A Field Guide to Gen Bank and NCBI

A Field Guide to Gen. Bank and NCBI Molecular Biology Resources slightly modified from Peter Cooper ftp: //ftp. ncbi. nih. gov/pub/cooper/Field. Guide/ Eric Sayers ftp: //ftp. ncbi. nih. gov/pub/sayers/Field_Guide/U_Penn/

NCBI Resources • About NCBI • NCBI Sequence Databases – Primary Database – Gen. Bank – Derivative Databases - Ref. Seq • Entrez Databases and Text Searching • BLAST Services • Genomic Resources

The National Center for Biotechnology Information (NCBI) • Created as a part of NLM in 1988 • • – – Establish public databases Perform research in computational biology Develop software tools for sequence analysis Disseminate biomedical information Tools: BLAST(1990), Entrez (1992) Gen. Bank (1992) Free MEDLINE (Pub. Med, 1997) Human genome (2001)

NCBI Home Page http: //www. ncbi. nlm. nih. gov To learn more, visit the “Site Map” and “About NCBI” web pages

About NCBI
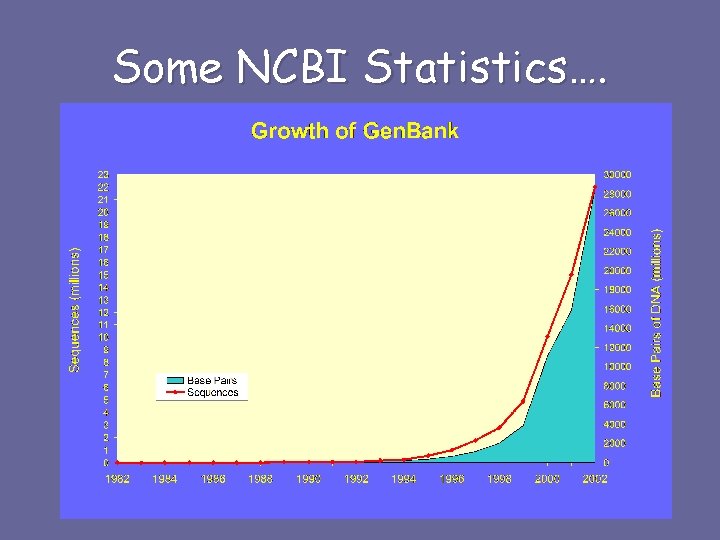
Some NCBI Statistics….
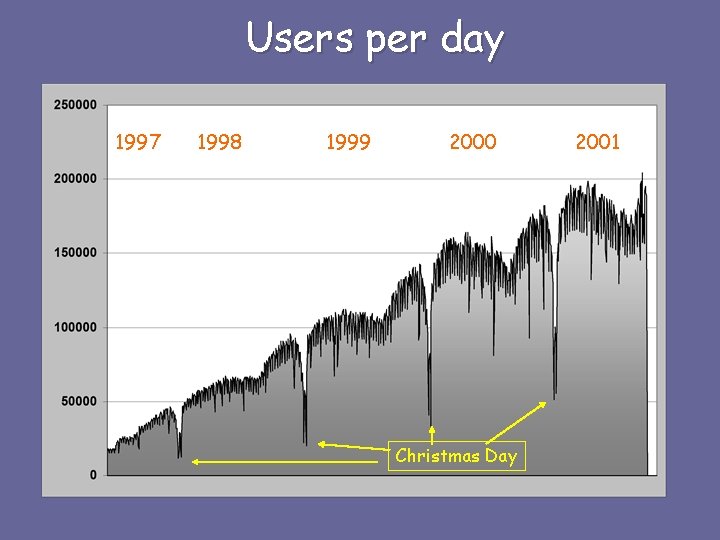
Users per day 1997 1998 1999 2000 Christmas Day 2001

Molecular Databases • Primary Databases – Original submissions by experimentalists – Database staff organize but don’t additional information • Example: Gen. Bank • Derivative Databases – Human curated • compilation and correction of data • Example: SWISS-PROT, NCBI Ref. Seq m. RNA – Computationally Derived • Example: Uni. Gene – Combinations • Example: NCBI Genome Assembly

What is Gen. Bank? NCBI’s Primary Sequence Database • Nucleotide only sequence database • Gen. Bank Data – Direct submissions individual records (Bank. It, Sequin) – Batch submissions via email (EST, GSS, STS) – ftp accounts established for sequencing centers • Data shared amongst three collaborating databases: – – – Gen. Bank DNA Database of Japan (DDBJ). European Molecular Biology Laboratory Database (EMBL)
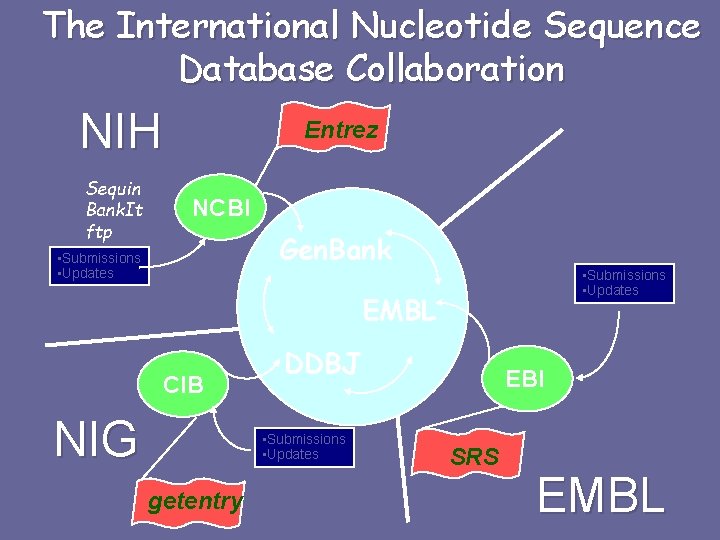
The International Nucleotide Sequence Database Collaboration NIH Sequin Bank. It ftp Entrez NCBI Gen. Bank • Submissions • Updates EMBL CIB NIG DDBJ • Submissions • Updates getentry EBI SRS EMBL

Gen. Bank: NCBI’s Primary Sequence Database Release 133 22, 318, 883 28, 507, 990, 166 110, 000 + December 2002 Records Nucleotides Species • full release every two months • incremental and cumulative updates daily • available only through internet ftp: //ftp. ncbi. nih. gov/genbank/ >90 Gigabytes of data
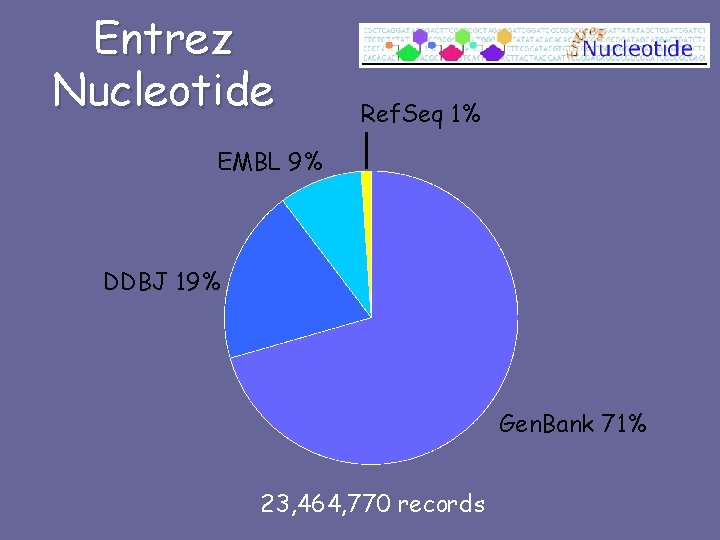
Entrez Nucleotide Ref. Seq 1% EMBL 9% DDBJ 19% Gen. Bank 71% 23, 464, 770 records
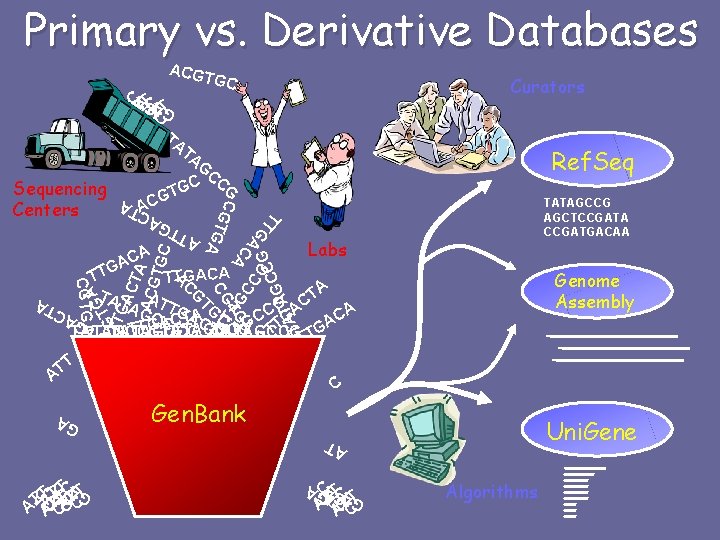
Primary vs. Derivative Databases ACGT GC C TC ATCATCT TA Curators GAG A A TA G C AT T GC C A AC T A G T AT G CTGA CT A ACG TG TT G A C A TTGACA C A TG CG C TG G A CGTGA G TT TA T A A AC G G CC G ATA T G AC CG TG Sequencing Centers Ref. Seq G CC C G G TATAGCCG AGCTCCGATA CCGATGACAA Labs Genome Assembly TA T TA Gen. Bank Uni. Gene AT C TC ATCATCT GAG A AC T G AT T GA TACTTTCTT T A ATCA C TA A C AG TTG CGGA A A CA CC C T C C A A G T G G TTATAGCCG A TA AT TATAGCCG ATT TATAGCCG TG T A T T AT C Algorithms GAGA GAG A

Traditional Gen. Bank Divisions • Direct Submissions (Sequin and Bank. It) • Accurate • Well characterized BCT INV MAM PHG PLN PRI ROD SYN VRL VRT Bacterial and Archeal Invertebrate Mammalian (ex. ROD and PRI) Phage Plant and Fungal Primate Rodent Synthetic (cloning vectors) Viral Other Vertebrate
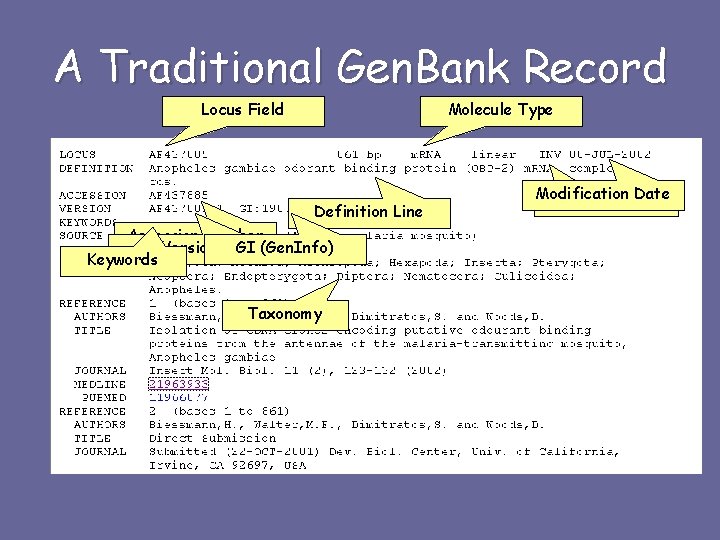
A Traditional Gen. Bank Record Locus Field Molecule Type Definition Line Accession Number Version GI (Gen. Info) Keywords Taxonomy Modification Date Gen. Bank Division

A Traditional Gen. Bank Record

Bulk Sequence Divisions of Gen. Bank • Batch Submissions (email and ftp) • Inaccurate • Poorly Characterized EST STS GSS HTG HTC Expressed Sequence Tagged Site Genome Survey Sequence High Throughput Genomic High Throughput c. DNA
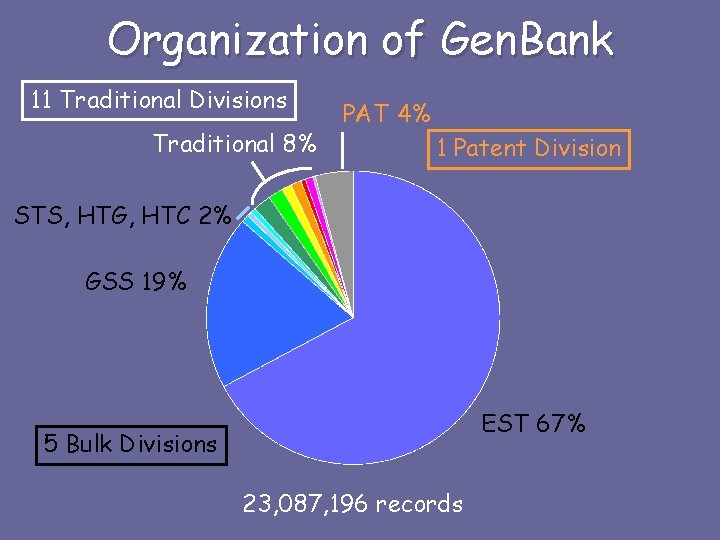
Organization of Gen. Bank 11 Traditional Divisions Traditional 8% PAT 4% 1 Patent Division STS, HTG, HTC 2% GSS 19% EST 67% 5 Bulk Divisions 23, 087, 196 records

What is Uni. Gene? A gene-oriented view of sequence entries • Mega. Blast-based automated sequence clustering • Nonredundant set of gene-oriented clusters • Each cluster represents a unique gene • Provides information on tissue-specific expression and map locations • Includes well-characterized genes and novel ESTs • Useful for gene discovery and selection of mapping reagents
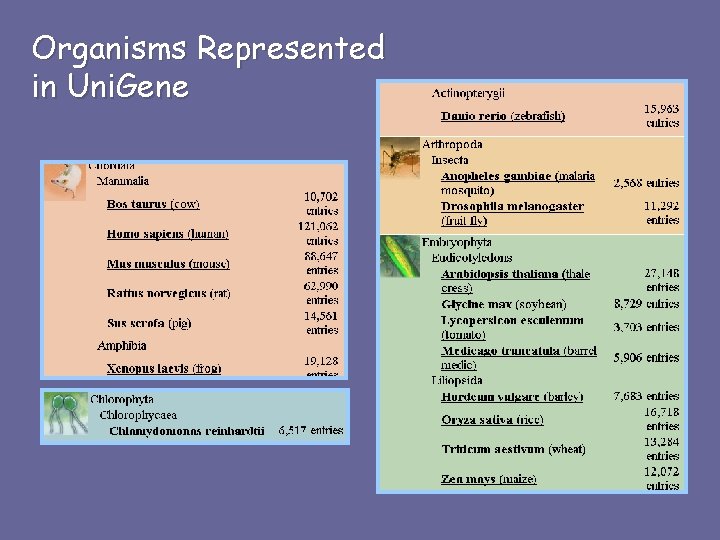
Organisms Represented in Uni. Gene
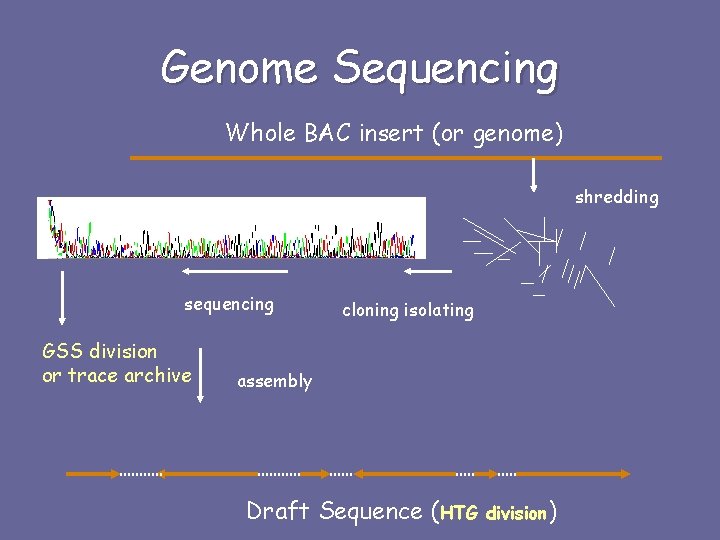
Genome Sequencing Whole BAC insert (or genome) shredding sequencing GSS division or trace archive cloning isolating assembly Draft Sequence (HTG division)
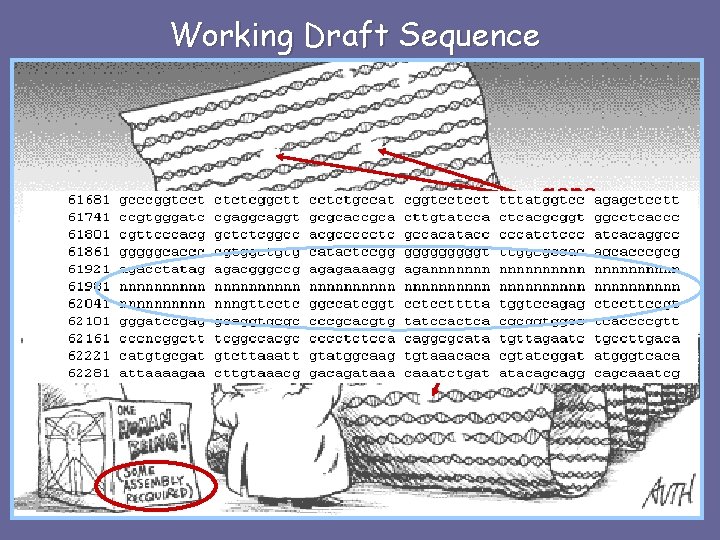
Working Draft Sequence gaps
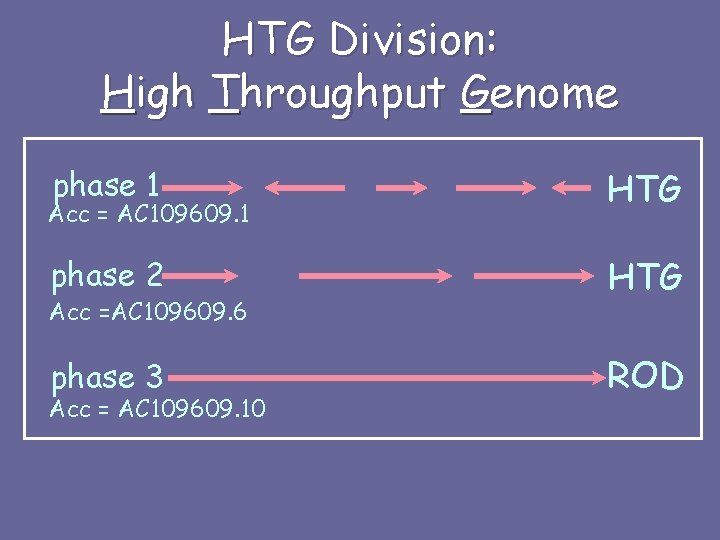
HTG Division: High Throughput Genome phase 1 HTG phase 2 HTG phase 3 ROD Acc = AC 109609. 1 Acc =AC 109609. 6 Acc = AC 109609. 10

HTG Division: High Throughput Genome

NCBI’s Third Party Annotation (TPA) Database NEW • NCBI now accepts the submission of new annotations of existing Gen. Bank sequences; • Facilitates the annotation of genomes by experts;

A Sample TPA record

Ref. Seq: NCBI’s Derivative Sequence Database • Curated transcripts and proteins – reviewed – human, mouse, rat, fruit fly, zebrafish, arabidopsis • Human model transcripts and proteins • Assembled Genomic Regions (contigs) – draft human genome – mouse genome • Chromosome records – – – Microbial viral organelle

The Ref. Seq Accession Numbers m. RNAs and Proteins NM_123456 NP_123456 NR_123456 XM_123456 XP_123456 XR_123456 Gene Records NG_ 123456 Assemblies NT_ 123456 NW_123456 NC_ 123456 NR_ 123456 human Curated m. RNA mouse rat Curated Protein fruit fly Curated non-coding RNA zebrafish Predicted Transcript (human, mouse) Arabidopsis Predicted Protein (human, mouse) Predicted non-coding RNA Reference Genomic Sequence (human) Contig (Mouse and Human) Supercontig (Mouse) Chromosome (Microbial, Viral, Arabidopsis ) Interim Identifier for Microbial Chromosomes

Curated Ref. Seq Records: NM_, NP_

Entrez: Linking and Neighboring

The Entrez Databases
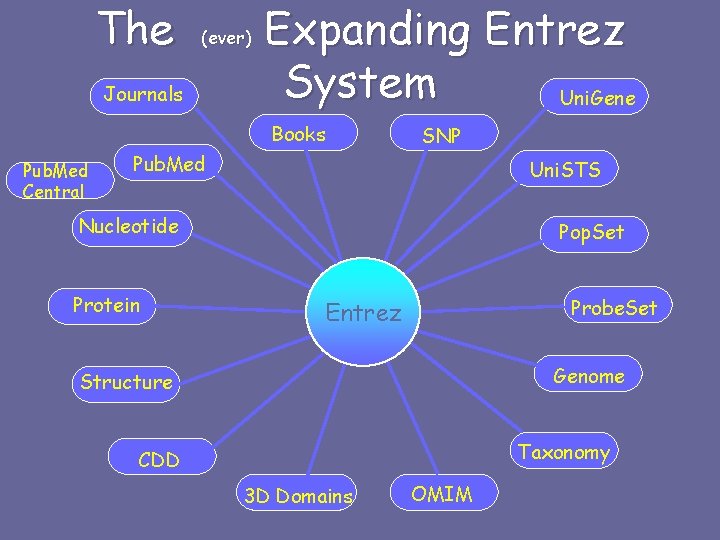
The (ever) Journals Expanding Entrez System Uni. Gene Books Pub. Med Central SNP Pub. Med Uni. STS Nucleotide Protein Pop. Set Probe. Set Entrez Genome Structure Taxonomy CDD 3 D Domains OMIM
Entrez Nucleotides glucose 6 phosphate dehydrogenase
![Document Summaries: glucose 6 phosphate dehydrogenase[All Fields] = 748 hits Document Summaries: glucose 6 phosphate dehydrogenase[All Fields] = 748 hits](http://slidetodoc.com/presentation_image_h/000aa34676aefab181e33ad356b471a6/image-34.jpg)
Document Summaries: glucose 6 phosphate dehydrogenase[All Fields] = 748 hits
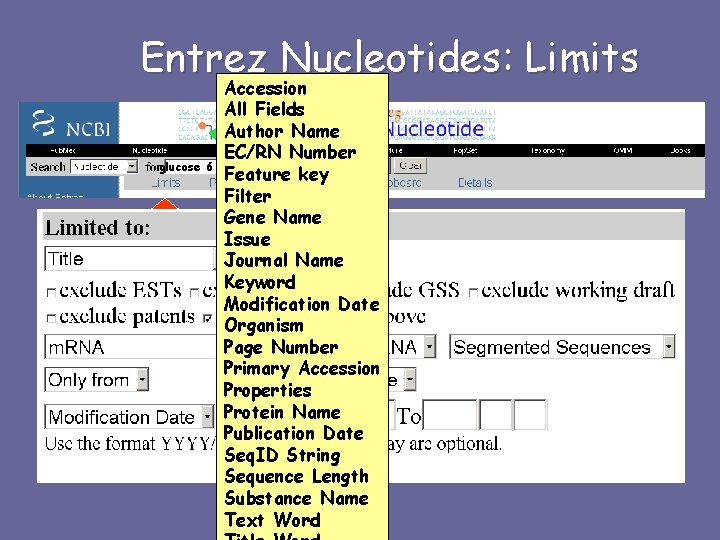
Entrez Nucleotides: Limits Accession All Fields Author Name EC/RN Number glucose 6 phosphate dehydrogenase Feature key Filter Gene Name Issue Journal Name Keyword Modification Date Organism Page Number Primary Accession Properties Protein Name Publication Date Seq. ID String Sequence Length Substance Name Text Word
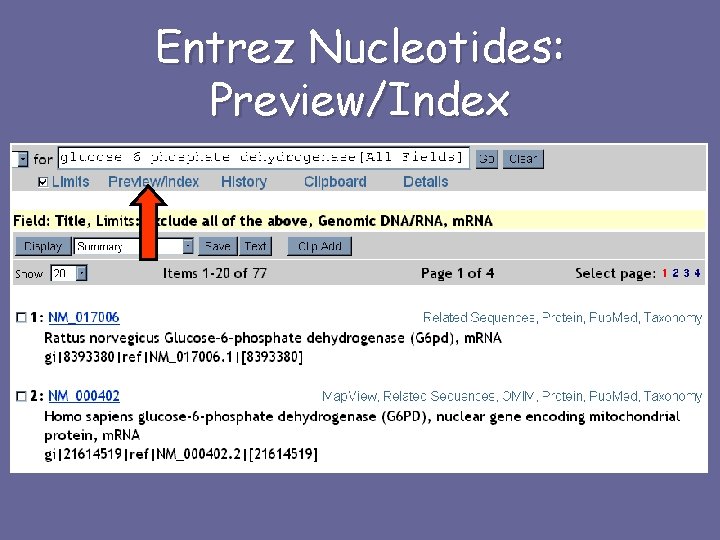
Entrez Nucleotides: Preview/Index
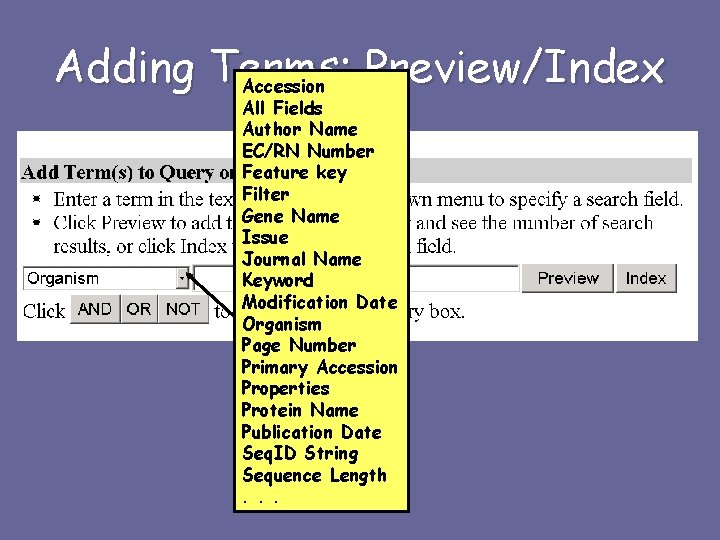
Adding Terms: Preview/Index Accession All Fields Author Name EC/RN Number Feature key Filter Gene Name Issue Journal Name Keyword Modification Date Organism Page Number Primary Accession Properties Protein Name Publication Date Seq. ID String Sequence Length. . .

Plant G 6 PD m. RNAs
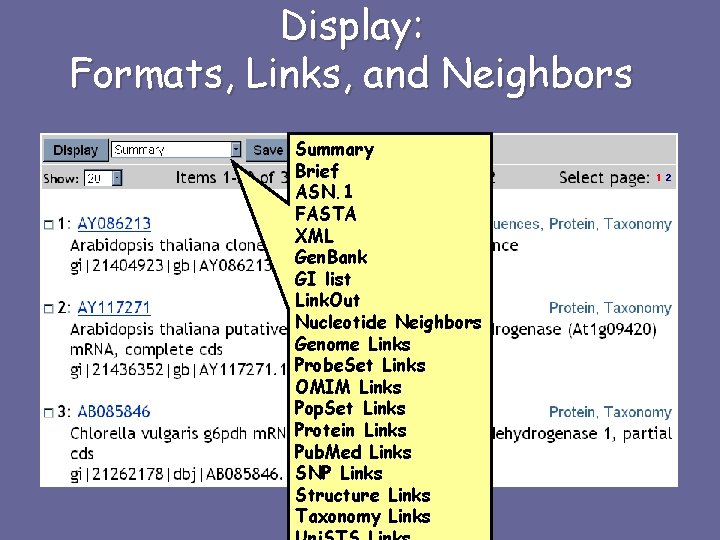
Display: Formats, Links, and Neighbors Summary Brief ASN. 1 FASTA XML Gen. Bank GI list Link. Out Nucleotide Neighbors Genome Links Probe. Set Links OMIM Links Pop. Set Links Protein Links Pub. Med Links SNP Links Structure Links Taxonomy Links
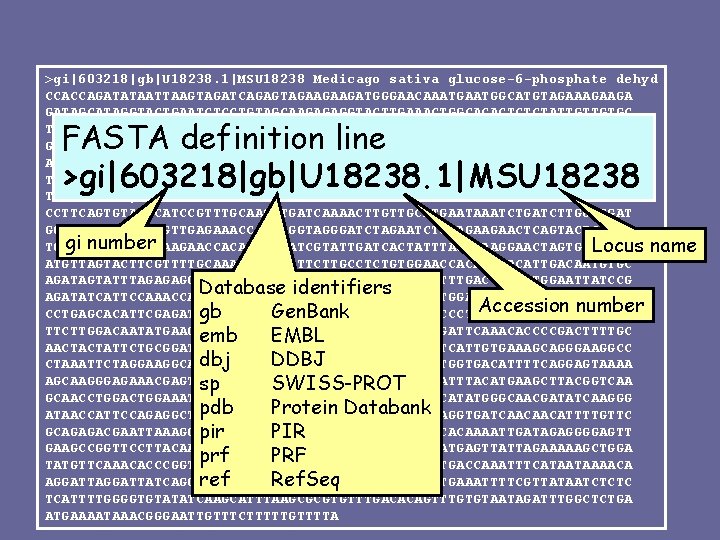
>gi|603218|gb|U 18238. 1|MSU 18238 Medicago sativa glucose-6 -phosphate dehyd CCACCAGATATAATTAAGTAGATCAGAGTAGAAGAAGATGGGAACAAATGGCATGTAGAAGA GATAGCATAGGTACTGAATCTCCTGTAGCAAGAGAGGTACTTGAAACTGGCACACTCTCTATTGTTGTGC TTGGTGCTTCTGGTGATCTTGCCAAGAAGAAGACTTTTCCTGCACTTTTTCACTTATATAAACAGGAATT GTTGCCACCTGATGAAGTTCACATTTTTGGCTATGCAAGGTCAAAGATCTCCGATGATGAATTGAGAAAC AAATTGCGTAGCTATCTTGTTCCAGAGAAAGGTGCTTCTCCTAAACAGTTAGATGATGTATCAAAGTTTT TACAATTGGTTAAATATGTAAGTGGCCCTTATGATTCTGAAGATGGATTTCGCTTGTTGGATAAAGAGAT TTCAGAGCATGAATATTTGAAAAATAGTAAAGAGGGTTCATCTCGGAGGCTTTTCTATCTTGCACTTCCT > CCTTCAGTGTATCCGTTTGCAAGATGATCAAAACTTGTTGCATGAATAAATCTGATCTTGGTGGAT GGACACGCGTTGTTGTTGAGAAACCCTTTGGTAGGGATCTAGAATCTGCAGAAGAACTCAGTACTCAGAT gi number TGGAGAGTTATTTGAAGAACCACAGATTTATCGTATTGATCACTATTTAGGAACTAGTGCAAAAC Locus name ATGTTAGTACTTCGTTTTGCAAATCGGTTCTTCTTGCCTCTGTGGAACCACATTGACAATGTGC AGATAGTATTTAGAGAGGATTTTGGAACTGATGGTCGTGGTGGATATTTTGACCAATATGGAATTATCCG Database identifiers AGATATCATTCCAAACCATCTGTTGCAGGTTCTTTGCTTGATTGCTATGGAAAAACCCGTTTCTCTCAAG Accession number gb Gen. Bank CCTGAGCACATTCGAGATGAGAAAGTGAAGGTTCTTGAATCAGTACTCCCTATTAGAGATGATGAAGTTG TTCTTGGACAATATGAAGGCTATACAGATGACCCAACTGTACCGGACGATTCAAACACCCCGACTTTTGC emb EMBL AACTACTATTCTGCGGATACACAATGAAAGATGGGAAGGTGTTCCTTTCATTGTGAAAGCAGGGAAGGCC dbj DDBJ CTAAATTCTAGGAAGGCAGAGATTCGGGTTCAAGGATGTTCCTGGTGACATTTTCAGGAGTAAAA AGCAAGGGAGAAACGAGTTTGTTATCCGCCTACAACCTTCAGAAGCTATTTACATGAAGCTTACGGTCAA sp SWISS-PROT GCAACCTGGAAATGTCTGCAGTTCAAAGTGAACTAGACTTGTCATATGGGCAACGATATCAAGGG pdb Protein Databank ATAACCATTCCAGAGGCTTATGAGCGTCTAATTCTCGACACAATTAGAGGTGATCAACAACATTTTGTTC GCAGAGACGAATTAAAGGCATCATGGCAAATATTCACACCACTTTTACACAAAATTGATAGAGGGGAGTT pir PIR GAAGCCGGTTCCTTACAACCCGGGAAGTAGAGGTCCTGCAGAAGCAGATGAGTTATTAGAAAAAGCTGGA prf PRF TATGTTCAAACACCCGGTTATATATGGATTCCTCCTACCTTATAGAGTGACCAAATTTCATAATAAAACA ref Ref. Seq AGGATTATCAGGAGCTTATAAGTCTTCAATAAGCTTGTGAAATTTTCGTTATAATCTCTC TCATTTTGGGGTGTATATCAAGCATTTAAGCGCGTGTTTGACACAGTTTGTGTAATAGATTTGGCTCTGA ATGAAAATAAACGGGAATTGTTTCTTTTTGTTTTA FASTA definition line >gi|603218|gb|U 18238. 1|MSU 18238
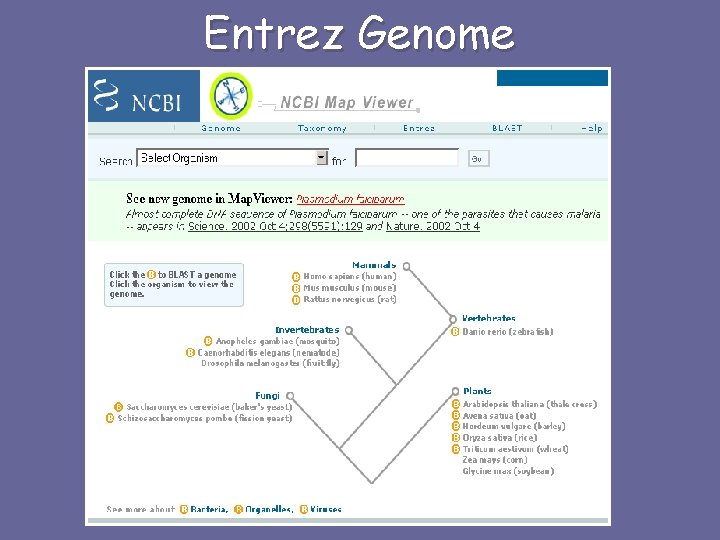
Entrez Genome
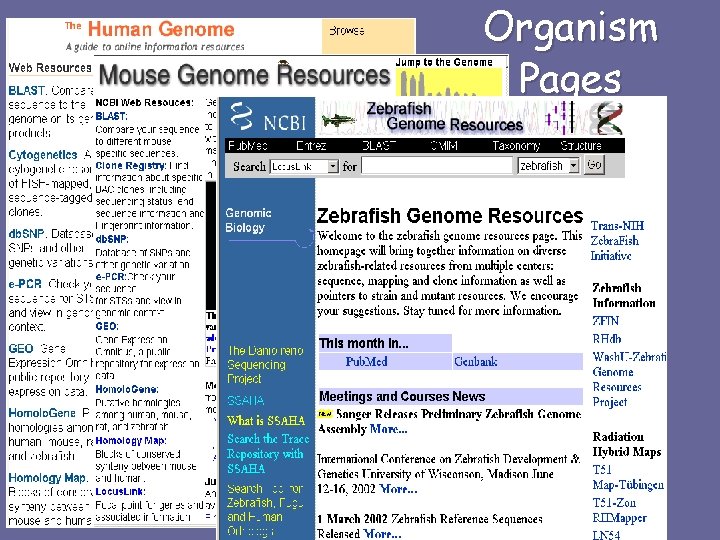
Organism Pages

The Map Viewer: a common platform for integrated display
The Map Viewer

Entrez Pub. Med
Online Books

Entrez Specialized Databases Taxonomy Searchable taxonomic tree having nodes for all species with records in an Entrez database OMIM Online Mendelian Inheritance in Man: A database of genetically linked human diseases Probe. Set Expression data (GEO) and microarray datasets
Entrez Taxonomy
Entrez OMIM

Entrez Probe. Set
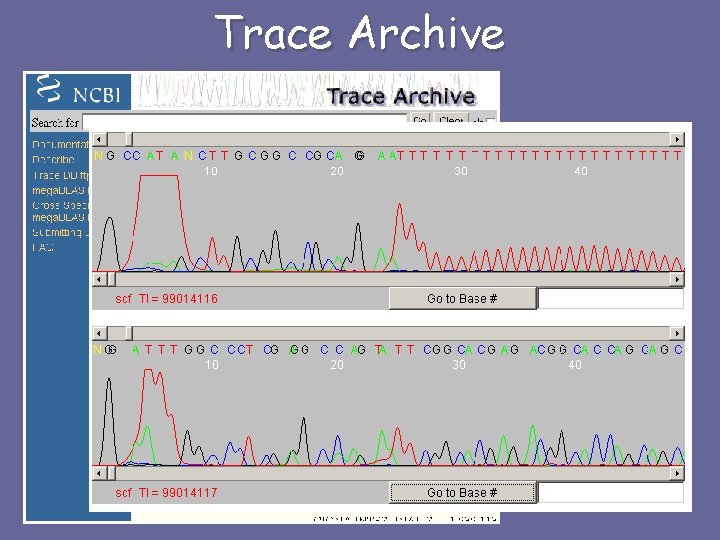
Trace Archive
Entrez Structure

Structure Summary Cn 3 D viewer Related Structures Conserved Domains
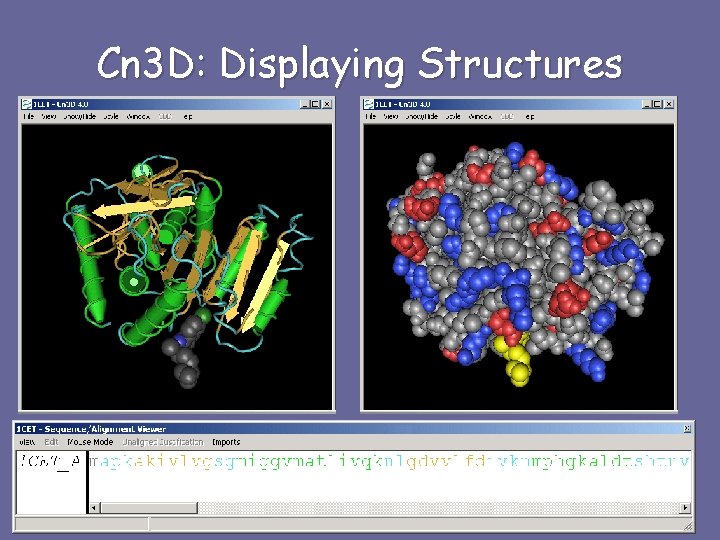
Cn 3 D: Displaying Structures
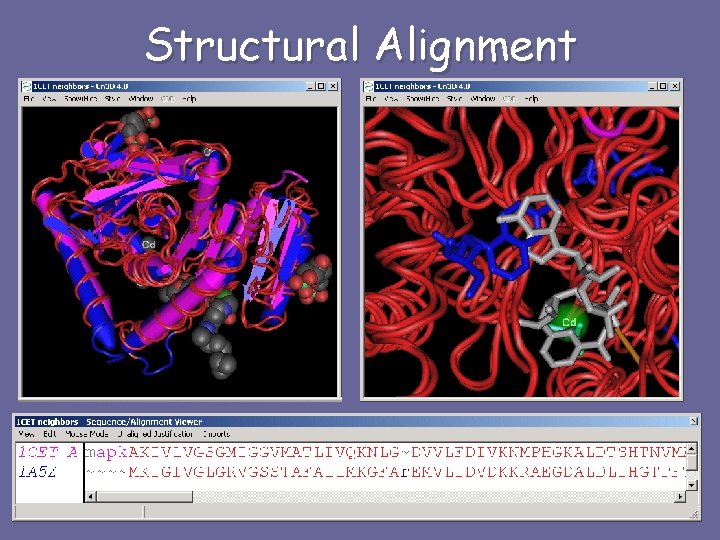
Structural Alignment
- Slides: 56