A comparative genomic strategy for target discovery and









- Slides: 22

A comparative genomic strategy for target discovery and evolutionary inferences in pathogenic bacteria: Stenotrophomonas sp. as a case study

Stenotrophomonas • Soil and plants are their main reservoirs • Beneficial interactions with plants • Gamma proteobacteria > Xanthomonadaceae • 12 identified species with sequenced genomes

S. maltophilia • Human oportunistic pathogen (colonization rather than infection) • High level of natural antibiotic resistance • Secretion of proteases, lipases, etc • Quorum-sensing regulation • Adherence, motility and biofilm formation • High GC content genome coding for 4. 3 k proteins • High genetic variability and horizontal gene transfer (this makes a troubled taxonomy for this group)

Objectives • Clarify phylogenetic relationships inside Stenotrophomonas genus • Identify shared proteins between clusters of relevant species / strains • Identify S. maltophilia coreproteome • Identify virulence / drug resistance factors

Massive comparative genomics at CDS level Species S. maltophilia S. sp S. acidaminiphila S. ginsengisoli S. humi S. koreensis S. panacihumi S. chelatiphaga S. nitritireducens S. daejeonensis S. pavanii S. terrae S. rhizophila Selected code genomes Sma 43 Ssp 5 Sac 2 Sgi 1 Shu 1 Sko 1 Spa 1 Sch 1 Sni 1 Sda 1 Spv 1 Ste 1 Srh 1 Total = 60 All vs All Proteome Comparison (UCSC Blat)

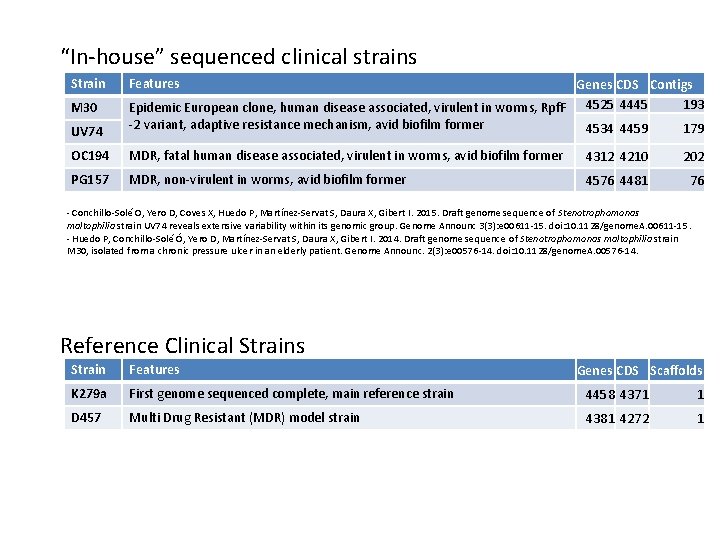
“In-house” sequenced clinical strains Strain Features UV 74 Genes CDS Contigs 193 Epidemic European clone, human disease associated, virulent in worms, Rpf. F 4525 4445 -2 variant, adaptive resistance mechanism, avid biofilm former 4534 4459 179 OC 194 MDR, fatal human disease associated, virulent in worms, avid biofilm former 4312 4210 202 PG 157 MDR, non-virulent in worms, avid biofilm former 4576 4481 76 M 30 - Conchillo-Solé O, Yero D, Coves X, Huedo P, Martínez-Servat S, Daura X, Gibert I. 2015. Draft genome sequence of Stenotrophomonas maltophilia strain UV 74 reveals extensive variability within its genomic group. Genome Announc 3(3): e 00611 -15. doi: 10. 1128/genome. A. 00611 -15. - Huedo P, Conchillo-Solé Ó, Yero D, Martínez-Servat S, Daura X, Gibert I. 2014. Draft genome sequence of Stenotrophomonas maltophilia strain M 30, isolated from a chronic pressure ulcer in an elderly patient. Genome Announc. 2(3): e 00576 -14. doi: 10. 1128/genome. A. 00576 -14. Reference Clinical Strains Strain Features K 279 a First genome sequenced complete, main reference strain 4458 4371 1 D 457 Multi Drug Resistant (MDR) model strain 4381 4272 1 Genes CDS Scaffolds

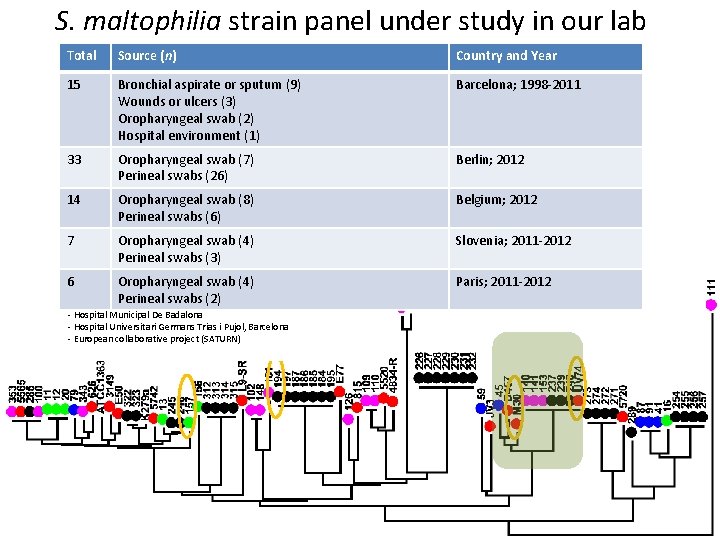
S. maltophilia strain panel under study in our lab Total Source (n) Country and Year 15 Bronchial aspirate or sputum (9) Wounds or ulcers (3) Oropharyngeal swab (2) Hospital environment (1) Barcelona; 1998 -2011 33 Oropharyngeal swab (7) Perineal swabs (26) Berlin; 2012 14 Oropharyngeal swab (8) Perineal swabs (6) Belgium; 2012 7 Oropharyngeal swab (4) Perineal swabs (3) Slovenia; 2011 -2012 6 Oropharyngeal swab (4) Perineal swabs (2) Paris; 2011 -2012 - Hospital Municipal De Badalona - Hospital Universitari Germans Trias i Pujol, Barcelona - European collaborative project (SATURN)

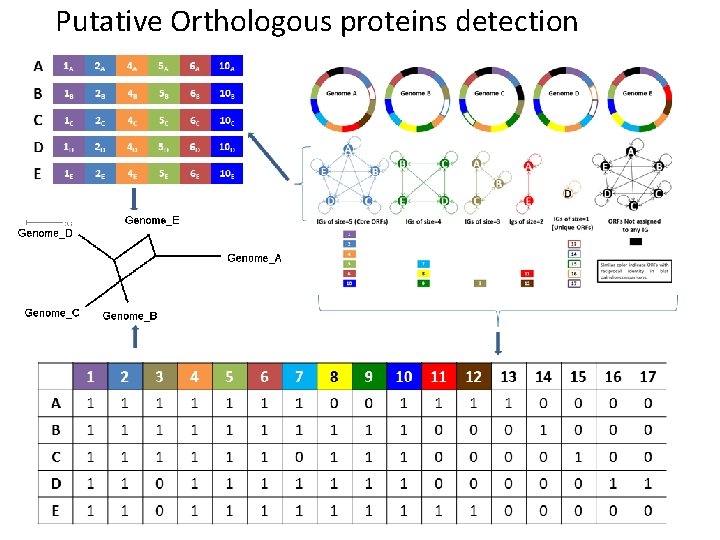
Putative Orthologous proteins detection

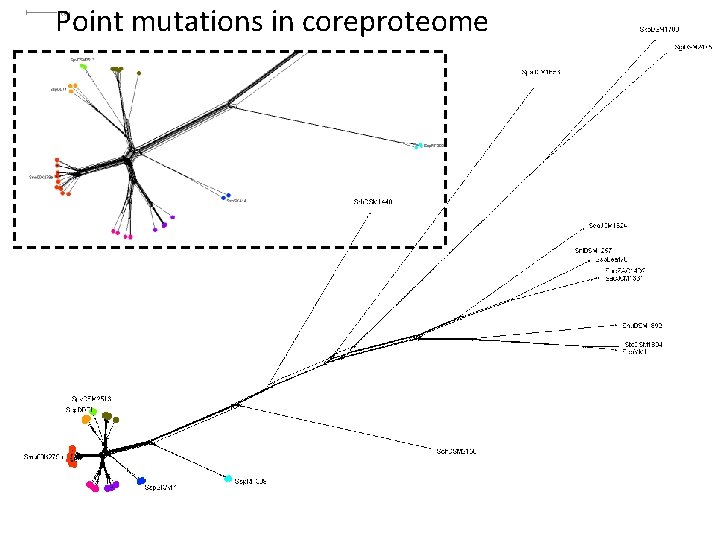
Point mutations in coreproteome

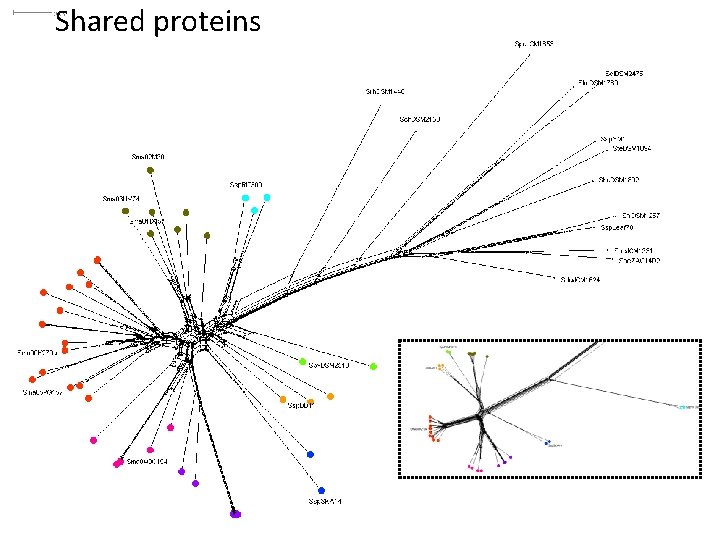
Shared proteins

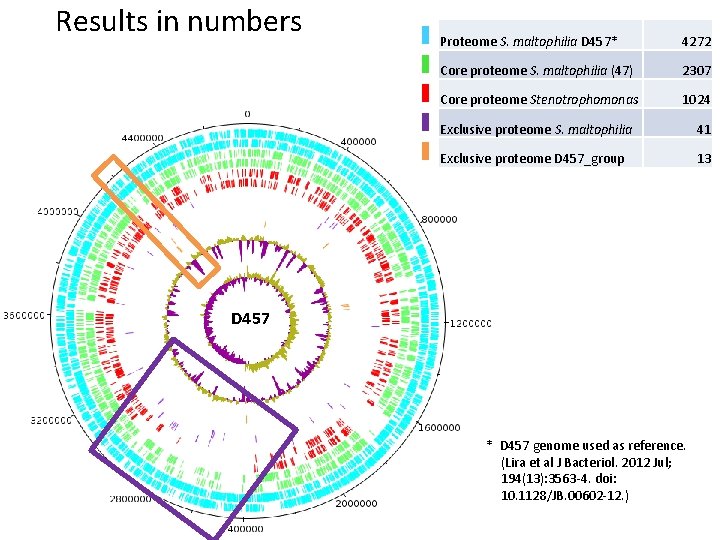
Results in numbers Proteome S. maltophilia D 457* 4272 Core proteome S. maltophilia (47) 2307 Core proteome Stenotrophomonas 1024 Exclusive proteome S. maltophilia 41 Exclusive proteome D 457_group 13 D 457 * D 457 genome used as reference. (Lira et al J Bacteriol. 2012 Jul; 194(13): 3563 -4. doi: 10. 1128/JB. 00602 -12. )

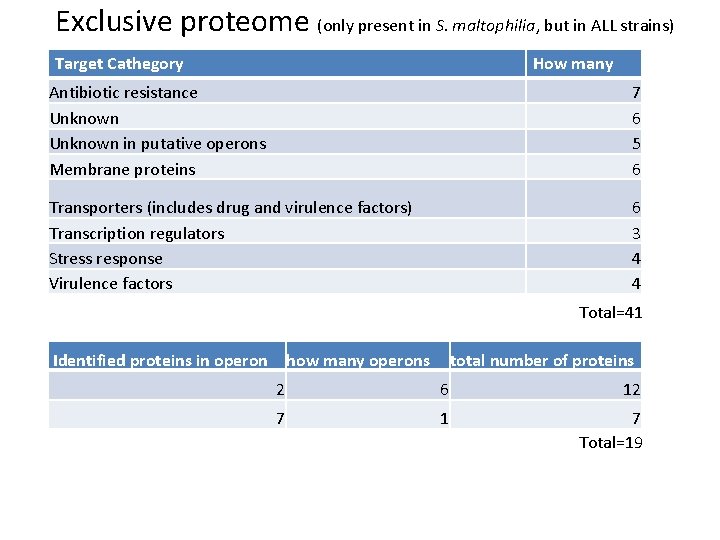
Exclusive proteome (only present in S. maltophilia, but in ALL strains) Target Cathegory How many Antibiotic resistance Unknown in putative operons Membrane proteins 7 6 5 6 Transporters (includes drug and virulence factors) Transcription regulators Stress response Virulence factors 6 3 4 4 Total=41 Identified proteins in operon how many operons total number of proteins 2 6 12 7 1 7 Total=19

S. maltophilia exclusive. Selected Examples Description Membrane protein Transcriptional regulator, XRE family Chaperone Hch. A(Heat shock protein HSP 31) Tet. R family transcriptional regulator Coments Multi Regulator (phague, development, modification proteins, antitoxin, methylases) Next to Lux. R, Stress response Control of multidrug efflux pumps, biosynthesis of antibiotics, osmotic stress, toxic chemicals, catabolic pathways, differentiation processes, pathogenicity Transmembrane protein (Transport protein) Membrane protein Hypothetical exported protein (Membrane protein) Putative lipoprotein (Phosphonate ABC P uptake as virulence factor transporter phosphate-binding periplasmic component) Putative lipoprotein ABC-type antimicrobial peptide transport system. Permease component (Multidrug ABC transporter substrate-binding protein) Zinc-binding protein Metal uptake as a virulence factor Uncharacterized protein (Mer. C mercury Metal uptake as a virulence factor resistance) Azaleucine resistance protein Azl. C, Branched. Peptidic antibiotic resistance chain amino acid ABC transporter permease Putative transmembrane amino acid Peptidic antibiotic resistance transporter protein, Branched-chain amino acid transport Uncharacterized protein (GCN 5 -related NIntervenes in many process including stress acetyltransferase, Spermidine N(1)acetyltransferase) Ara. C family transcriptional regulator, HTH-type Stress and pathogenesis transcriptional activator rha. S

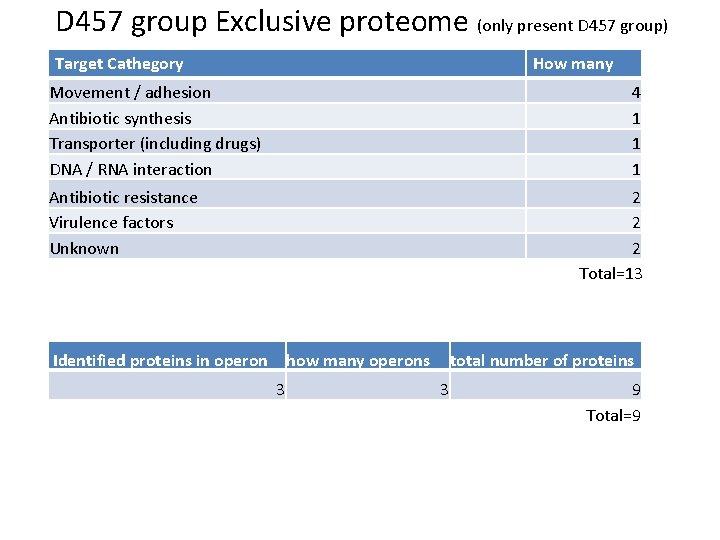
D 457 group Exclusive proteome (only present D 457 group) Target Cathegory How many Movement / adhesion Antibiotic synthesis Transporter (including drugs) DNA / RNA interaction 4 1 1 1 Antibiotic resistance Virulence factors Unknown 2 2 2 Total=13 Identified proteins in operon how many operons 3 total number of proteins 3 9 Total=9

D 457 group (only present D 457 group) Selected Examples Description Comments Uncharacterized protein ( ACR family protein cold shock protein, possible role in PARTIAL_MATCH: Tad. E family protein (pili related) ) extracellular sugar processing Uncharacterized protein (VERY_LOW_SCORE_PARTIAL_MATCH: Flagellar ALL THREE movement / anchor RELATED biosynthesis protein Fli. J) Uncharacterized protein (Lipoprotein, VERY_LOW_SCORE_PARTIAL_MATCH: Beta-propeller ALL THREE movement / anchor RELATED domain-containing protein, ) Uncharacterized protein (MULTIDOMAIN Adhesin, PARTIAL Bacterial Ig-like domain family protein, PARTIAL Adhesion Biofilm-associated ) Membrane translocator, serine/threonine protein Antibiotic sinthesis kinase, Lanthionine synthetase ABC transporter (ABC-type multidrug transport system, ATPase and permease component)

Accomplished Objectives: • Clarify phylogenetic relationships inside Stenotrophomonas genus • Identify shared proteins between clusters of relevant species / strains • Identify S. maltophilia coreproteome • Identify virulence / drug resistance factors Everything done with a single program (and its helper scripts): Fi. Sh. ORF (Find Shared ORF)

Daniel Yero Sònia Martínez Servat Xavi Coves Isidre Gibert Thanks to: Aniol Valera Xavier Daura And everybody else in the group


Exclusive proteome (only present in S. maltophilia, but in ALL strains) Description Coments Uncharacterized protein (Beta-lactamase, Cag pathogenicity island protein Cag 5, Snoa. L-like (biosynthesis of polyketide antibiotics)) Putative esterase, Beta-lactamase (Esterase est. B) Rec Psel yes no no yes Lana protein Uncharacterized methyltransferase Yba. J, Methyltransferase type 11 no no yes Membrane protein no yes Uncharacterized protein no no ABC transporter permease Membrane protein yes yes Transcriptional regulator, XRE family Chaperone protein Hch. A (Heat shock protein HSP 31) Next to Lux. R, Stress response Control of multidrug efflux pumps, biosynthesis of antibiotics, osmotic stress, Tet. R family transcriptional regulator toxic chemicals, catabolic pathways, differentiation processes, pathogenicity Transmembrane protein (Transport protein) no yes no no yes yes Uncharacterized protein Putative transmembrane protein (Membrane protein) no yes no no Multi Regulator (phague, development, modification proteins, antitoxin, methylases)

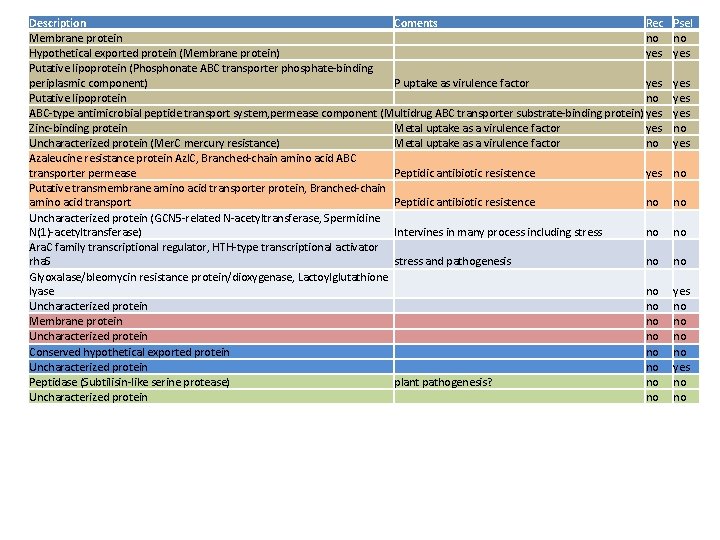
Description Coments Rec Membrane protein no Hypothetical exported protein (Membrane protein) yes Putative lipoprotein (Phosphonate ABC transporter phosphate-binding periplasmic component) P uptake as virulence factor yes Putative lipoprotein no ABC-type antimicrobial peptide transport system, permease component (Multidrug ABC transporter substrate-binding protein) yes Zinc-binding protein Metal uptake as a virulence factor yes Uncharacterized protein (Mer. C mercury resistance) Metal uptake as a virulence factor no Azaleucine resistance protein Azl. C, Branched-chain amino acid ABC transporter permease Peptidic antibiotic resistence yes Putative transmembrane amino acid transporter protein, Branched-chain amino acid transport Peptidic antibiotic resistence no Uncharacterized protein (GCN 5 -related N-acetyltransferase, Spermidine N(1)-acetyltransferase) Intervines in many process including stress no Ara. C family transcriptional regulator, HTH-type transcriptional activator rha. S stress and pathogenesis no Glyoxalase/bleomycin resistance protein/dioxygenase, Lactoylglutathione lyase no Uncharacterized protein no Membrane protein no Uncharacterized protein no Conserved hypothetical exported protein no Uncharacterized protein no Peptidase (Subtilisin-like serine protease) plant pathogenesis? no Uncharacterized protein no Psel no yes yes no no no no yes no no

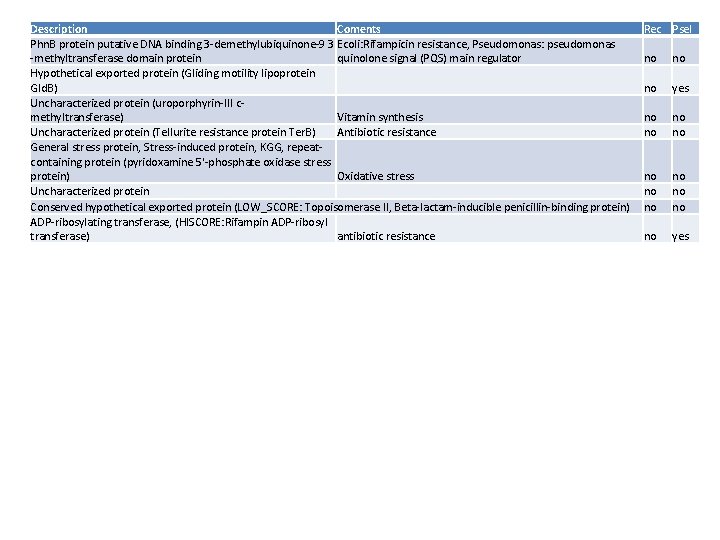
Description Coments Phn. B protein putative DNA binding 3 -demethylubiquinone-9 3 Ecoli: Rifampicin resistance, Pseudomonas: pseudomonas -methyltransferase domain protein quinolone signal (PQS) main regulator Hypothetical exported protein (Gliding motility lipoprotein Gld. B) Uncharacterized protein (uroporphyrin-III cmethyltransferase) Vitamin synthesis Uncharacterized protein (Tellurite resistance protein Ter. B) Antibiotic resistance General stress protein, Stress-induced protein, KGG, repeatcontaining protein (pyridoxamine 5'-phosphate oxidase stress protein) Oxidative stress Uncharacterized protein Conserved hypothetical exported protein (LOW_SCORE: Topoisomerase II, Beta-lactam-inducible penicillin-binding protein) ADP-ribosylating transferase, (HISCORE: Rifampin ADP-ribosyl transferase) antibiotic resistance Rec Psel no no no yes no no no yes

D 457 group Exclusive proteome (only present D 457 group) In: description Uncharacterized protein ( ACR family protein 7 PARTIAL_MATCH: Tad. E family protein (pili related) ) Uncharacterized protein (VERY_LOW_SCORE_PARTIAL_MATCH: Flagellar biosynthesis 7 protein Fli. J) Uncharacterized protein (Lipoprotein, VERY_LOW_SCORE_PARTIAL_MATCH: Beta-propeller domain 7 containing protein, ) Uncharacterized protein (MULTIDOMAIN Adhesin, PARTIAL Bacterial Ig-like domain family protein, PARTIAL Biofilm 7 associated ) Membrane translocator, serine/threonine protein kinase, 7 Lanthionine synthetase Comments cold shock protein, possible role in extracellular sugar processing ALL THREE movement / anchor RELATED UV 74 ALL THREE movement / anchor RELATED Adhesion Antibiotic sinthesis 7 ABC transporter (ABC-type multidrug transport system, ATPase and permease component) 6 superfamily I DNA/RNA helicase protein 5 NAD-dependent / UDP-glucose epimerase Antibiotic resistance (ciclopirox) Uncharacterized protein ( Putative membrane protein, LOWER_SCORE: Putative sugar transferase, TPR repeat-containing 5) virulence Bactoprenol-linked glucose translocase -O antigen modification- 5 (Gtr. A family protein) virulence 4 Uncharacterized protein (Membrane protein) M 30 Antibiotic (Vancomycin) resistance D 457 JV 3