A HandsOn Tutorial High Performance Computing in Bioinformatics

A Hands-On Tutorial: High Performance Computing in Bioinformatics Medical Center Information Technology High-Performance Computing Core September 7 th 2018 Michael Costantino

Introduction High Performance Computing (HPC) generally refers to the practice of aggregating computing power in a way that delivers much higher performance than what one could get out of a typical desktop computer. In NYU Langone the High Performance Computing (HPC) Core is the resource for performing computational research at scale and for analyzing big data. HPC is a Shared Resource. Big. Purple is a HPC cluster running on Redhat 7. 4 Operating system. It is comprised of 90 compute nodes (40 CPU cores each) of which 32 include the total of 156 GPU cards; 16 service nodes; Infini. Band EDR 100 Gb interconnect. The attached storage runs on GPFS file system with. 5 PB flash scratch (non replicated) and 4 PB spinning disk permanent (replicated). Bright Computing is the cluster manager suite. Slurm is chosen as the resource manager and Job scheduler with the “cgroup” feature deployed on Big. Purple. 2
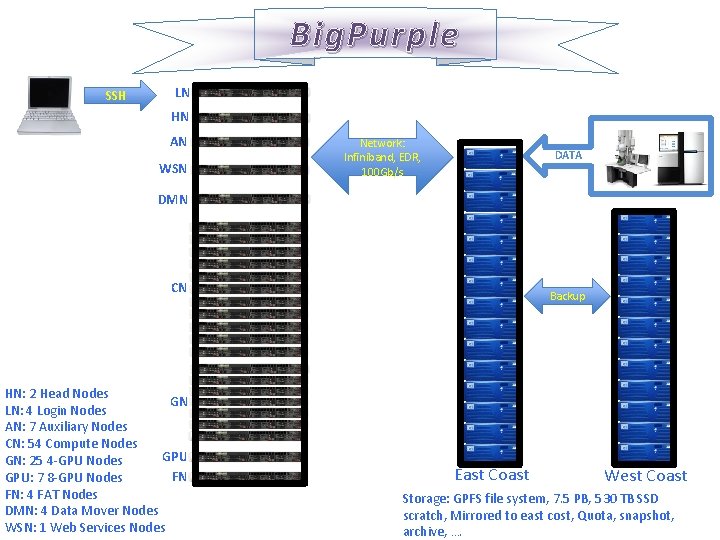
Big. Purple SSH LN HN AN WSN Network: Infiniband, EDR, 100 Gb/s DATA DMN CN HN: 2 Head Nodes GN LN: 4 Login Nodes AN: 7 Auxiliary Nodes CN: 54 Compute Nodes GPU GN: 25 4 -GPU Nodes FN GPU: 7 8 -GPU Nodes FN: 4 FAT Nodes DMN: 4 Data Mover Nodes WSN: 1 Web Services Nodes Backup East Coast West Coast Storage: GPFS file system, 7. 5 PB, 530 TB SSD scratch, Mirrored to east cost, Quota, snapshot, archive, ….

Node Specifications Ø Head Nodes: 2×intel Xion Skylake 2. 4 GHz (total of 40 cores), 384 GB RAM, EDR Infiniband interconnected, 2 TB SATA and 2 TB SSD local disks. Ø Login Nodes: 2×intel Xion Skylake 2. 4 GHz (total of 40 cores), 384 GB RAM, EDR Infiniband interconnected and 40 Gb/s outbound connect, 2 TB SATA and 2 TB SSD local disks. Ø Auxiliary Nodes: 2×intel Xion Skylake 2. 4 GHz (total of 40 cores), 384 GB RAM, EDR Infiniband interconnected and 40 Gb/s outbound connect, 2 TB SATA and 2 TB SSD local disks. Ø Compute Nodes: 2×intel Xion Skylake 2. 4 GHz (total of 40 cores), 384 GB RAM, EDR Infiniband interconnected, 2 TB SATA and 2 TB SSD local disks. Ø 4 -way GPU Nodes: 2×intel Xion Skylake 2. 4 GHz (total of 40 cores), 384 GB RAM, 4 NVIDIA® Tesla V 100 GPUs, GPU-to-GPU NVLINK, Supports GPUDirect RDMA , EDR Infiniband interconnected, 2 TB SATA and 2 TB SSD local disks. Ø 8 -way GPU Nodes: 2×intel Xion Skylake 2. 4 GHz (total of 40 cores), 768 GB RAM, 8 NVIDIA® Tesla V 100 GPUs, GPU-to-GPU NVLINK, Supports GPUDirect RDMA , EDR Infiniband interconnected, 2 TB SATA and 4 TB SSD local disks. Ø FAT Nodes: 2×intel Xion Skylake 2. 4 GHz (total of 40 cores), 1536 GB RAM, EDR Infiniband interconnected, 2 TB SATA and 4 TB SSD local disks. Ø Data Mover Nodes: 2×intel Xion Skylake 2. 4 GHz (total of 40 cores), 384 GB RAM, EDR Infiniband interconnected and 40 Gb/s outbound connect, 2 TB SATA and 2 TB SSD local disks. Ø Web Services Node: 2×intel Xion Skylake 2. 4 GHz (total of 40 cores), 384 GB RAM, EDR Infiniband interconnected and 40 Gb/s outbound connect, 2 TB SATA and 2 TB SSD local disks.

Accessing Big. Purple • Login to The System – Linux/Mac • Open Terminal / Command Line • ssh -X <username>@bigpurple. nyumc. org – Windows • Download install Putty https: //www. putty. org/ • Hostname=bigpurple. nyumc. org • User name = NYU Langone Kerberos ID • Password = NYU Langone password (same as email) • More Information at: http: //bigpurple-ws. nyumc. org/wiki/index. php/Getting-Started#Accessing_Big. Purple
Splash Screen

Course Environment • • Home Directory: /gpfs/home/<username> Scratch Space: /gpfs/scratch/<username> Course Directory: /gpfs/data/courses/bmsc 4449 Examples (RO): /gpfs/data/courses/bmsc 4449/examples Sample Datasets (RO): /gpfs/data/courses/bmsc 4449/samples Course Shared (RW): /gpfs/data/courses/bmsc 4449/shared Course Group: course_bmsc 4449

Utilizing Big. Purple • Plan your job: Test it and beware about how many cores, how much RAM it needs and also estimate how long it takes for the job to finish. Then scale up gradually. • Login to the system: ssh –X your_kerberos_id@bigpurple. nyumc. org Unless you want to move data, run a gui, use an auxilary node, …. Navigate and Prepare: Linux commands: cd, ls, mkdir, cp, rm, cat, which, head, tail, …, Familiarize yourself with man pages. Familiarize yourself with an editor in linux (nano). Beware of hidden characters when you cp/paste in a Windows environment Determine where (using scratch disk, stage-in-stage-out, …)and how (parallel, serial, interactive, queued, GUI, …) you want to run your job. • • Setup your environment: Setup your needed modules. module list, module avail, module load, module rm, …. • Utilize queuing system slurm to submit, control and monitor your job: submision script, partitions, fare-share, ….

Slurm is an open source, fault-tolerant, and highly scalable cluster management and job scheduling system for large and small Linux clusters. • Job submission, job monitoring, job control: sinfo, sbatch, srun, scontrol, squeue, sacct, … • Partitions, fair-share • Interactive jobs: srun -p cpu_short --mem-per-cpu=4 G —time=1: 00 --x 11 --pty bash

Hands-on Lab • • ssh –X siavoa 01@bigpurple. nyumc. org pwd ls mkdir project 1 cd project 1 ls nano #!/bin/bash #SBATCH --job-name=serial_job_test # Job name #SBATCH --mail-type=END, FAIL # Mail events (NONE, BEGIN, END, FAIL, ALL) #SBATCH --mail-user=first. last@nyumc. org # Where to send mail #SBATCH --ntasks=1 # Run on a single CPU #SBATCH --mem=1 gb # Job memory request #SBATCH --time=00: 05: 00 # Time limit hrs: min: sec #SBATCH --output=serial_test_%j. log # Standard output and error log pwd; hostname; date sleep 300 date

• • • module avail module load your_required_module list sinfo squeue –u your_kerberos_id sbatch. job squeue –u your_kerberos_id scontrol show jobid=job_id scontrol update Job. Id=job_id Partition=cpu_short squeue –u your_kerberos_id

Examples 1. Login to Big. Purple 2. Execute the Following Commands: • cd /gpfs/data/courses/bmsc 4449/examples/0_Prepare. Environment/ • ls • cat README

Points To Remember • A good source of information: http: //bigpurple-ws. nyumc. org/wiki • email HPC Administrators for problem/questions: hpc_admins@nyulangone. org when you are sending us an email please include the error and the command you issued. It is vital for troubleshooting. • Remember that this is a shared resource. Login nodes are not for running jobs, instead use compute nodes, data mover nodes, auxiliary nodes, ….
- Slides: 13