A next generation HLA sequencing method provides robust

A next generation HLA sequencing method provides robust reliable class II typing of an HSCT cohort Anajane G. Smith 1, 2 Shalini E. Pereira 1, 2 Dan E. Geraghty 1, 2 John A. Hansen 1 1 Fred Hutchinson Cancer Research Center, Seattle, WA 2 Scisco Genetics, Inc. , Seattle, WA

Challenges of current HLA SBT The vast array of HLA alleles often generates ambiguous typing results. § Inability to assign the phase of sequence polymorphisms. DRB 1*11: 01, 13: 01 or, DRB 1*11: 04, 13: 02 or, 16 other possible allele combinations. § Polymorphism exists outside of exon 2, the traditionally typed class II antigen recognition site (ARS). DRB 1*12: 01/12: 06/12: 10/12: 17 (exon 3 and signal peptide).
NG-HLA sequencing procedure § Locus specific PCR amplification of genomic DNA. § DRB 1/B 3/B 4/B 5, DQB 1, DQA 1, DPB 1: exons 2 and 3. § 4 primer pairs for DRB genes. § 3 primer pairs for DQB 1. § 2 primer pairs each for DQA 1 and DPB 1. § 1 primer pair for DPA 1 exon 2. § Pool all amplicons into a single aliquot per sample. § Barcode PCR to attach a unique label on each sample. § Pool all reactions in a single tube.

NGS analysis tools § Mi. Seq paired end reads matched with “. fasta” sequences. § Phase between exons determined by coincidence with established databases. § Statistics for the likelyhood of each allele match. § Number of paired end reads with a perfect allele match. § Number of paired end reads with a single mismatch. § Number of unique pairs for a given type. § Consistency between independently derived exon data. § Aggregate quality values of base calls at each

Legacy DRB 1 -DQB 1 typing Typing technology evolved over 20 years. § DRB 1 by SSOP and SBT. § DQB 1 by SSOP, SBT, and serology. § SSOP typed for the alleles known at the time. § Assessed exon 2 polymorphism (ARS). § Reported at the 4 digit allele level or with letter codes. § DRB 1*11: 01/11: 02 = DRB 1*11: 01. § DRB 1*12: 01/12: 06/12: 10/12: 17 = DRB 1*12 DUKV.

Study Population Fred Hutchinson Cancer Research Center § HSCT recipients and related or unrelated donors. § 86% Caucasian. § 14% Asian/AFA/Hispanic/Native American. § 2, 605 DNA specimens plus 83 no DNA controls.

Legacy versus NGS DRB 1 DQB 1 § Alleles compared at the 4 digit level of legacy typing. § Discordant allele resolution. § Original clinical typing records. § NG raw sequencing data. § 96% of the typing results were concordant. § 47 (0. 5%) of NGS allele assignments were discordant. § 350 (3. 5%) of legacy allele assignments were discordant.

Legacy discordant: DRB 1 -DQB 1 § 34 discordant alleles due to exon 2 polymorphism § Alleles not known when SSOP typing was done. § Early SSO probe problems. § 316 discordant alleles due to exon 3 polymorphism. § Legacy DRB 1*14: 01 versus NGS DRB 1*14: 54. § Legacy DQB 1*03: 01 versus NGS DQB 1*03: 19. § Legacy DQB 1*02: 01 versus NGS DQB 1*02: 02. § Above includes 9 new allele sequences by

NGS discordant: DRB 1 -DQB 1 § 27 due to low read sequences. § One allele not assigned in heterozygous samples. § Setting limits for acceptable data is critical for accuracy. § 20 of analysis software errors. § Extra allele assigned in homozygous samples. § One allele not assigned in heterozygous samples. § Errors resolved with software improvements.
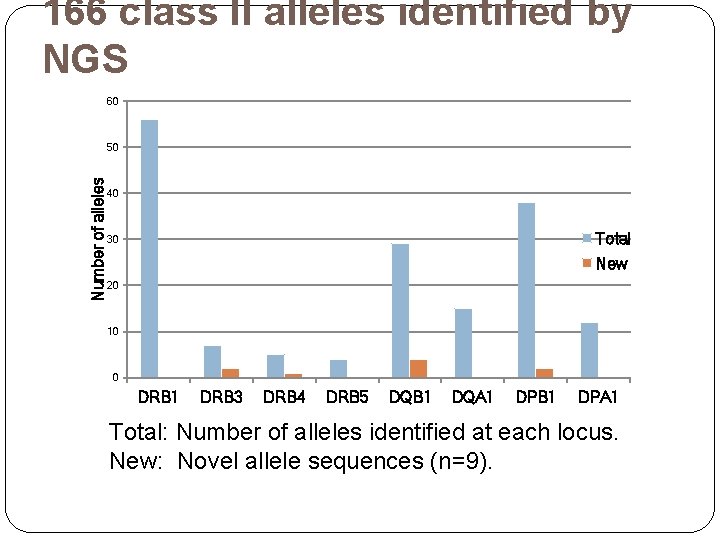
166 class II alleles identified by NGS 60 Number of alleles 50 40 Total 30 New 20 10 0 DRB 1 DRB 3 DRB 4 DRB 5 DQB 1 DQA 1 DPB 1 DPA 1 Total: Number of alleles identified at each locus. New: Novel allele sequences (n=9).

Summary § Our study demonstrates the robust reliability of next generation sequencing for HLA typing. § 99. 5% accuracy of NGS. § Identification of novel allele sequences. § NGS eliminates the complex, time-consuming testing required for resolution of ambiguous typing results. § This technology has been extended to HLA-class I exons 2, 3, and 4 (see poster 1004 -LBP).

Conclusion Next generation sequencing, with fully phased typing by parallel sequencing of individual transcripts, is no longer next, it is now the technology for HLA typing.

Acknowledgements § John A. Hansen § Dan E. Geraghty § Chul-Woo Pyo § Wyatt Nelson § Shalini E. Pereira § Sandy Warnock § Maggie Sprague § Entire staff, past and present, of the FHCRC/SCCA Clinical Immunogenetics Laboratory for the legacy typing.
- Slides: 13