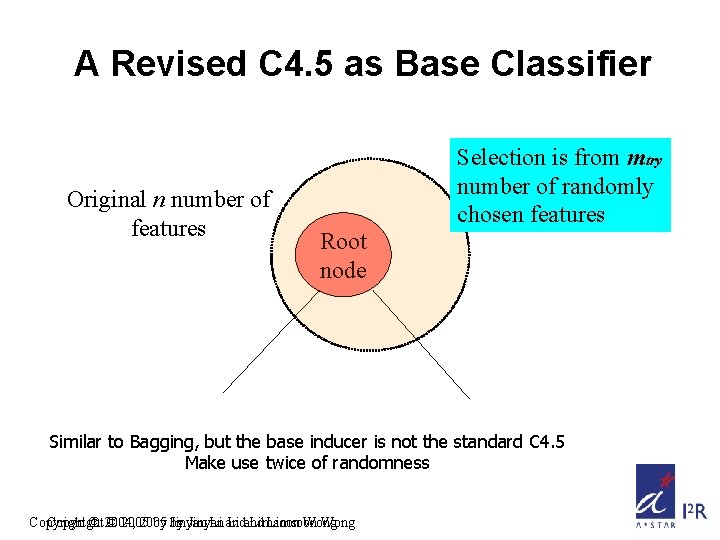
A Revised C 4. 5 as Base Classifier Original n number of features Selection is from mtry number of randomly chosen features Root node Similar to Bagging, but the base inducer is not the standard C 4. 5 Make use twice of randomness Copyright © 2004, © 2005 by by Jinyan Li and Limsoon Wong
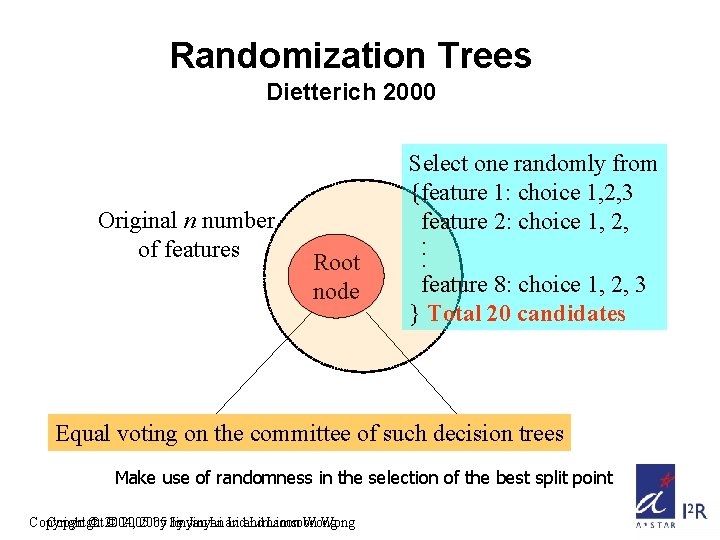
Randomization Trees Dietterich 2000 Original n number of features Root node Select one randomly from {feature 1: choice 1, 2, 3 feature 2: choice 1, 2, . . . feature 8: choice 1, 2, 3 } Total 20 candidates Equal voting on the committee of such decision trees Make use of randomness in the selection of the best split point Copyright © 2004, © 2005 by by Jinyan Li and Limsoon Wong
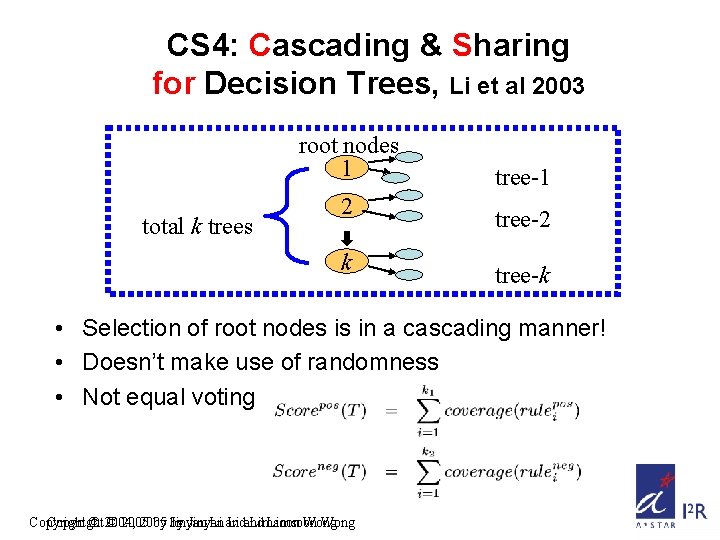
CS 4: Cascading & Sharing for Decision Trees, Li et al 2003 root nodes 1 total k trees 2 k tree-1 tree-2 tree-k • Selection of root nodes is in a cascading manner! • Doesn’t make use of randomness • Not equal voting Copyright © 2004, © 2005 by by Jinyan Li and Limsoon Wong
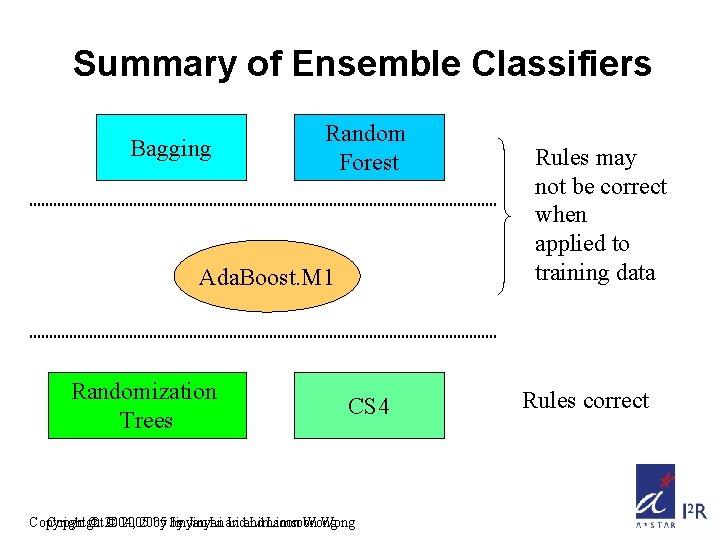
Summary of Ensemble Classifiers Bagging Random Forest Ada. Boost. M 1 Randomization Trees CS 4 Copyright © 2004, © 2005 by by Jinyan Li and Limsoon Wong Rules may not be correct when applied to training data Rules correct

Example of Decision Tree Given a test sample, at most 3 of the 4 genes’ expression values are needed to make a decision! • Yeoh et al. , Cancer Cell 1: 133 -143, 2002; Differentiating MLL subtype from other subtypes of childhood leukemia • Training data (14 MLL vs 201 others), Test data (6 MLL vs 106 others), Number of features: 12558 Copyright © 2004, © 2005 by by Jinyan Li and Limsoon Wong
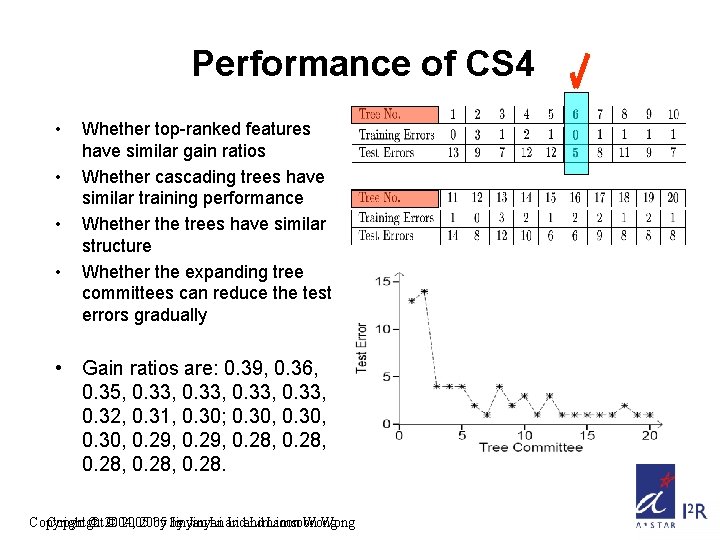
Performance of CS 4 • • Whether top-ranked features have similar gain ratios Whether cascading trees have similar training performance Whether the trees have similar structure Whether the expanding tree committees can reduce the test errors gradually • Gain ratios are: 0. 39, 0. 36, 0. 35, 0. 33, 0. 32, 0. 31, 0. 30; 0. 30, 0. 29, 0. 28, 0. 28. Copyright © 2004, © 2005 by by Jinyan Li and Limsoon Wong
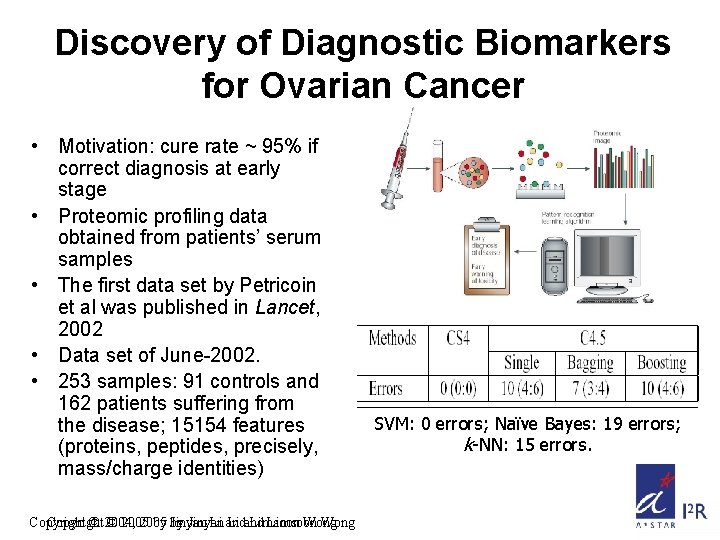
Discovery of Diagnostic Biomarkers for Ovarian Cancer • Motivation: cure rate ~ 95% if correct diagnosis at early stage • Proteomic profiling data obtained from patients’ serum samples • The first data set by Petricoin et al was published in Lancet, 2002 • Data set of June-2002. • 253 samples: 91 controls and 162 patients suffering from the disease; 15154 features (proteins, peptides, precisely, mass/charge identities) Copyright © 2004, © 2005 by by Jinyan Li and Limsoon Wong SVM: 0 errors; Naïve Bayes: 19 errors; k-NN: 15 errors.

Part 7: Mining Errors from Bio Databases Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong

Data Cleansing, Koh et al, DBi. DB 2005 • 11 types & 28 subtypes of data artifacts – Critical artifacts (vector contaminated sequences, duplicates, sequence structure violations) – Non-critical artifacts (misspellings, synonyms) • > 20, 000 seq records in public contain artifacts • Identification of these artifacts are impt for accurate knowledge discovery Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong • Sources of artifacts – Diverse sources of data • • Repeated submissions of seqs to db’s Cross-updating of db’s – Data Annotation • • • Db’s have diff ways for data annotation Data entry errors can be introduced Different interpretations – Lack of standardized nomenclature • • Variations in naming Synonyms, homonyms, & abbrevn – Inadequacy of data quality control mechanisms • Systematic approaches to data cleaning are lacking
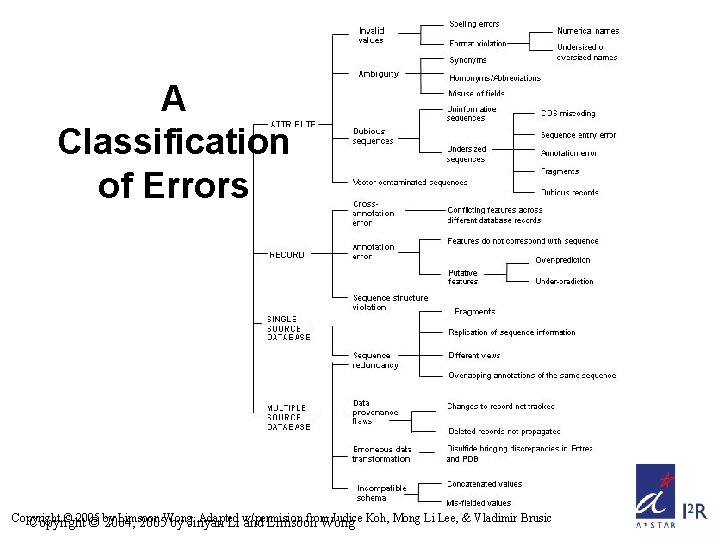
A Classification of Errors Copyright © 2005 Limsoon Adapted from Judice Koh, Mong Li Lee, & Vladimir Brusic Copyright © by 2004, 2005 Wong. by Jinyan Li w/permision and Limsoon Wong

Invalid values Uninformative sequences Undersized sequences ATTRIBUTE Ambiguity Dubious sequences Vector contaminated sequence Crossannotation error RECORD Annotation error Example Meaningless Seqs • Among the 5, 146, 255 protein records queried using Entrez to the major protein or translated nucleotide databases , 3, 327 protein sequences are shorter than four residues (as of Sep, 2004). • In Nov 2004, the total number of undersized protein sequences increases to 3, 350. • Among 43, 026, 887 nucleotide records queried using Entrez to major nucleotide databases, 1, 448 records contain sequences shorter than six bases (as of Sep, 2004). • In Nov 2004, the total number of undersized nucleotide sequences increases to 1, 711. Sequence structure violation SINGLE SOURCE DATABASE Sequence redundancy Data provenance flaws MULTIPLE SOURCE DATABASE Erroneous data transformation Incompatible schema Copyright © 2005 Limsoon Adapted from Judice Koh, Mong Li Lee, & Vladimir Brusic Copyright © by 2004, 2005 Wong. by Jinyan Li w/permision and Limsoon Wong
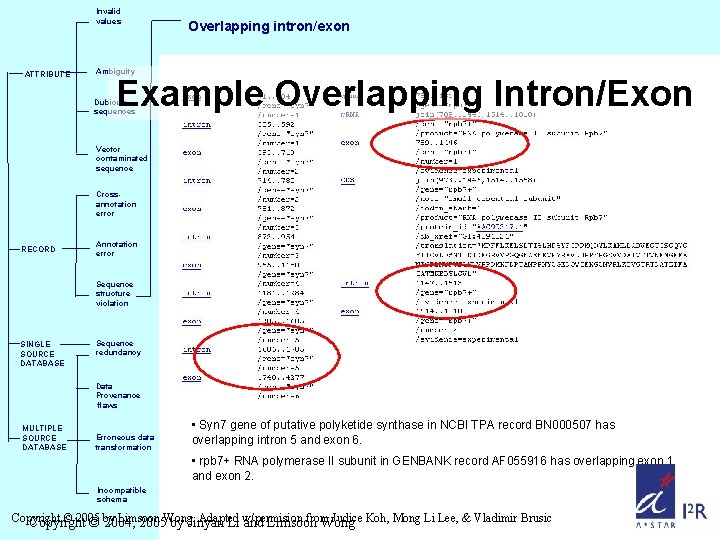
Invalid values ATTRIBUTE Overlapping intron/exon Ambiguity Example Overlapping Intron/Exon Dubious sequences Vector contaminated sequence Crossannotation error RECORD Annotation error Sequence structure violation SINGLE SOURCE DATABASE Sequence redundancy Data Provenance flaws MULTIPLE SOURCE DATABASE Erroneous data transformation • Syn 7 gene of putative polyketide synthase in NCBI TPA record BN 000507 has overlapping intron 5 and exon 6. • rpb 7+ RNA polymerase II subunit in GENBANK record AF 055916 has overlapping exon 1 and exon 2. Incompatible schema Copyright © 2005 Limsoon Adapted from Judice Koh, Mong Li Lee, & Vladimir Brusic Copyright © by 2004, 2005 Wong. by Jinyan Li w/permision and Limsoon Wong

Replication of sequence information Invalid values Different views Overlapping annotations of the same sequence ATTRIBUTE Ambiguity Dubious sequences Example Seqs w/ Identical Info Vector contaminated sequence Submission of the same sequence to different databases • Repeated submission of the same sequence to the same database • Initially submitted by different groups • Protein sequences may be translated from duplicate nucleotide sequences Crossannotation error RECORD Annotation error Sequence structure violation SINGLE SOURCE DATABASE Sequence redundancy Data provenance flaws MULTIPLE SOURCE DATABASE Erroneous data transformation Incompatible schema http: //www. ncbi. nlm. nih. gov/entrez/query. fcgi? cmd=Retrieve&db =protein&list_uids=11692005&dopt=Gen. Pept Copyright © 2005 Limsoon Adapted from Judice Koh, Mong Li Lee, & Vladimir Brusic Copyright © by 2004, 2005 Wong. by Jinyan Li w/permision and Limsoon Wong

Association Rule Mining for De-duplication Select matching criteria Compute similarity scores from known duplicate pairs Generate association rules Detect duplicates using the rules Copyright © 2005 Jinyan Li and Wong. Copyright © by 2004, 2005 by Limsoon Jinyan Li and Limsoon Wong

Features to Match Copyright © 2005 Limsoon Adapted from Judice Koh, Mong Li Lee, & Vladimir Brusic Copyright © by 2004, 2005 Wong. by Jinyan Li w/permision and Limsoon Wong

Association Rule Mining AAG 39642 AAG 39643 AC 0. 9 LE 1. 0 DB 1 SP 1 RF 1. 0 PD 0 FT 1. 0 SQ 1. 0 Similarity scores of known duplicate pairs AAG 39642 Q 9 GNG 8 AC 0. 1 LE 1. 0 DE 0. 4 DB 0 SP 1 RF 1. 0 PD 0 FT 0. 1 SQ 1. 0 P 00599 PSNJ 1 W AC 0. 2 LE 1. 0 DE 0. 4 DB 0 SP 1 RF 1. 0 PD 0 FT 1. 0 SQ 1. 0 P 01486 NTSREB AC 0. 0 LE 1. 0 DE 0. 3 DB 0 SP 1 RF 1. 0 PD 0 FT 1. 0 SQ 1. 0 O 57385 CAA 11159 AC 0. 1 LE 1. 0 DE 0. 5 DB 0 SP 1 RF 0. 0 PD 0 FT 0. 1 SQ 1. 0 S 32792 P 24663 AC 0. 0 LE 1. 0 DE 0. 4 DB 0 SP 1 RF 0. 5 PD 0 FT 1. 0 SQ 1. 0 P 45629 S 53330 AC 0. 0 LE 1. 0 DE 0. 2 DB 0 SP 1 RF 1. 0 PD 0 FT 1. 0 SQ 1. 0 Association rule mining Frequent item-set with support Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong LE 1. 0 PD 0 SQ 1. 0 (99. 7%) SP 1 PD 0 SQ 1. 0 (97. 1%) SP 1 LE 1. 0 PD 0 SQ 1. 0 (96. 8%) DB 0 PD 0 SQ 1. 0 (93. 1%) DB 0 LE 1. 0 PD 0 SQ 1. 0 (92. 8%) DB 0 SP 1 PD 0 SQ 1. 0 (90. 4%) DB 0 SP 1 LE 1. 0 PD 0 SQ 1. 0 (90. 1%) RF 1. 0 SP 1 LE 1. 0 PD 0 SQ 1. 0 (47. 6%) RF 1. 0 DB 0 LE 1. 0 PD 0 SQ 1. 0 (44. 0%)
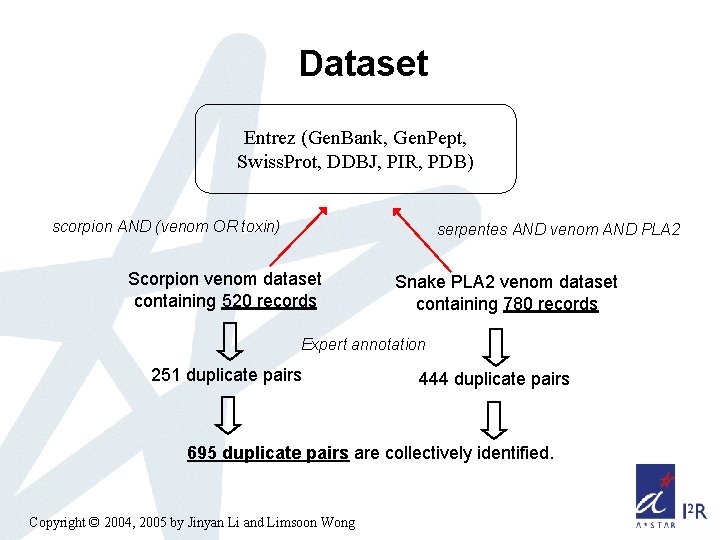
Dataset Entrez (Gen. Bank, Gen. Pept, Swiss. Prot, DDBJ, PIR, PDB) scorpion AND (venom OR toxin) serpentes AND venom AND PLA 2 Scorpion venom dataset containing 520 records Snake PLA 2 venom dataset containing 780 records Expert annotation 251 duplicate pairs 444 duplicate pairs 695 duplicate pairs are collectively identified. Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong
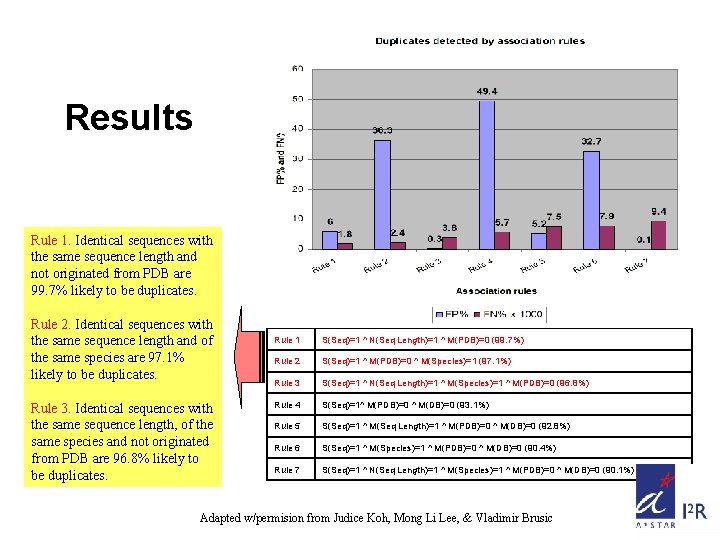
Results Rule 1. Identical sequences with the same sequence length and not originated from PDB are 99. 7% likely to be duplicates. Rule 2. Identical sequences with the same sequence length and of the same species are 97. 1% likely to be duplicates. Rule 3. Identical sequences with the same sequence length, of the same species and not originated from PDB are 96. 8% likely to be duplicates. Rule 1 S(Seq)=1 ^ N(Seq Length)=1 ^ M(PDB)=0 (99. 7%) Rule 2 S(Seq)=1 ^ M(PDB)=0 ^ M(Species)=1 (97. 1%) Rule 3 S(Seq)=1 ^ N(Seq Length)=1 ^ M(Species)=1 ^ M(PDB)=0 (96. 8%) Rule 4 S(Seq)=1^ M(PDB)=0 ^ M(DB)=0 (93. 1%) Rule 5 S(Seq)=1 ^ M(Seq Length)=1 ^ M(PDB)=0 ^ M(DB)=0 (92. 8%) Rule 6 S(Seq)=1 ^ M(Species)=1 ^ M(PDB)=0 ^ M(DB)=0 (90. 4%) Rule 7 S(Seq)=1 ^ N(Seq Length)=1 ^ M(Species)=1 ^ M(PDB)=0 ^ M(DB)=0 (90. 1%) Adapted w/permision from Judice Koh, Mong Li Lee, & Vladimir Brusic

Acknowledgements TIS Prediction Subgraphs in Protein Interactions • Huiqing Liu, Roland Yap, Fanfan Zeng • Haiquan Li, Donny Soh, Chris Tan, See Kiong Ng Treatment Optimization for Childhood ALL Mining Errors from Bio Databases • James Downing, Huiqing Liu, Allen Yeoh • Vladimir Brusic, Judice Koh, Mong Li Lee Prognosis based on Gene Expression Profiling • Huiqing Liu Analysis of Proteomics Data • Huiqing Liu Copyright © 2004, 2005 by Jinyan Li and Limsoon Wong
- Slides: 100