A multistrain highresolution mouse haplotype map reveals three












- Slides: 22

A multi-strain, high-resolution mouse haplotype map reveals three distinctive genetic signatures Laboratory of Population Genetics

Motivations 1. An accurate high-resolution haplotype map of the mouse genome enables prioritization of QTL candidate genes 2. Different haplotype block structures have been reported in different studies • • • >10 MB block size in GNF study (Wiltshire et al, PNAS 2003) 1. 0 -2. 0 Mb block size in WI study (Wade et al, Nature 2002) 100 -150 kb block size in a 8 MB region chr 19 (Park et al, Genome Research 2003) 3. Analysis of a 10 Mb region on chromosome 7 using the Celera mouse SNPs reveals a different genetic variation pattern 4. Celera mouse chromosome 16 SNP data are publicly available Laboratory of Population Genetics

Objectives 1. Develop an integrated, high resolution, multi-strain mouse haplotype map 2. Compare the haplotype structure derived from highdensity SNPs with those derived from low density markers 3. Perform experimental validation in regions of conflict and in regions of interest across 20 inbred strains 4. Analyze biological factors that have contributed to the formation mouse genetic variation patterns Laboratory of Population Genetics

Data Sources 1. Chromosome 16 reference sequence • MGSCv 3 (NCBI build 30, Feb. 2003) 2. SNP Data Laboratory of Population Genetics

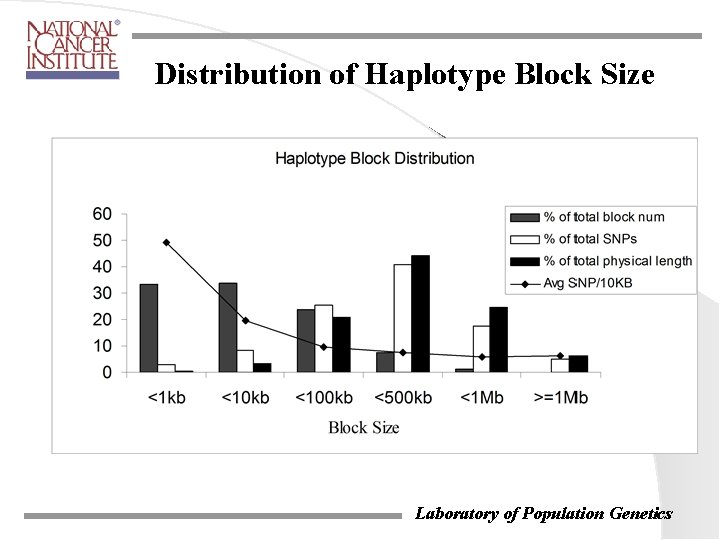
Construction of Multi-Strain Haplotype Blocks with High Density SNP Markers Method • • • Greedy algorithm that starts with two-haplotype per block Seed: a minimum of two adjacent SNPs with no-ambiguity in haplotype assignment Singleton SNP that breaks the two-haplotype configuration does not affect block extension Results • • 2, 083 blocks 65, 068 (95% ) Celera SNPs in 5 laboratory inbred strains. Laboratory of Population Genetics

Distribution of Haplotype Block Size Laboratory of Population Genetics

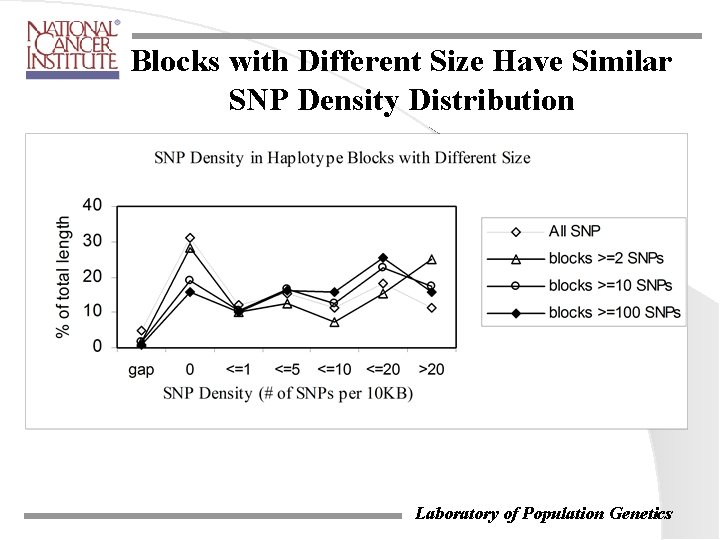
Blocks with Different Size Have Similar SNP Density Distribution Laboratory of Population Genetics

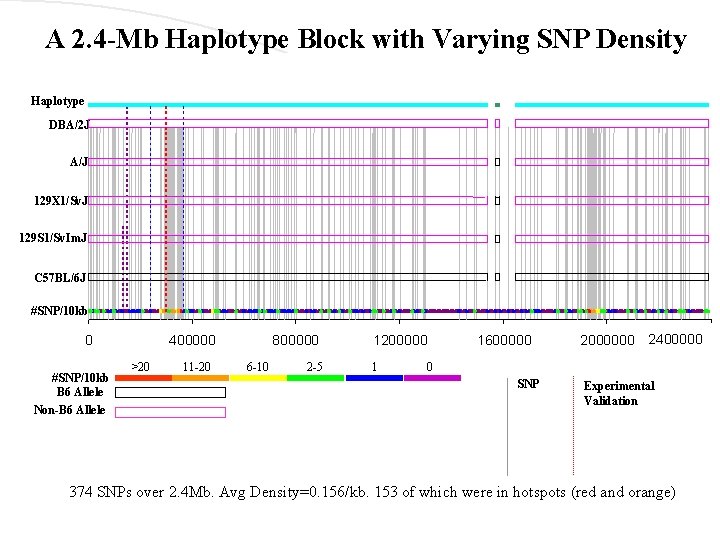
A 2. 4 -Mb Haplotype Block with Varying SNP Density Haplotype DBA/2 J A/J 129 X 1/Sv. J 129 S 1/Sv. Im. J C 57 BL/6 J #SNP/10 kb 0 #SNP/10 kb B 6 Allele Non-B 6 Allele 400000 >20 11 -20 800000 6 -10 2 -5 1200000 1 1600000 2000000 2400000 0 SNP Experimental Validation 374 SNPs over 2. 4 Mb. Avg Density=0. 156/kb. 153 of which were in hotspots (red and orange)

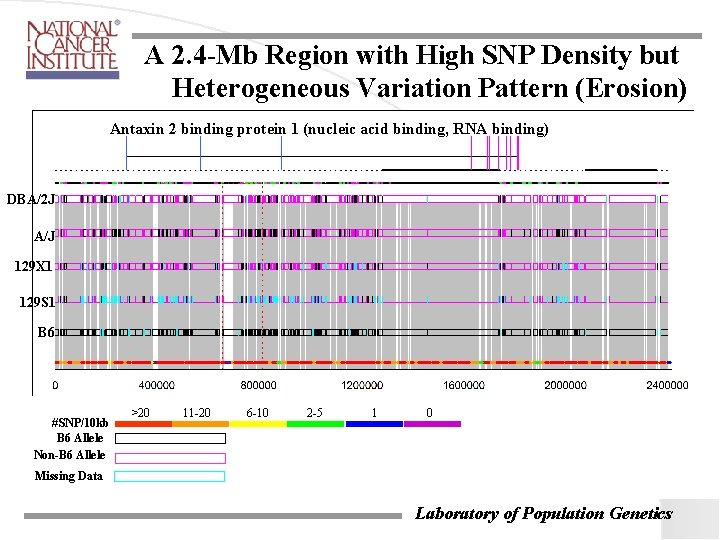
A 2. 4 -Mb Region with High SNP Density but Heterogeneous Variation Pattern (Erosion) Antaxin 2 binding protein 1 (nucleic acid binding, RNA binding) DBA/2 J A/J 129 X 1 129 S 1 B 6 #SNP/10 kb B 6 Allele Non-B 6 Allele >20 11 -20 6 -10 2 -5 1 0 Missing Data Laboratory of Population Genetics

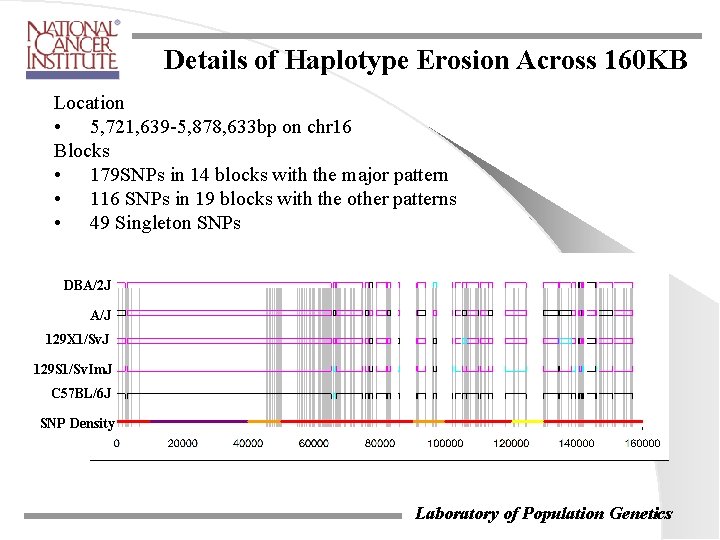
Details of Haplotype Erosion Across 160 KB Location • 5, 721, 639 -5, 878, 633 bp on chr 16 Blocks • 179 SNPs in 14 blocks with the major pattern • 116 SNPs in 19 blocks with the other patterns • 49 Singleton SNPs DBA/2 J A/J 129 X 1/Sv. J 129 S 1/Sv. Im. J C 57 BL/6 J SNP Density Laboratory of Population Genetics

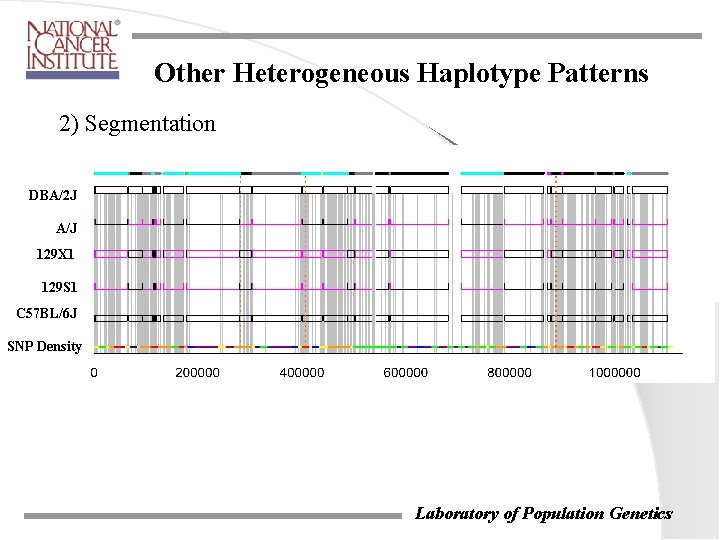
Other Heterogeneous Haplotype Patterns 2) Segmentation DBA/2 J A/J 129 X 1 129 S 1 C 57 BL/6 J SNP Density Laboratory of Population Genetics

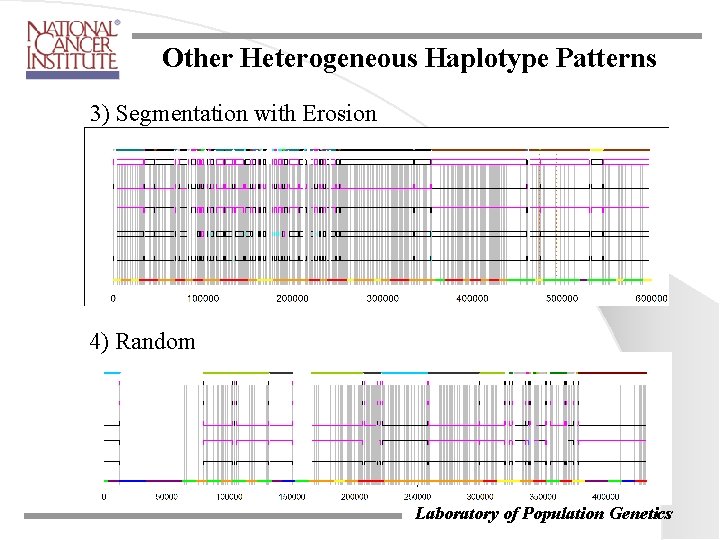
Other Heterogeneous Haplotype Patterns 3) Segmentation with Erosion 4) Random Laboratory of Population Genetics

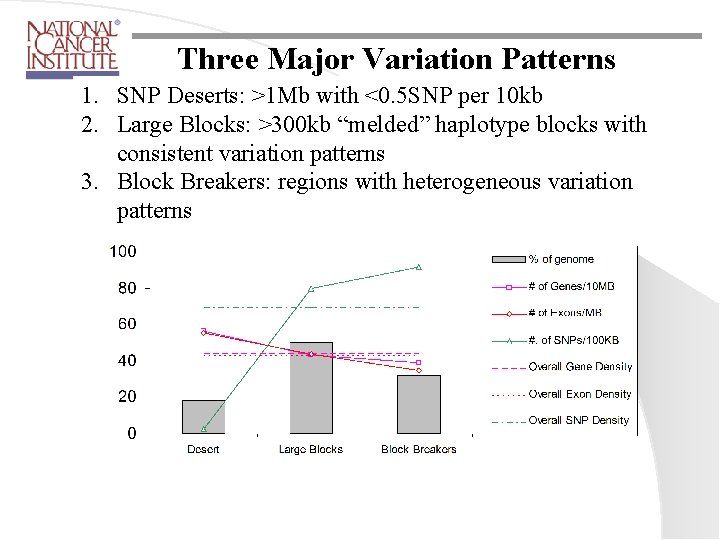
Three Major Variation Patterns 1. SNP Deserts: >1 Mb with <0. 5 SNP per 10 kb 2. Large Blocks: >300 kb “melded” haplotype blocks with consistent variation patterns 3. Block Breakers: regions with heterogeneous variation patterns

Predictive Power of Haplotype Structures • Test the ability to use the haplotype structure in one study to predict allelic variations in another study • Our Haploytpe Blocks • 98% accuracy on WI B 6/129 S 1 SNPs that do not overlap with Celera SNPs • 92% accuracy on GNF B 6/129 S 1/AJ/DBA haplotypes • WI B 6/129 S 1 Haplotype Blocks • 74% accuracy on Celera B 6/129 S 1 genotypes • 85% accuracy on GNF B 6/129 S 1 genotypes • 80% GNF markers are non-polymorphic across inbred strains used in Celera and WI shotgun sequencing Laboratory of Population Genetics

SNP Deserts in Chromosome 16 • 6 >1 Mb SNP deserts in the five inbred strains used for Celera shotgun sequencing • All 6 SNP deserts overlap with WI SNP deserts conserved across all WI strains • 0. 21% WI B 6/Sv. J SNPs in our SNP deserts • 0. 97% WI all SNPs in our SNP deserts • 5 out of the 6 deserts have at least one end as part of large haplotype blocks • SNP deserts are not genetically homogeneous • There are STRP polymorphisms • There are indel polymorphisms Laboratory of Population Genetics

Validation of a SNP Desert 000010000000000000000000000000000000000000000000000000000000000 00 1000 11111 01 0000000000000000000000000000000000000000000000000000000000000 00 1110100011111111111111111111111111111111111111111111 011111101111101 000111 N 0101 11 1111111101111111 NNNNNN 1111111111111 01111111011111 NN 011111 N 10001111111111111111111 0 110001 • • • 11 Other Lab inbred B 6, AKRJ Skive Czech 5 STRPs and 1 SNP in a 15 kb SNP desert in all laboratory inbred strains The STRPs and the 1 SNP have the same variation pattern as the neighboring regions with high SNP density among the laboratory inbred strains Additional 120 SNPs discovered between the laboratory inbred strains and feral inbred strains Laboratory of Population Genetics

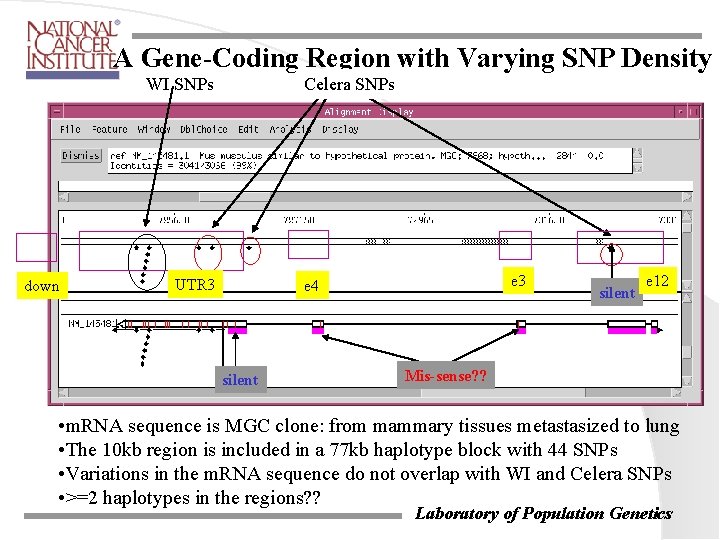
A Gene-Coding Region with Varying SNP Density WI SNPs down Celera SNPs UTR 3 e 4 silent e 12 Mis-sense? ? • m. RNA sequence is MGC clone: from mammary tissues metastasized to lung • The 10 kb region is included in a 77 kb haplotype block with 44 SNPs • Variations in the m. RNA sequence do not overlap with WI and Celera SNPs • >=2 haplotypes in the regions? ? Laboratory of Population Genetics

Results of Experimental Validation >down hap 1 011110 129 s 1; 129 x 1; AJ; BALB; C 3 HHe; DBA hap 2 000000 AKRJ; C 57 BL hap 3 110001 Czech; Skive >UTR 3 /num_SNP=25 /num_strain=10 /num_hap=4 hap 1 1100000010000010 129 s 1; 129 x 1; AJ; BALB; C 3 HHe; DBA 2 J; hap 4 110001010011100 N 11000 AKRJ; hap 3 111111001110101 Czech; Skive; hap 2 0000000000000 C 57 BL; >NM_145481_e 12 /num_SNP=7 /num_strain=10 /num_hap=4 >e 4 hap 1 0000100 129 s 1; 129 x 1; AJ; BALB; hap 1 0 Others hap 2 0010001 AKRJ; hap 2 1 Czech; Skive hap 3 0000000 C 3 HHe; C 57 BL; DBA 2 J; hap 4 1101011 Czech; Skive; Silent SNP validates Mis-sense SNP does not validate

Mouse c. SNPs • Synonymous: 185 • Non-synonymous: 100 Laboratory of Population Genetics

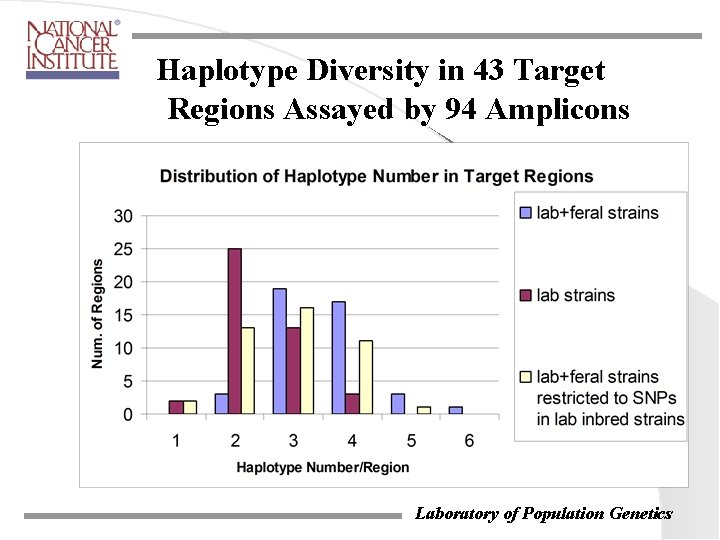
Haplotype Diversity in 43 Target Regions Assayed by 94 Amplicons Laboratory of Population Genetics

Conclusions • We have compiled an accurate, multi-strain, high-resolution haplotype map for mouse chromosome 16 • We have discovered three distinctive genetic variation patterns for laboratory inbred mouse: SNP deserts, large blocks and block breakers • Large haplotype blocks may consist regions with varying SNP density • Selection in inbreeding may have an effect on SNP distribution in protein coding regions as well as SNP rate in gene coding regions • Our method is scalable for whole-genome analysis Laboratory of Population Genetics

Acknowledgement Laboratory of Population Genetics • Ken Buetow • Kent Hunter • Michael Gandolph • Bill Rowe • Michael Edmonson • Jenny Kelly University of Wisconsin • Rob Williams Laboratory of Population Genetics