A clustering method for repeat analysis in DNA














- Slides: 19

A clustering method for repeat analysis in DNA sequences Pusan National University Interdisciplinary Program of Bioinformatics Molecular Biology & Phylogeny Laboratory 석사 1년 김우연 2021 -10 -15 1

A clustering method for repeat analysis in DNA sequences Natalia Volfovsky, Brian J Haas and Steven L Salzberg l The Institute for Genomic Research, USA l Genome Biology 2001 l Molecular Biology & Phylogeny Laboratory Pusan Bioinformatics & Biocomplexity Research Center 2

Abstract Molecular Biology & Phylogeny Laboratory Pusan Bioinformatics & Biocomplexity Research Center 3

Suffix Trie l Definition a 123456 b b a # 1 # 5 a 6 a b # # 3 2 Pusan Bioinformatics & Biocomplexity Research Center # 4 Molecular Biology & Phylogeny Laboratory ¨ Tree: 한 개 이상의 node 로 구성된 유한집합 ¨ Suffix: 각 위치에서 시작하는 가장 긴 substring ¨ Suffix tree: 모든 suffix 를 표현하는 trie ¨ 예: T = ababa# 4

Suffix Tree l Definition T = ababa# P = aba 123456 a # ba 6 • Edge : label # ba # • Internal node # 5 • Sibling edge ba# 4 • Leaf node <=> Suffix 1 3 2 Pusan Bioinformatics & Biocomplexity Research Center Molecular Biology & Phylogeny Laboratory ¨ Suffix tree: 모든 suffix 를 표현하는 compacted trie ¨ 예: 5

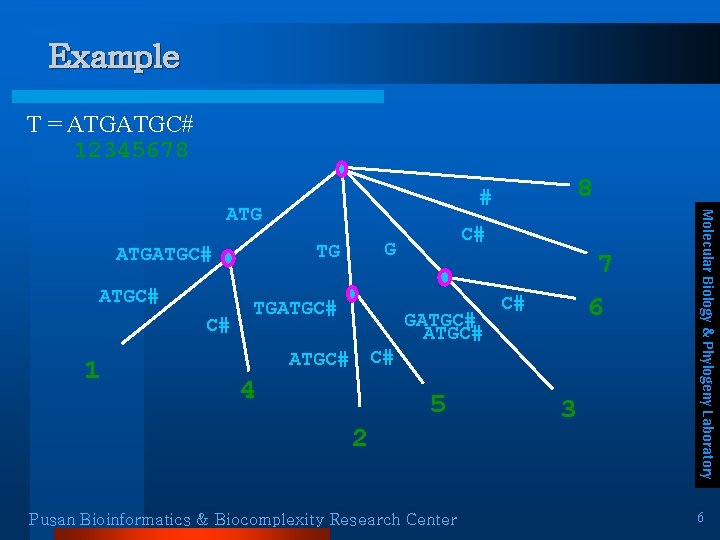
Example T = ATGATGC# 12345678 ATGC# C# 1 C# G TG TGATGC# 7 GATGC# 6 C# C# ATGC# 4 5 2 Pusan Bioinformatics & Biocomplexity Research Center 3 Molecular Biology & Phylogeny Laboratory ATGATGC# 8 # 6

Numerous methods for detecting repeats l Repeat. Masker ¨ Using a database of known repeat sequences and implements a string-matching algorithm Masker. Aid ¨ Same approach ¨ More rapid than Repeat. Masker l WU-BLAST ¨ Using the BLAST engine l Based on suffix trees ¨ Repeat. Match, REPuter, Repeat. Finder ¨ Finding all exact repeats ¨ 10 -100 megabases (Mb) Pusan Bioinformatics & Biocomplexity Research Center Molecular Biology & Phylogeny Laboratory l 7

Definitions l An exact repeat ¨ A subsequence occurring in DNA seqeunce at least twice A maximal repeat ¨ Can’t be extended in either direction without incurring a mismatch Pusan Bioinformatics & Biocomplexity Research Center Molecular Biology & Phylogeny Laboratory l 8

Exact repeats Molecular Biology & Phylogeny Laboratory Pusan Bioinformatics & Biocomplexity Research Center 9

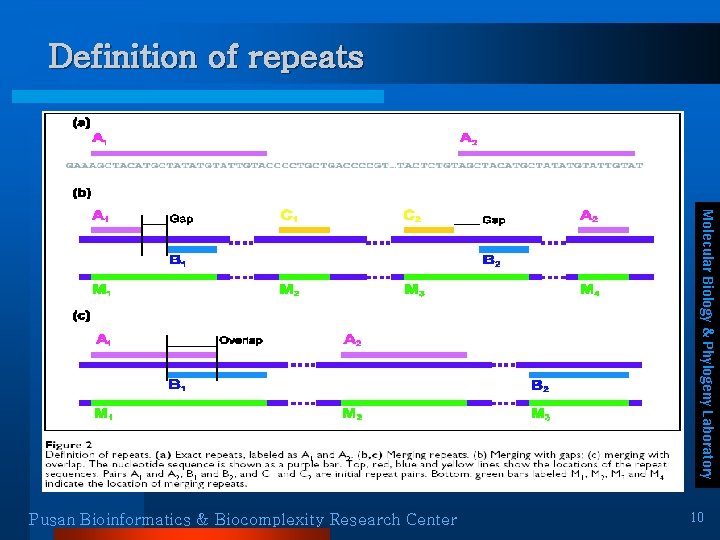
Definition of repeats Molecular Biology & Phylogeny Laboratory Pusan Bioinformatics & Biocomplexity Research Center 10

Algorithm description l Using either of two suffix tree method ¨ Repeat. Match, REPuter Step 1: Selection and pre-processing l Step 2: Merging procedure l Step 3: Classification l Step 4: BLAST searches and repeat class updates l Pusan Bioinformatics & Biocomplexity Research Center Molecular Biology & Phylogeny Laboratory Based on first identifying all exact repeats l Defining repeat classes by merging and extending l 11

STEP 1: Selection and pre-processing Interpreting a partition of the original genome sequence F: forward RC: reverse complement l: length Pusan Bioinformatics & Biocomplexity Research Center Molecular Biology & Phylogeny Laboratory By output of Repeat. Match or REPuter 12

STEP 2: Merging procedure Pusan Bioinformatics & Biocomplexity Research Center Molecular Biology & Phylogeny Laboratory Merging two exact repeats that either overlap or that occur within A limited distance ( a gap ) of each other 13

STEP 3: Classification Pusan Bioinformatics & Biocomplexity Research Center Molecular Biology & Phylogeny Laboratory One step of the classification procedure 14

STEP 4: BLAST searches and further merging Pusan Bioinformatics & Biocomplexity Research Center Molecular Biology & Phylogeny Laboratory If a class appears in multiple similarity pairs, all these similar classes are merged with the original class. 15

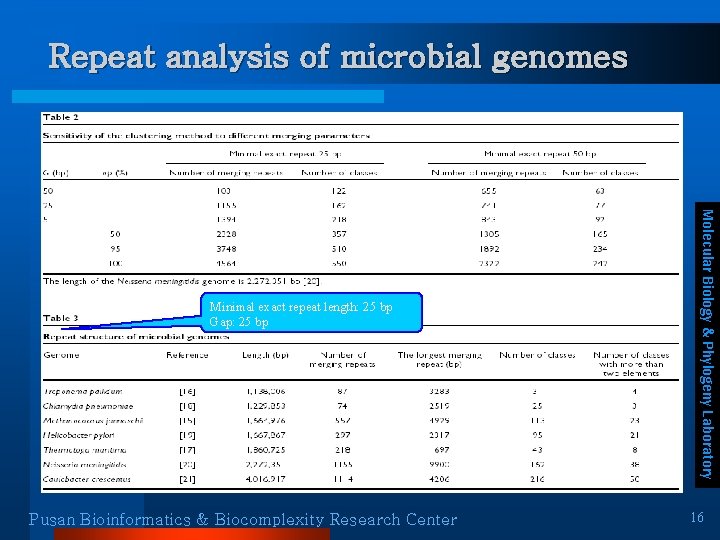
Repeat analysis of microbial genomes Pusan Bioinformatics & Biocomplexity Research Center Molecular Biology & Phylogeny Laboratory Minimal exact repeat length: 25 bp Gap: 25 bp 16

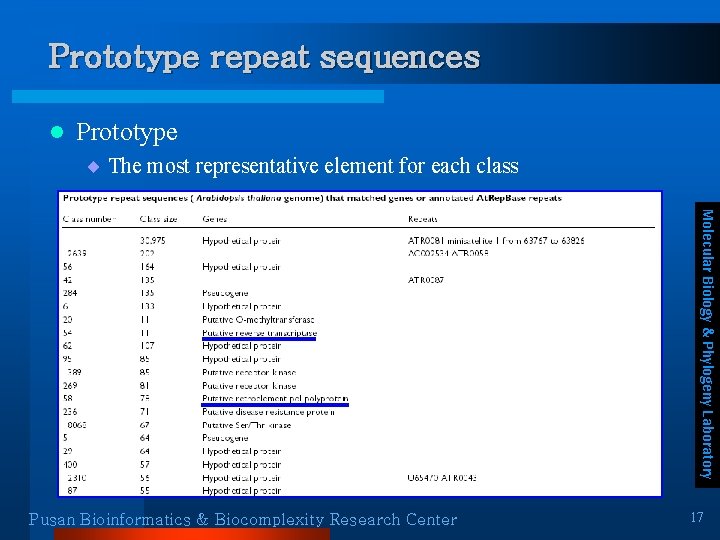
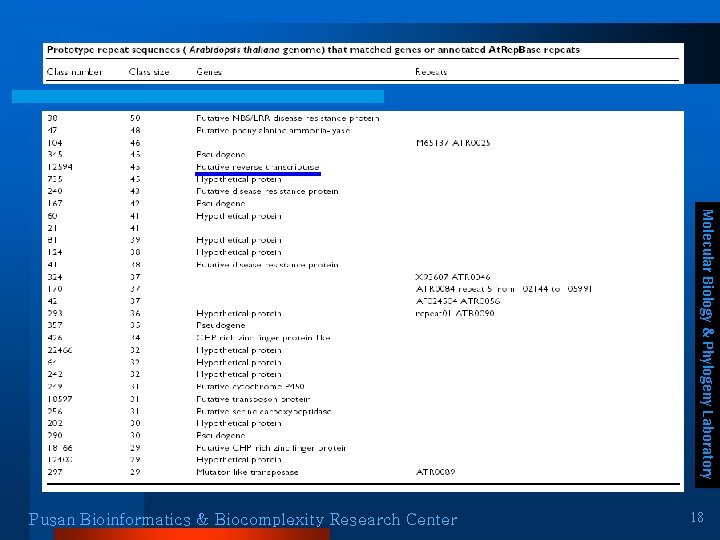
Prototype repeat sequences l Prototype ¨ The most representative element for each class Molecular Biology & Phylogeny Laboratory Pusan Bioinformatics & Biocomplexity Research Center 17

Molecular Biology & Phylogeny Laboratory Pusan Bioinformatics & Biocomplexity Research Center 18

l Finding new HERVs by Suffix Tree Molecular Biology & Phylogeny Laboratory Pusan Bioinformatics & Biocomplexity Research Center 19