A clickerbased case study that untangles student thinking

A clicker-based case study that untangles student thinking about the processes in the central dogma. Karen N. Pelletreau 1*, Tessa Andrews 2, Norris Armstrong 2, Mary A. Bedell 2, Farahad Dastoor 1, Neta Dean 3, Susan Erster 3, Cori Fata- Hartley 4, Nancy Guild 5, Hamish Greig 1, David Hall 2, Jennifer K. Knight 5, Donna Koslowsky 4, Paula P. Lemons 6, Jennifer Martin 5, Jill Mc. Court 6, John Merrill 4, Rosa Moscarella 7, Ross Nehm 8, Robert Northington 1, Brian Olsen 1, Luanna Prevost 9, Jon Stoltzfus 10, Mark Urban-Lurain 7, Michelle K. Smith 1 Affiliations: 1 School of Biology and Ecology, University of Maine. 2 Department of Genetics, University of Georgia. 3 Department of Biochemistry and Cell Biology, Stony Brook University. 4 Department of Microbiology and Molecular Genetics, Michigan State University. 5 Department of Molecular, Cellular and Developmental Biology, University of Colorado Boulder. 6 Department of Biochemistry and Molecular Biology, University of Georgia. 7 Collaborative Research in Education, Assessment, and Teaching Environments for the fields of Science, Technology, Engineering and Mathematics (CREATE 4 STEM), Michigan State University. 8 Center for Science and Mathematics Education and Department of Ecology and Evolution, Stony Brook University. 9 Department of Integrative Biology, University of South Florida. 10 Department of Biochemistry and Molecular Biology, Michigan State University. *Correspondence to: School of Biology and Ecology, 5751 Murray Hall, Orono ME 04469. karen. pelletreau@maine. edu 1
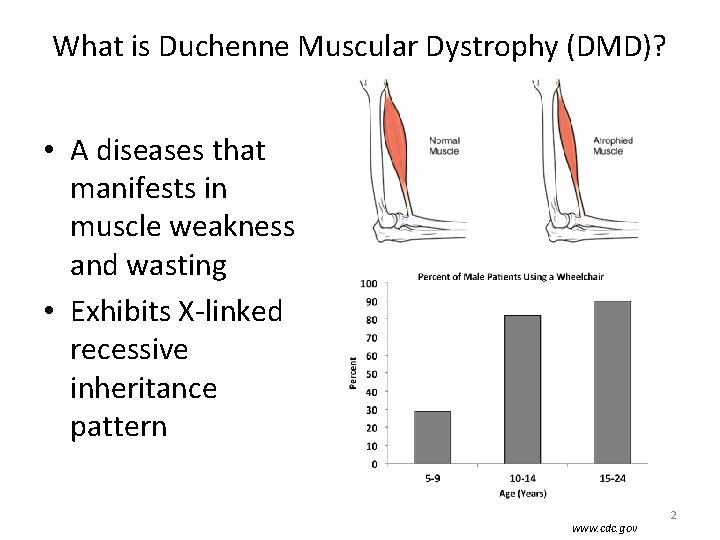
What is Duchenne Muscular Dystrophy (DMD)? • A diseases that manifests in muscle weakness and wasting • Exhibits X-linked recessive inheritance pattern www. cdc. gov 2
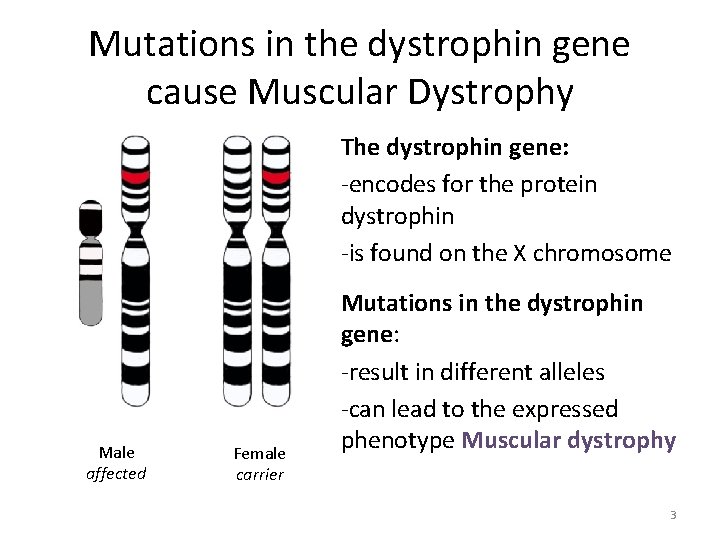
Mutations in the dystrophin gene cause Muscular Dystrophy The dystrophin gene: -encodes for the protein dystrophin -is found on the X chromosome Male affected Female carrier Mutations in the dystrophin gene: -result in different alleles -can lead to the expressed phenotype Muscular dystrophy 3
The dystrophin gene is large and can have many different mutations along the DNA. X chromosome Dystrophin gene 4
Case Study: Siblings living with Duchenne Muscular Dystrophy (DMD) • Liam: affected with DMD • Elijah: unaffected Liam Elijah affected unaffected Your goal: Use the information provided to explore what nucleotide changes could result in Liam developing DMD 5
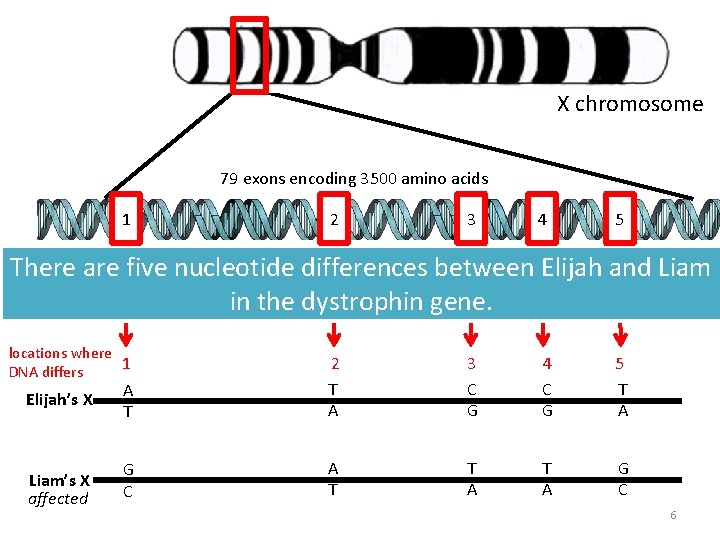
X chromosome 79 exons encoding 3500 amino acids 1 2 3 4 5 There are five nucleotide differences between Elijah and Liam in the dystrophin gene. locations where 1 DNA differs Elijah’s X Liam’s X affected A T 2 T A 3 C G 4 C G 5 T A G C A T T A G C 6
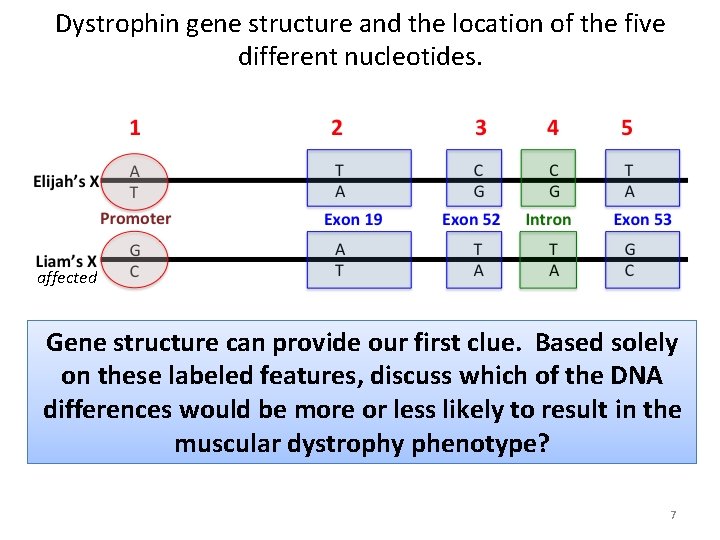
Dystrophin gene structure and the location of the five different nucleotides. affected Gene structure can provide our first clue. Based solely on these labeled features, discuss which of the DNA differences would be more or less likely to result in the muscular dystrophy phenotype? 7
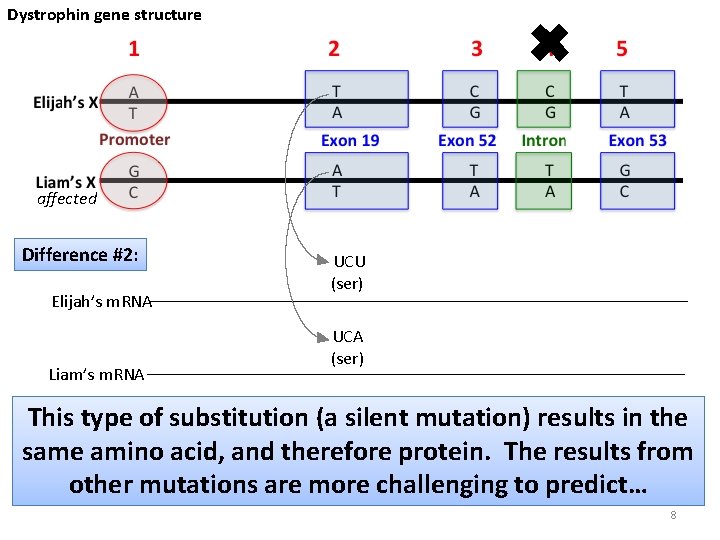
Dystrophin gene structure ✖ affected Difference #2: Elijah’s m. RNA Liam’s m. RNA UCU (ser) UCA (ser) This type of substitution (a silent mutation) results in the same amino acid, and therefore protein. The results from other mutations are more challenging to predict… 8
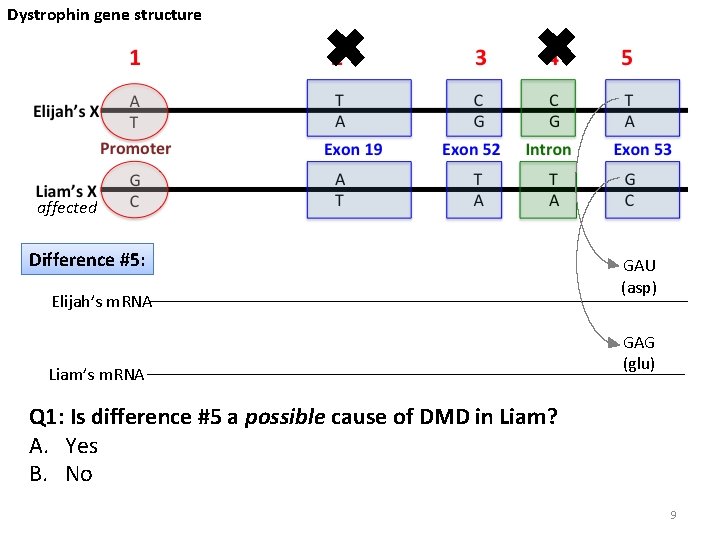
Dystrophin gene structure ✖ ✖ affected Difference #5: Elijah’s m. RNA Liam’s m. RNA GAU (asp) GAG (glu) Q 1: Is difference #5 a possible cause of DMD in Liam? A. Yes B. No 9
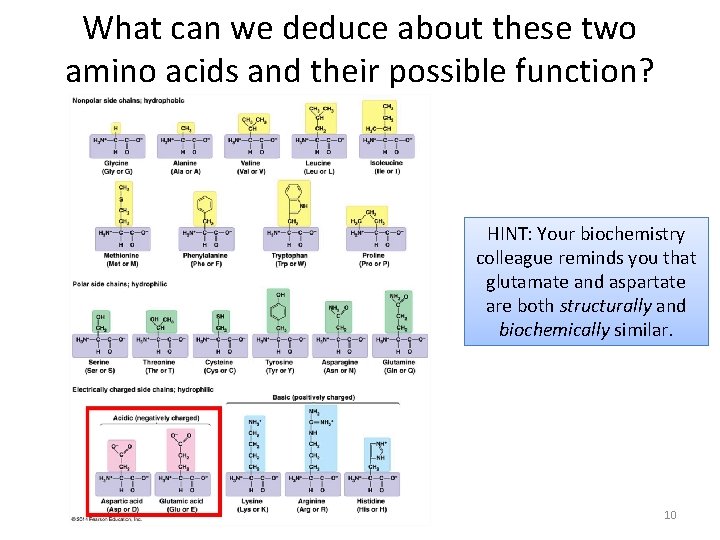
What can we deduce about these two amino acids and their possible function? HINT: Your biochemistry colleague reminds you that glutamate and aspartate are both structurally and biochemically similar. 10
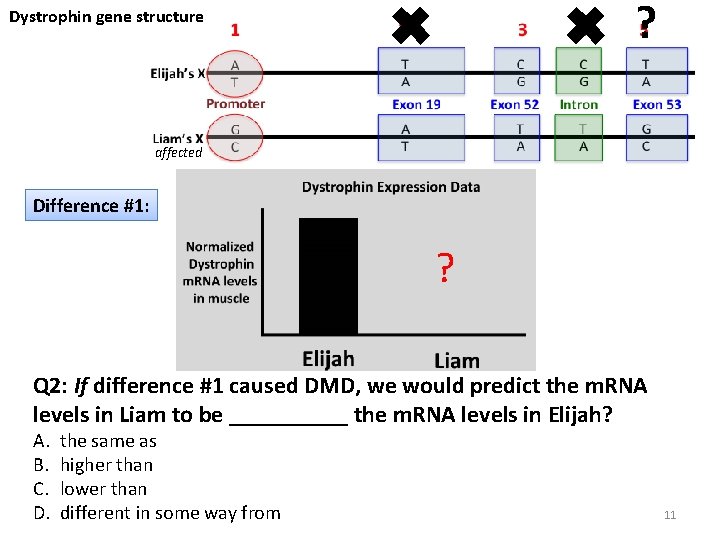
Dystrophin gene structure ✖ ✖ ? affected Difference #1: ? Q 2: If difference #1 caused DMD, we would predict the m. RNA levels in Liam to be _____ the m. RNA levels in Elijah? A. B. C. D. the same as higher than lower than different in some way from 11
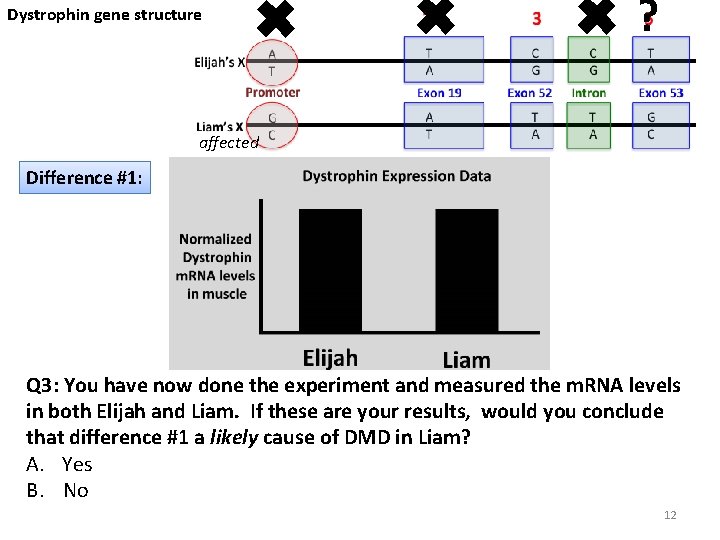
Dystrophin gene structure ✖ ✖ ✖? affected Difference #1: Q 3: You have now done the experiment and measured the m. RNA levels in both Elijah and Liam. If these are your results, would you conclude that difference #1 a likely cause of DMD in Liam? A. Yes B. No 12
13
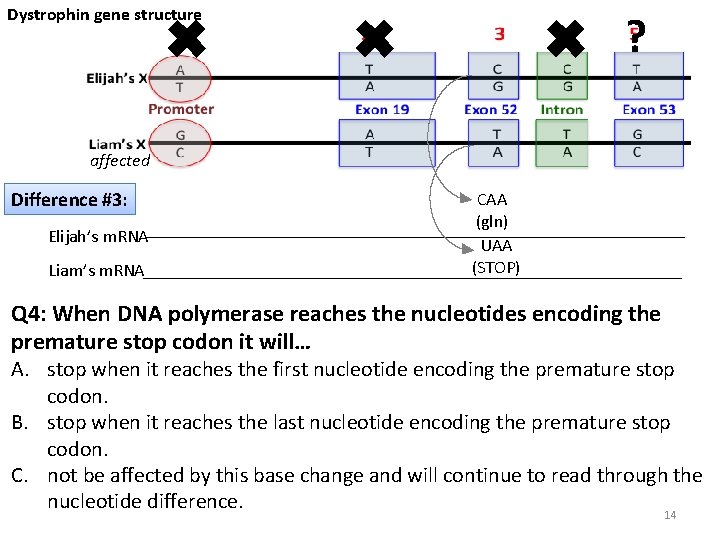
Dystrophin gene structure ✖ ✖ ? ✖ affected Difference #3: Elijah’s m. RNA Liam’s m. RNA CAA (gln) UAA (STOP) Q 4: When DNA polymerase reaches the nucleotides encoding the premature stop codon it will… A. stop when it reaches the first nucleotide encoding the premature stop codon. B. stop when it reaches the last nucleotide encoding the premature stop codon. C. not be affected by this base change and will continue to read through the nucleotide difference. 14
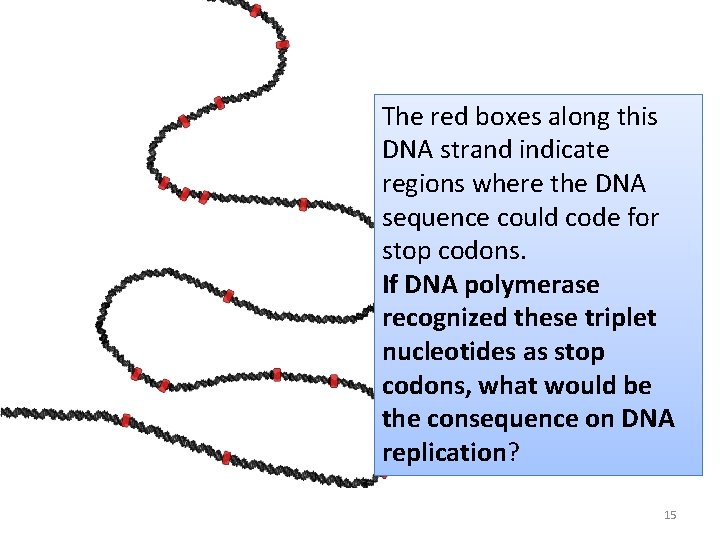
The red boxes along this DNA strand indicate regions where the DNA sequence could code for stop codons. If DNA polymerase recognized these triplet nucleotides as stop codons, what would be the consequence on DNA replication? 15
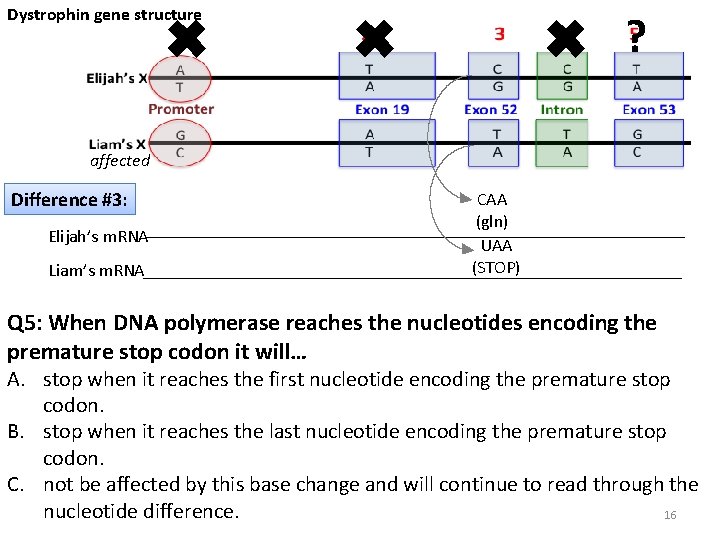
Dystrophin gene structure ✖ ✖ ? ✖ affected Difference #3: Elijah’s m. RNA Liam’s m. RNA CAA (gln) UAA (STOP) Q 5: When DNA polymerase reaches the nucleotides encoding the premature stop codon it will… A. stop when it reaches the first nucleotide encoding the premature stop codon. B. stop when it reaches the last nucleotide encoding the premature stop codon. C. not be affected by this base change and will continue to read through the nucleotide difference. 16

Polymerases read only one base at a time, not triplet codons. The DNA polymerase would not be affected by this base change in Liam’s DNA and replication would proceed normally. 17
NO EFFECT ? ? 18
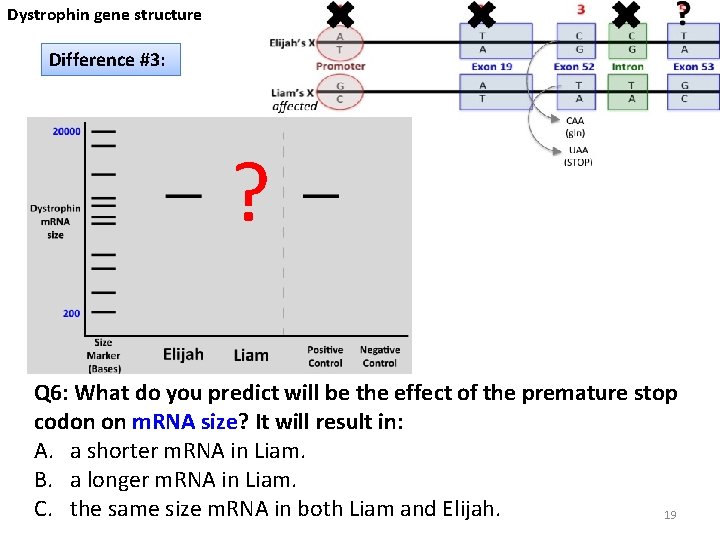
Dystrophin gene structure Difference #3: ? Q 6: What do you predict will be the effect of the premature stop codon on m. RNA size? It will result in: A. a shorter m. RNA in Liam. B. a longer m. RNA in Liam. C. the same size m. RNA in both Liam and Elijah. 19

Liam’s DNA (affected) coding strand template strand Q 7: The above DNA sequence is being transcribed by an RNA polymerase (red square). The premature stop codon mutation in Liam’s DNA is indicated in blue. What will the RNA polymerase do when it reaches the nucleotides encoding the premature stop codon? It will: A. stop when it reaches the first nucleotide encoding the premature stop codon. B. stop when it reaches the last nucleotide encoding the premature stop codon. C. not be affected by this base change and will continue to read through nucleotide difference. 20
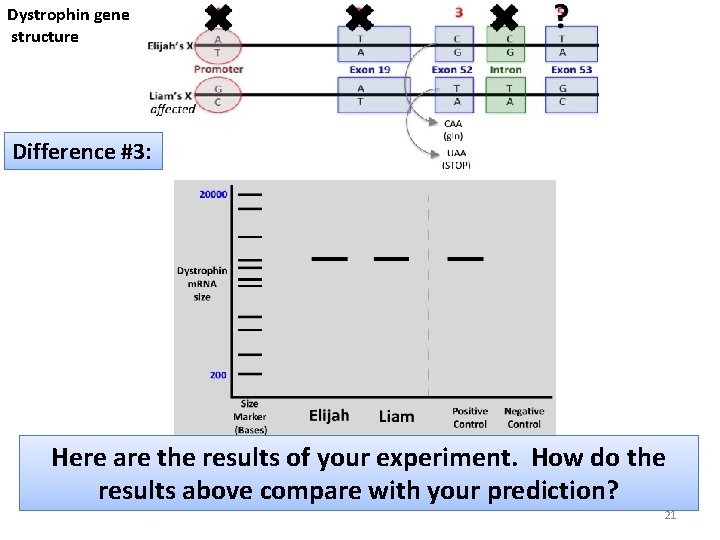
Dystrophin gene structure Difference #3: Here are the results of your experiment. How do the results above compare with your prediction? 21
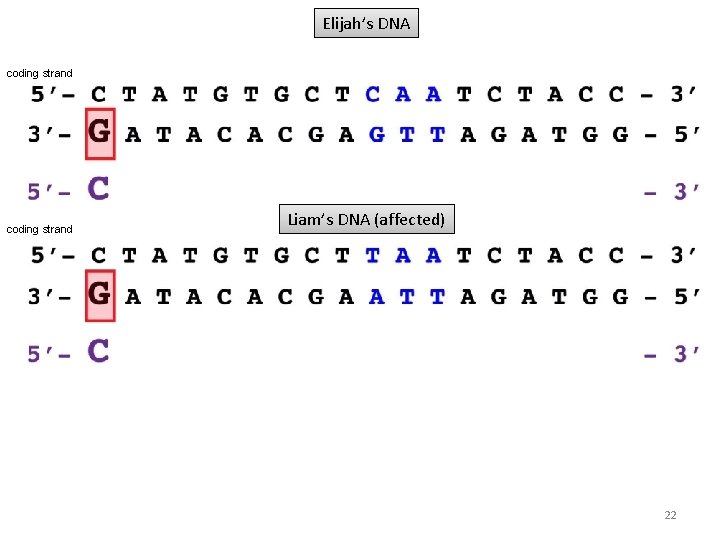
Elijah’s DNA coding strand Liam’s DNA (affected) 22

Elijah’s DNA Liam’s DNA (affected) Polymerases read only one base at a time, not triplet codons. The RNA polymerase would not be affected by this base change in Liam’s DNA and transcription would proceed normally. 23
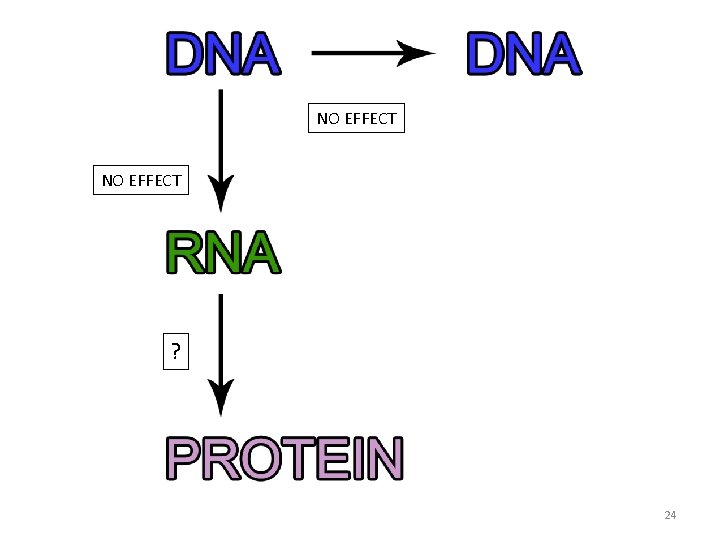
NO EFFECT ? 24
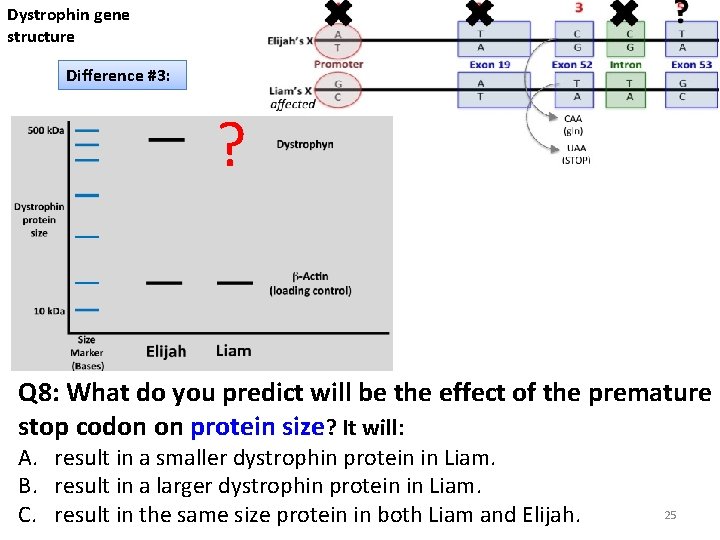
Dystrophin gene structure Difference #3: ? Q 8: What do you predict will be the effect of the premature stop codon on protein size? It will: A. result in a smaller dystrophin protein in Liam. B. result in a larger dystrophin protein in Liam. C. result in the same size protein in both Liam and Elijah. 25
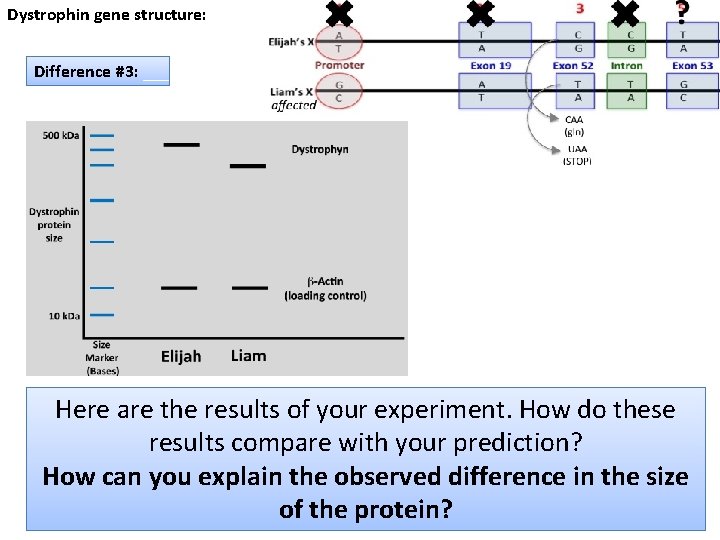
Dystrophin gene structure: Difference #3: Here are the results of your experiment. How do these results compare with your prediction? How can you explain the observed difference in the size of the protein? 26
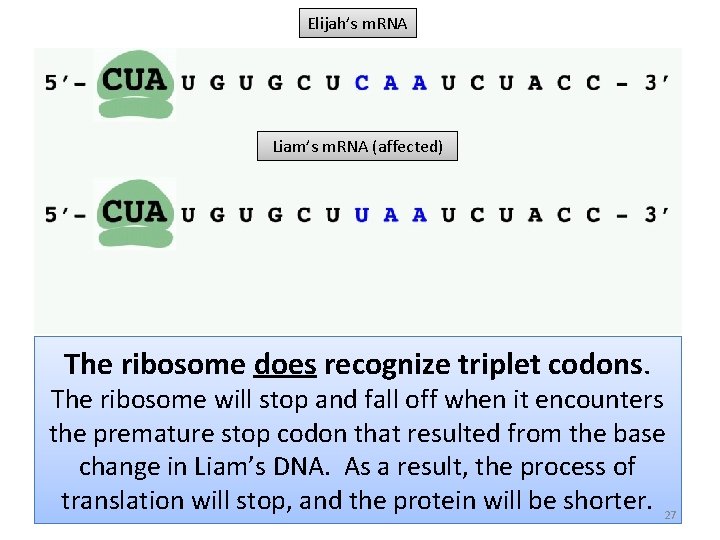
Elijah’s m. RNA Liam’s m. RNA (affected) The ribosome does recognize triplet codons. The ribosome will stop and fall off when it encounters the premature stop codon that resulted from the base change in Liam’s DNA. As a result, the process of translation will stop, and the protein will be shorter. 27
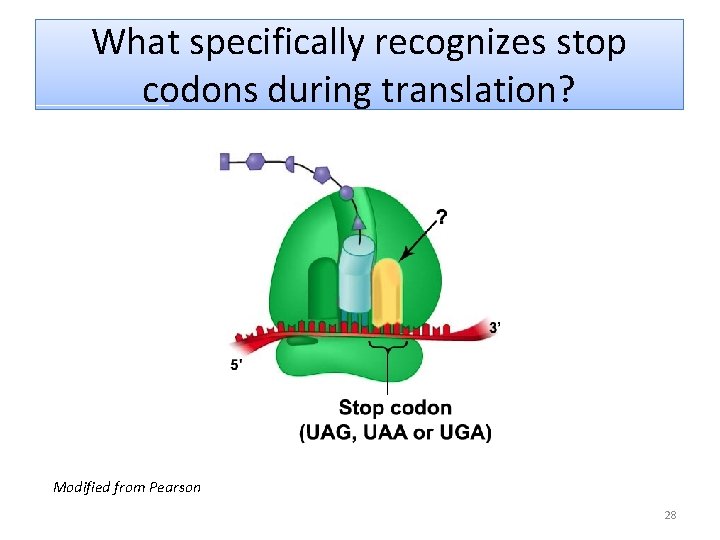
What specifically recognizes stop codons during translation? Modified from Pearson 28
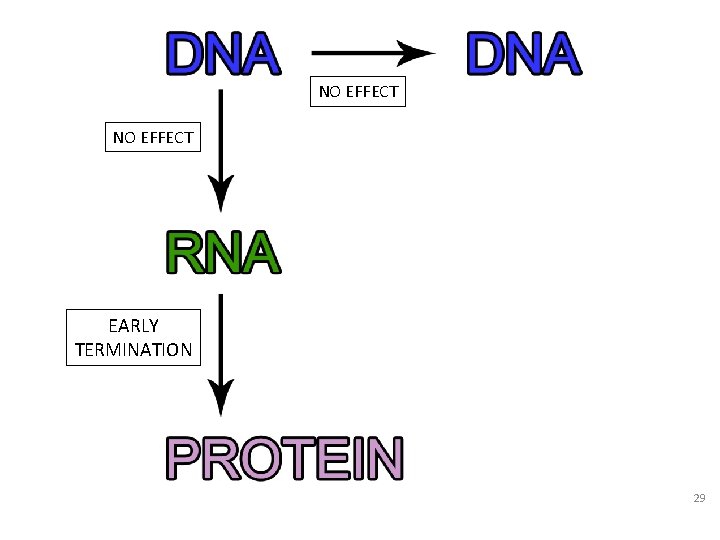
NO EFFECT EARLY TERMINATION 29
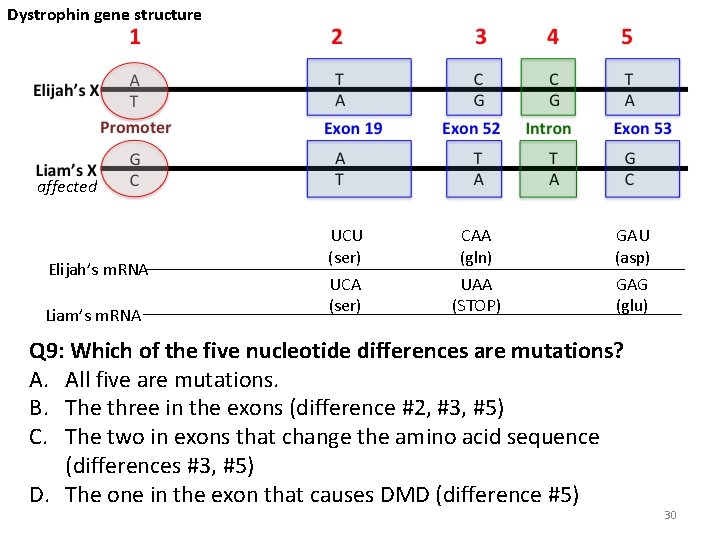
Dystrophin gene structure affected Elijah’s m. RNA Liam’s m. RNA UCU (ser) UCA (ser) CAA (gln) UAA (STOP) GAU (asp) GAG (glu) Q 9: Which of the five nucleotide differences are mutations? A. All five are mutations. B. The three in the exons (difference #2, #3, #5) C. The two in exons that change the amino acid sequence (differences #3, #5) D. The one in the exon that causes DMD (difference #5) 30
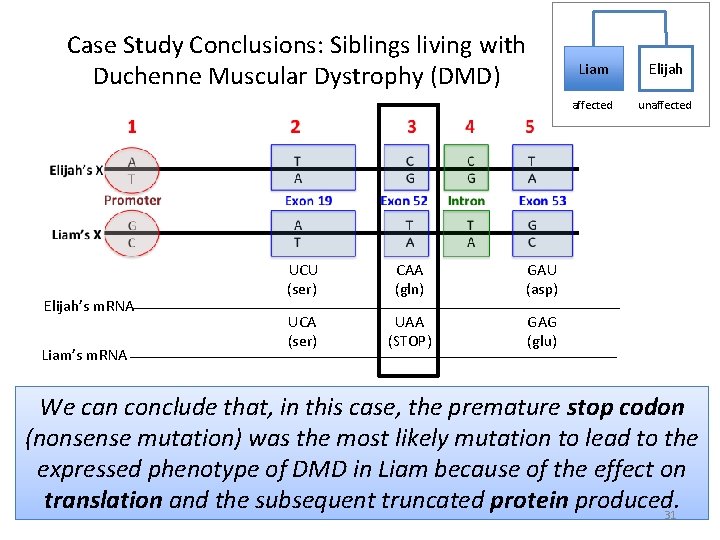
Case Study Conclusions: Siblings living with Duchenne Muscular Dystrophy (DMD) Elijah’s m. RNA Liam’s m. RNA UCU (ser) CAA (gln) GAU (asp) UCA (ser) UAA (STOP) GAG (glu) Liam Elijah affected unaffected We can conclude that, in this case, the premature stop codon (nonsense mutation) was the most likely mutation to lead to the expressed phenotype of DMD in Liam because of the effect on translation and the subsequent truncated protein produced. 31
- Slides: 31