A Clustered Particle Swarm Algorithm for Retrieving all



















- Slides: 33

A Clustered Particle Swarm Algorithm for Retrieving all the Local Minima of a function C. Voglis & I. E. Lagaris Computer Science Department University of Ioannina, GREECE

Presentation Outline Global Optimization Problem n Particle Swarm Optimization n Clustering Approach n n Modifying Particle Swarm to form clusters Modifying the affinity matrix Putting the pieces together Determining the number of minima n Identification of the clusters n n Preliminary results – Future research

Global Optimization n The goal is to find the Global minimum inside a bounded domain: n One way to do that, is to find all the local minima and choose among them the global one (or ones). Popular methods of that kind are Multistart, MLSL, TMLSL*, etc. n *M. Ali

Particle Swarm Optimization q Developed in 1995 by James Kennedy and Russ Eberhart. q It was inspired by social behavior of bird flocking or fish schooling. q PSO applies the concept of social interaction to problem solving. q Finds a global optimum.

PSO-Description The method allows the motion of particles to explore the space of interest. n Each particle updates its position in discrete unit time steps. n The velocity is updated by a linear combination of two terms: n The first along the direction pointing to the best position discovered by the particle n The second towards the overall best position. n

PSO - Relations Where: is the position of the ith Swarm’s best position particle at step k is its velocity is the best position visited by the ith particle is the overall best position ever visited is the constriction factor Particle’s best position

PS+Clustering Optimization If the global component is weakened the swarm is expected to form clusters around the minima. n If a bias is added towards the steepest descent direction, this will be accelerated. n Locating the minima then may be tackled, to a large extend, as a Clustering Problem (CP). n However is not a regular CP, since it can benefit from information supplied by the objective function. n

Modified PSO Global component is set to zero. n A component pointing towards the steepest descent direction* is added to accelerate the process. n So the swarm motion is described by: n *A. Ismael F. Vaz, M. G. P. Fernantes

Modified PSO movie

Clustering problem: “Partition a data set into M disjoint subsets containing points with one or more properties in common” n A commonly used property refers to topographical grouping based on distances. n Plethora of Algorithms: n n K-Means, Hierarchical -Single linkage-Quantum. Newtonian clustering.

Global k-means n Minimize the clustering error n It is an incremental procedure using the k-Means algorithm repeatedly Independent of the initialization choice. Has been successfully applied to many problems. n n A. Likas

Global K-Means movie

Spectral Clustering Algorithms that cluster points using eigenvectors of matrices derived from the data n Obtain data representation in the lowdimensional space that can be easily clustered n Variety of methods that use the eigenvectors differently n Useful information can be extracted from the eigenvalues n

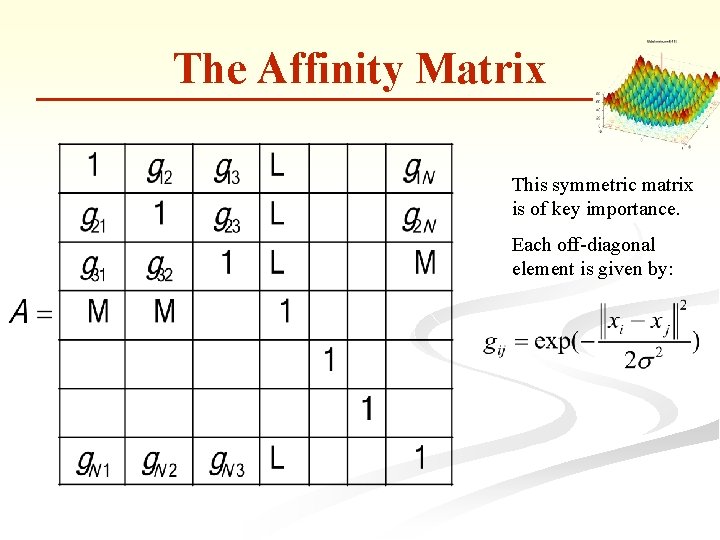
The Affinity Matrix This symmetric matrix is of key importance. Each off-diagonal element is given by:

The Affinity Matrix n Let The Matrix and for is diagonalized and let be its eigenvalues sorted in descending order. The gap which is biggest, identifies the number of clusters (k).

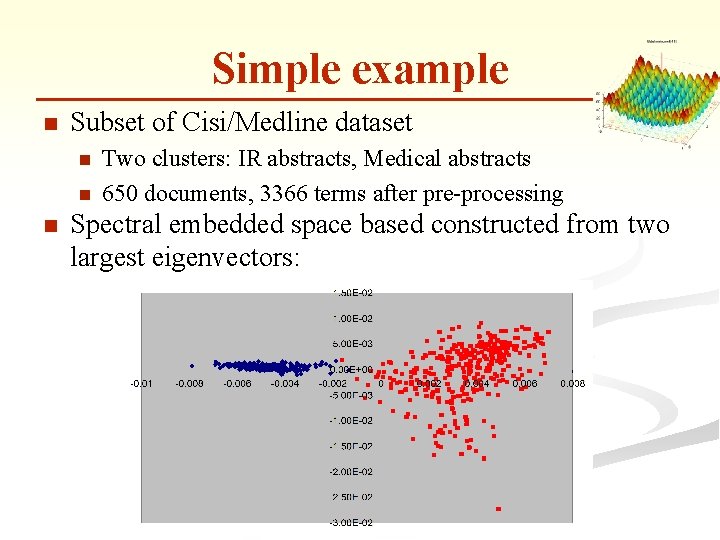
Simple example n Subset of Cisi/Medline dataset n n n Two clusters: IR abstracts, Medical abstracts 650 documents, 3366 terms after pre-processing Spectral embedded space based constructed from two largest eigenvectors:

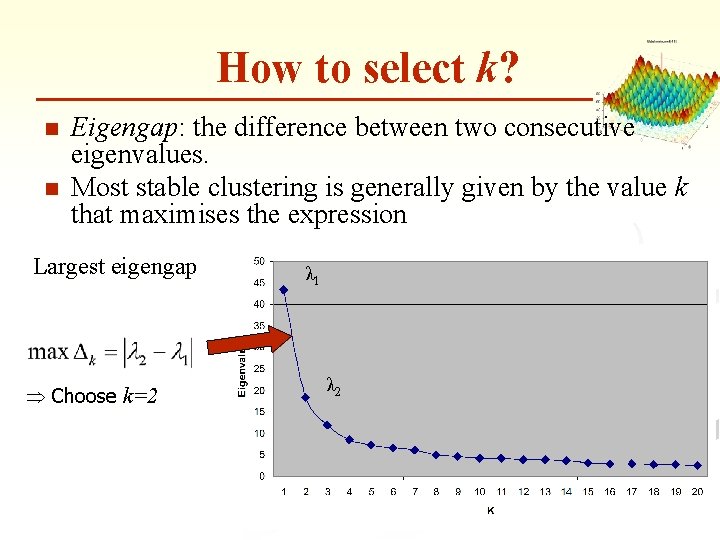
How to select k? n n Eigengap: the difference between two consecutive eigenvalues. Most stable clustering is generally given by the value k that maximises the expression Largest eigengap Þ Choose k=2 λ 1 λ 2

Putting the pieces together 1. 2. Apply modified particle swarm to form clusters around the minima Construct the affinity matrix A and compute the eigenvalues of M. A. B. 3. 4. Use only distance information Add gradient information Find the largest eigengap and identify k. Perform global k-means using the determined k A. B. Use pairwise distances and centroids Use affinity matrix and medoids (with gradient

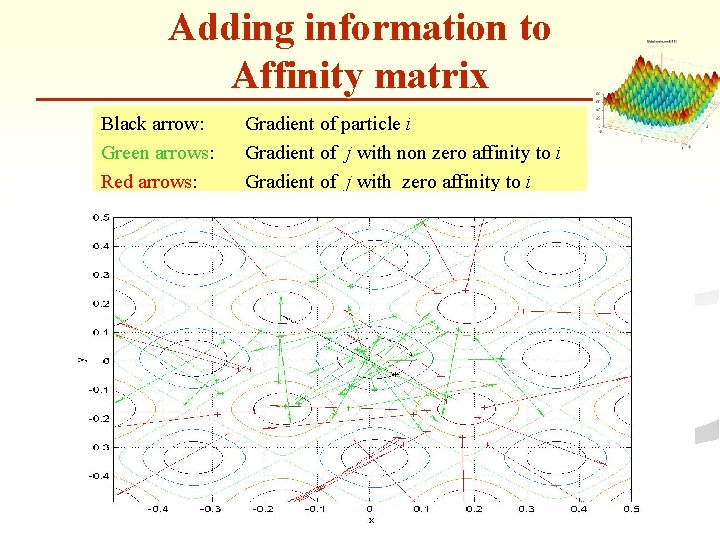
Adding information to Affinity matrix n n n Use the gradient vectors to zero out pairwise affinities. New formula : Do not associate particles that would become more distant if they would follow the negative gradient.

Adding information to Affinity matrix Black arrow: Green arrows: Red arrows: Gradient of particle i Gradient of j with non zero affinity to i Gradient of j with zero affinity to i

From global k-means to global k-medoids n Original global k-means

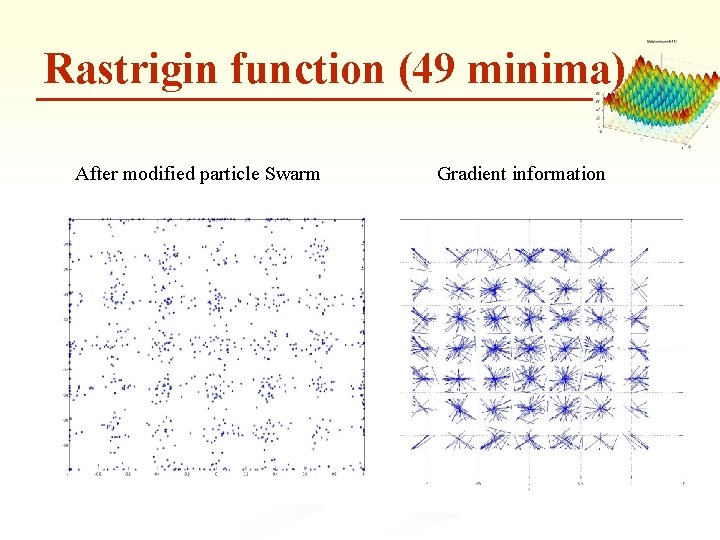
Rastrigin function (49 minima) After modified particle Swarm Gradient information

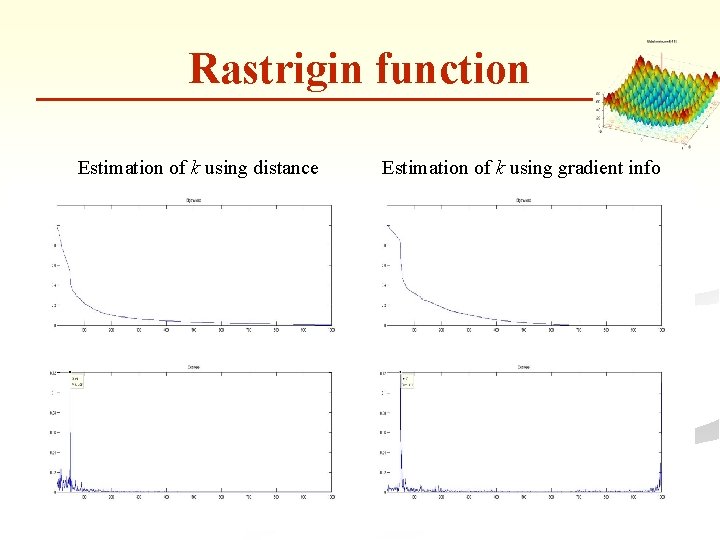
Rastrigin function Estimation of k using distance Estimation of k using gradient info

Rastrigin function Global k-means

Rastrigin function Global k-medoids

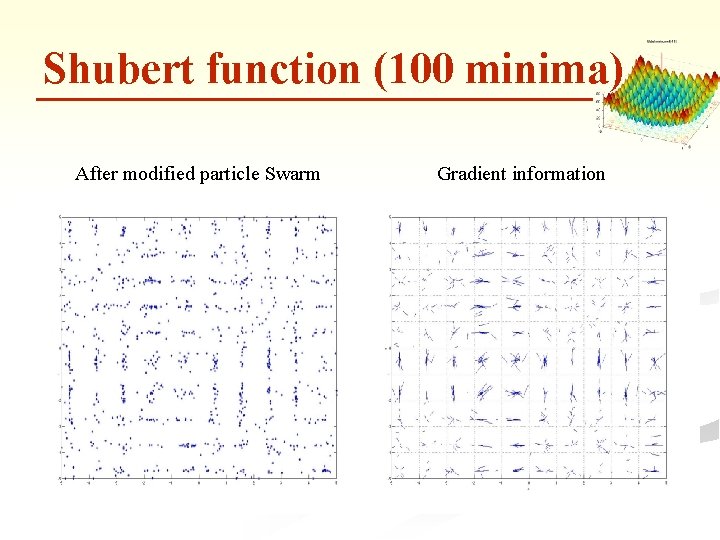
Shubert function (100 minima) After modified particle Swarm Gradient information

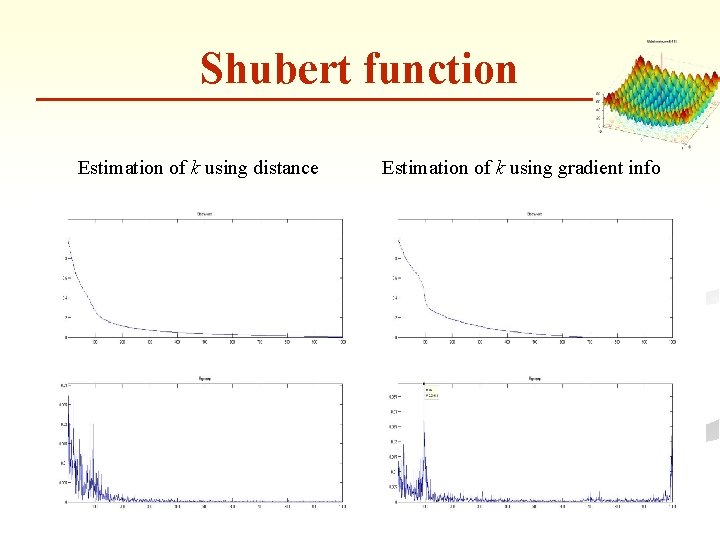
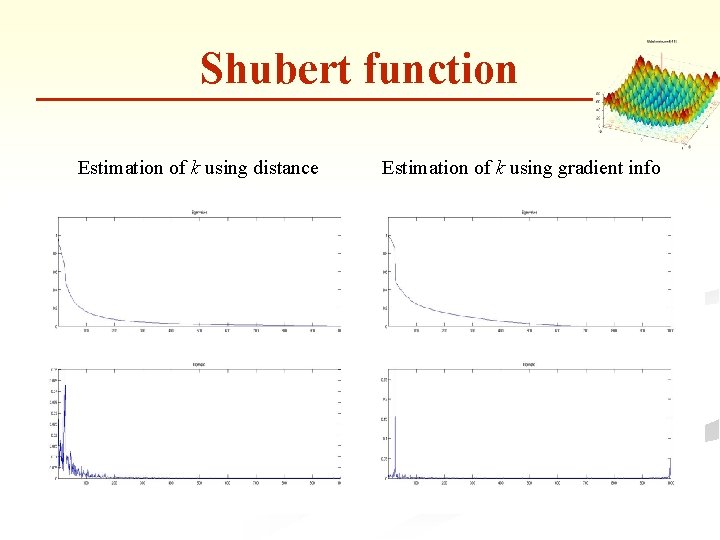
Shubert function Estimation of k using distance Estimation of k using gradient info

Shubert function Global k-means

Shubert function Global k-medoids

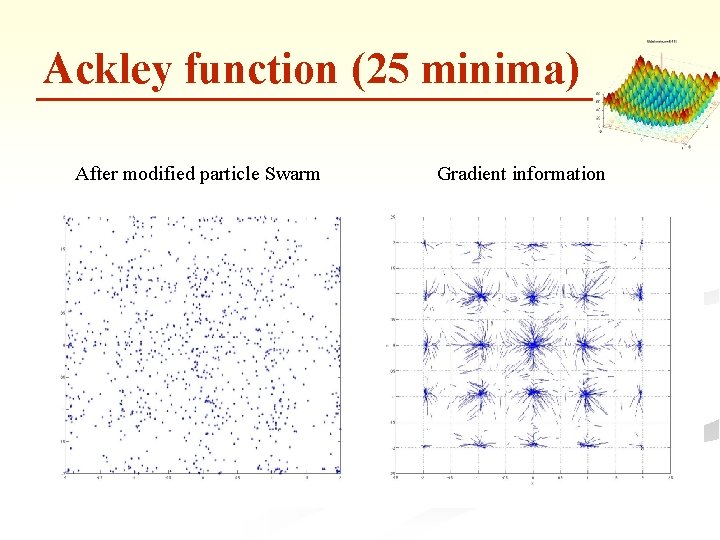
Ackley function (25 minima) After modified particle Swarm Gradient information

Shubert function Estimation of k using distance Estimation of k using gradient info

Shubert function Global k-means

Shubert function Global k-medoids