A Eukaryotic Transcriptional Activator Bearing the DNA Specificity

A Eukaryotic Transcriptional Activator Bearing the DNA Specificity of a Prokaryotic Repressor By Roger Brent and Mark Ptashne Cell (1985) 43: 729 -736 Presented by N. Kuldell and R. Weiss for 20. 382 02. 10

Principles of gene regulation Hypothesize: Tx’n is regulated with modular components Hypothesize: Eukaryotic and prokaryotic systems share common themes for control – – Binding Protein-protein contact to activate Cooperativity Modularity Why we care: enable synthetic control systems
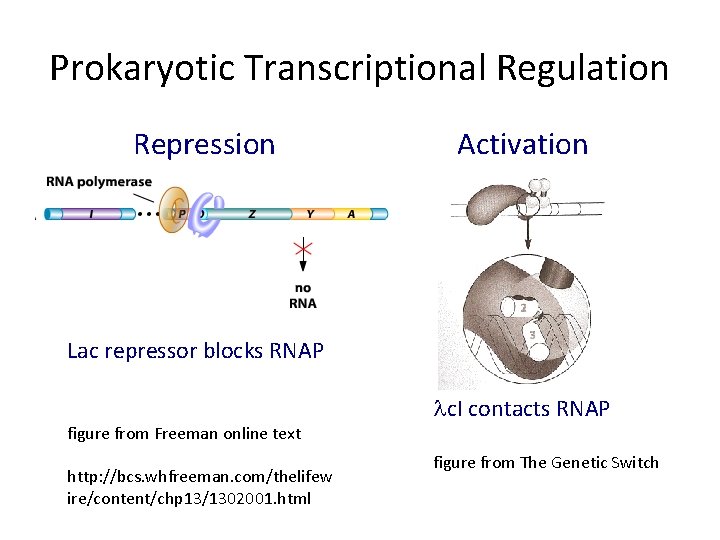
Prokaryotic Transcriptional Regulation Repression Activation Lac repressor blocks RNAP figure from Freeman online text http: //bcs. whfreeman. com/thelifew ire/content/chp 13/1302001. html lc. I contacts RNAP figure from The Genetic Switch
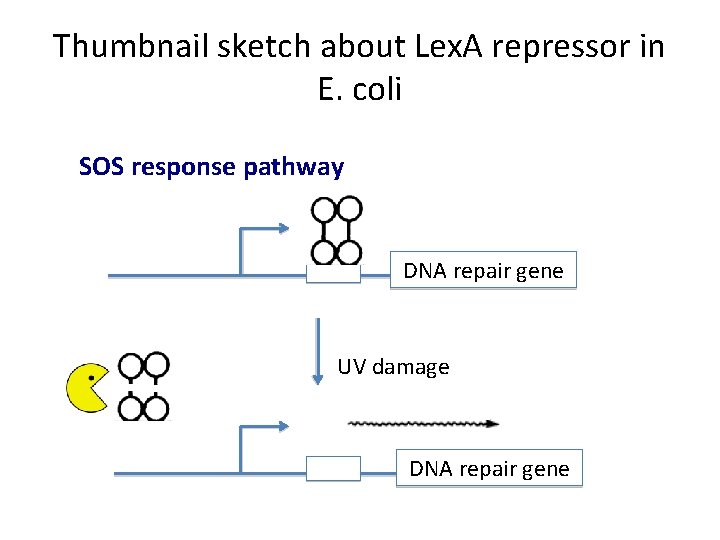
Thumbnail sketch about Lex. A repressor in E. coli SOS response pathway DNA repair gene UV damage DNA repair gene
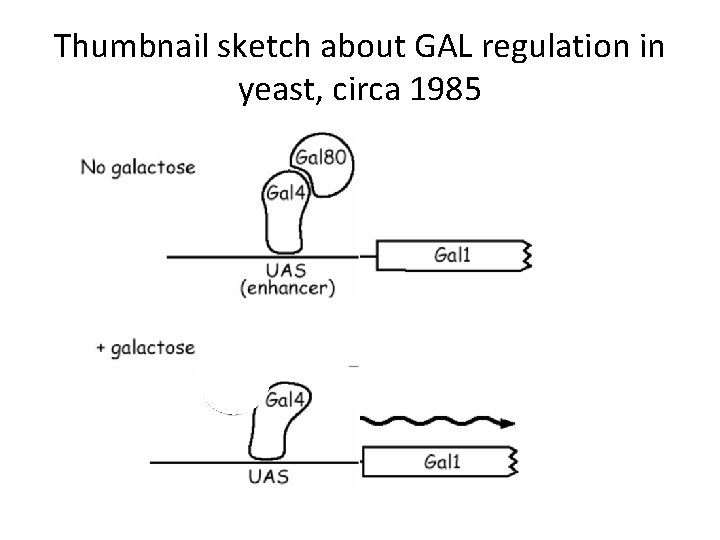
Thumbnail sketch about GAL regulation in yeast, circa 1985
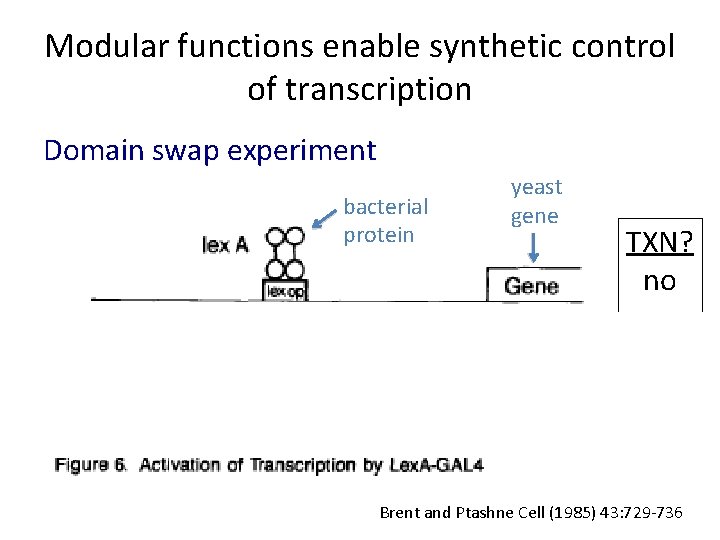
Modular functions enable synthetic control of transcription Domain swap experiment bacterial protein yeast gene TXN? no Brent and Ptashne Cell (1985) 43: 729 -736
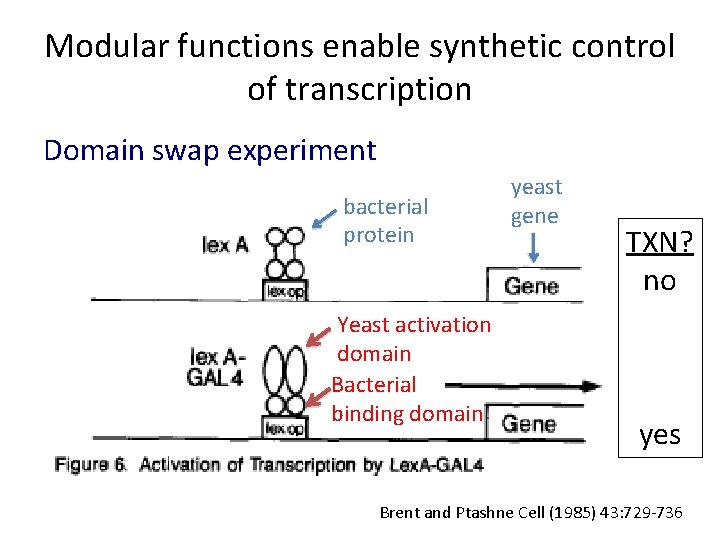
Modular functions enable synthetic control of transcription Domain swap experiment bacterial protein Yeast activation domain Bacterial binding domain yeast gene TXN? no yes Brent and Ptashne Cell (1985) 43: 729 -736

“what if”…eukaryotic activators work like l c. I You’re crazy…. • What about nuclear localization signals? • What about histones? Brent Nature (1984) 312: 612

“what if”…eukaryotic activators work like l c. I You’re crazy…. • What about nuclear localization signals? • What about histones? So knew that bacterial protein could function in eukaryotic nucleus… Brent Nature (1984) 312: 612
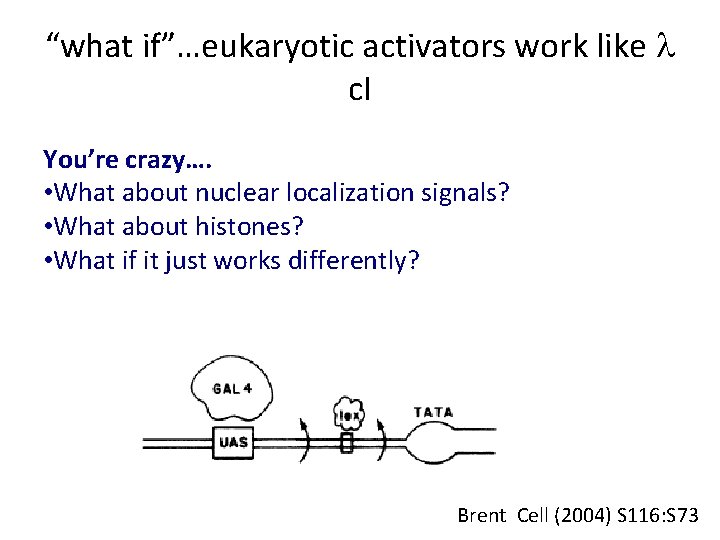
“what if”…eukaryotic activators work like l c. I You’re crazy…. • What about nuclear localization signals? • What about histones? • What if it just works differently? Brent Cell (2004) S 116: S 73
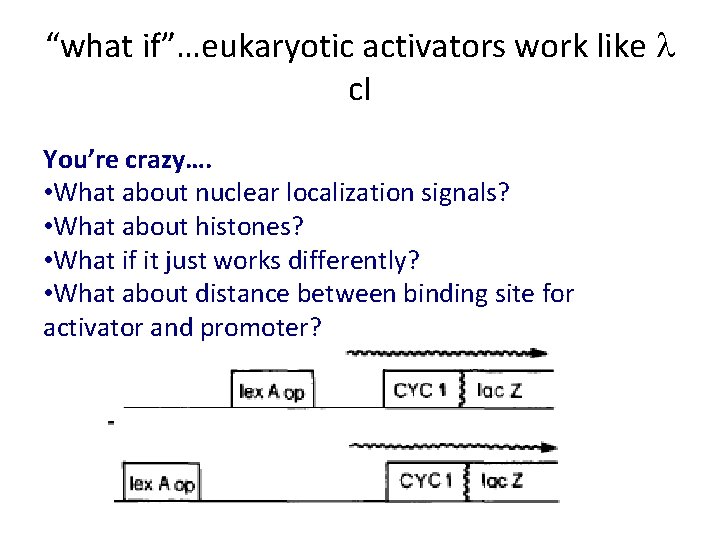
“what if”…eukaryotic activators work like l c. I You’re crazy…. • What about nuclear localization signals? • What about histones? • What if it just works differently? • What about distance between binding site for activator and promoter?

Lex. A-GAL 4 fusion protein construct Brent and Ptashne Cell (1985) 43: 729 -736

Lex. A-GAL 4 works in E. coli Brent and Ptashne Cell (1985) 43: 729 -736
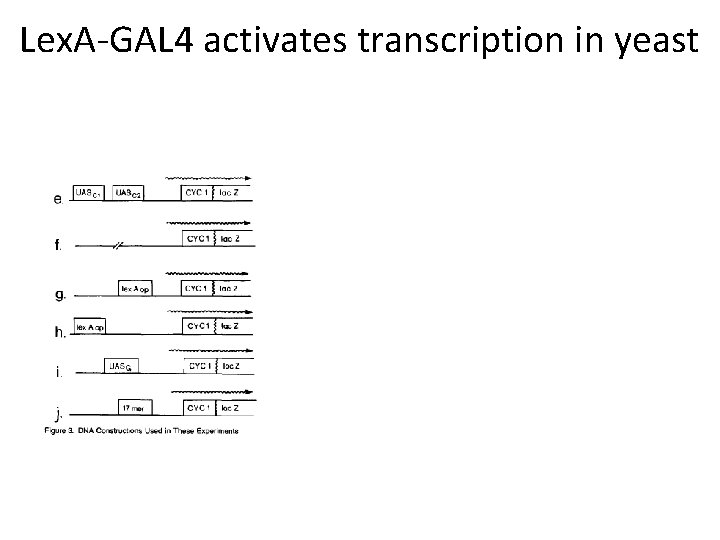
Lex. A-GAL 4 activates transcription in yeast

Lex. A-GAL 4 activates transcription in yeast

Mapping 5’ end of transcript to verify Brent and Ptashne Cell (1985) 43: 729 -736

Squelching by overexpression of GAL 4 Brent and Ptashne Cell (1985) 43: 729 -736

Downstream Activation as well! Figure 5 Brent and Ptashne Cell (1985) 43: 729 -736

Critique Key assumptions • protein functions are modular • eukaryotic/prokaryotic/whatever…. Biggest gaps • footprinting of protein on DNA? • RNAP contact? • nucleosome remodeling? • generalizable?
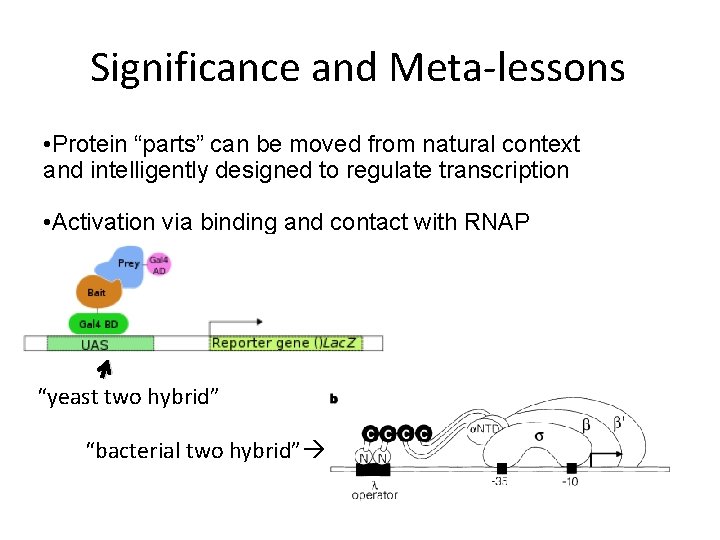
Significance and Meta-lessons • Protein “parts” can be moved from natural context and intelligently designed to regulate transcription • Activation via binding and contact with RNAP “yeast two hybrid” “bacterial two hybrid”
- Slides: 20