A brief introduction to nextgeneration DNA sequencing data

A brief introduction to nextgeneration DNA sequencing data processing methods Mark A. De. Pristo Manager, Medical and Population Genetics Analysis Medical and Population Genetics Program Broad Institute of Harvard and MIT March 28, 2010
Acknowledgments 2 Genome sequencing and analysis group (MPG) Kiran Garimella (Analysis Lead) Andrew Kernytsky Michael Melgar Chris Hartl Sherman Jia Eric Banks (Development lead) Ryan Poplin Matt Hanna Aaron Mc. Kenna Broad postdocs, staff, and faculty Anthony Philippakis Vineeta Agarwala Manny Rivas Jared Maguire Carrie Sougnez David Jaffe Nick Patterson Steve Schaffner Shamil Sunyaev Paul de Bakker 1000 Genomes project In general but notably: Matt Hurles Philip Awadalla Richard Durbin Goncalo Abecasis Richard Gibbs Gabor Marth Thomas Keane Gil Mc. Vean Gerton Lunter Heng Li Genome Sequencing Platform In general but notably: Lauren Ambrogio Illumina Production Team Tim Fennell Kathleen Tibbetts Alec Wysoker Toby Bloom Copy number group Bob Handsaker Jim Nemesh Josh Korn Steve Mc. Carroll Cancer genome analysis Kristian Cibulskis Andrey Sivachenko Gad Getz Integrative Genomics Viewer (IGV) Jim Robinson Helga Thorvaldsdottir MPG directorship Stacey Gabriel David Altshuler Mark Daly
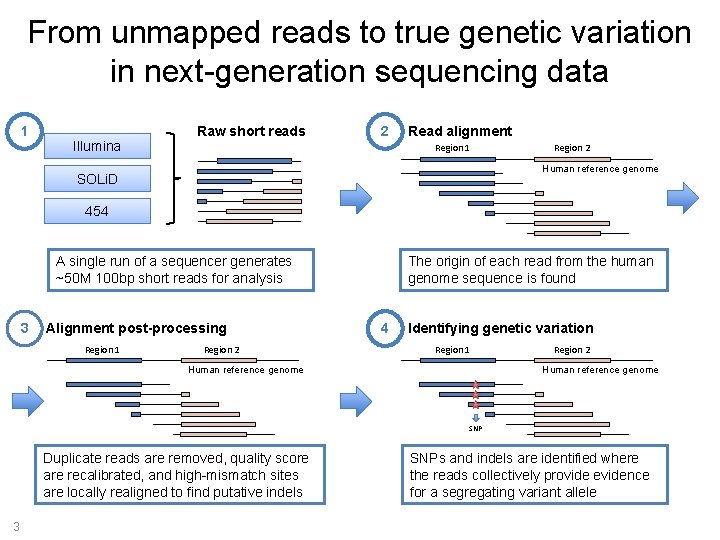
From unmapped reads to true genetic variation in next-generation sequencing data 1 Illumina Raw short reads 2 Read alignment Region 1 Region 2 Human reference genome SOLi. D 454 A single run of a sequencer generates ~50 M 100 bp short reads for analysis 3 Alignment post-processing Region 1 Region 2 The origin of each read from the human genome sequence is found 4 Identifying genetic variation Region 1 Human reference genome Region 2 Human reference genome SNP Duplicate reads are removed, quality score are recalibrated, and high-mismatch sites are locally realigned to find putative indels 3 SNPs and indels are identified where the reads collectively provide evidence for a segregating variant allele

A heterozygous haplotype with two synonymous variants in exon 1 of Notch 2 Coverage Reads at locus Individual reads Non-reference bases colored Reference bases in grey Ref. Seq 4 This is a screenshot of IGV: http: //www. broadinstitute. org/igv/
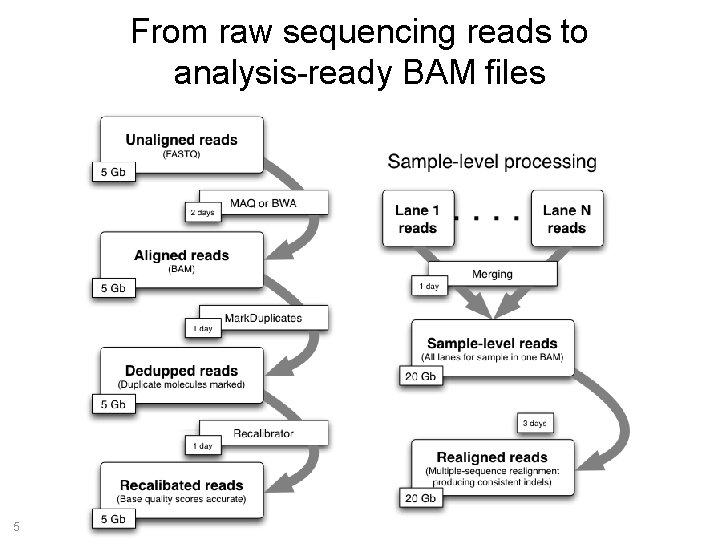
From raw sequencing reads to analysis-ready BAM files 5
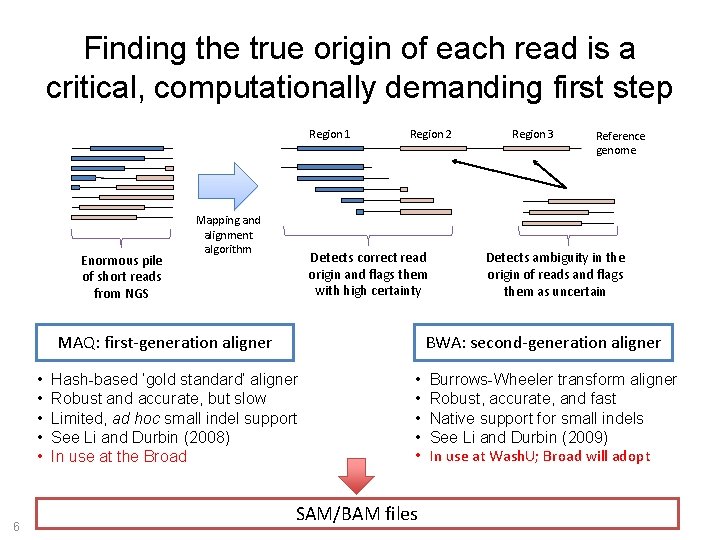
Finding the true origin of each read is a critical, computationally demanding first step Region 1 Enormous pile of short reads from NGS Mapping and alignment algorithm Region 2 Detects correct read origin and flags them with high certainty MAQ: first-generation aligner • • • 6 Region 3 Reference genome Detects ambiguity in the origin of reads and flags them as uncertain BWA: second-generation aligner Hash-based ‘gold standard’ aligner Robust and accurate, but slow Limited, ad hoc small indel support See Li and Durbin (2008) In use at the Broad • • • SAM/BAM files Burrows-Wheeler transform aligner Robust, accurate, and fast Native support for small indels See Li and Durbin (2009) In use at Wash. U; Broad will adopt

The BAM format facilitates sequenceragnostic analyses Bases, quality scores, (optionally) alignments, and meta data BAM file Read name Alignment gap information Quality scores (fastq format) SLX 1: 1: 127: 63: 4 … 1 10052169 … 23 M 6 N 10 M … GAAGATACTGGTTTTTTTCTTATGAGACGGAGT 768832'48: : : ; ; : /78$88818099897 SM: Z: JPTGBMN 01 … Locus BAM file allows us to represent the data of any sequencer. Analyses can then be conducted largely agnostic to the particular sequencer used. 7 Read sequence Meta data Data processing and analysis Courtesy of Kiran Garimella

Duplicate reads with early PCR errors offer false SNP evidence 1 KG 454 Pilot 3 lane Duplicate reads come from same molecular fragment. Including only one read for each fragment eliminates many false-positives. 1000 Genomes Pilot 3 NA 12878 hybrid capture For whole-exome data, 5 -10% of reads can be duplicates Excluding duplicates avoids calling many artifactual SNPs Duplicates not removed Duplicates removed Target size 2. 4 Mb Variants 2734 2082 % in db. SNP 59. 33% 77. 47% Only 10 of the 652 eliminated SNPs are in db. SNP; Almost all removed SNPs are FP since >90% of SNPs are in db. SNP 8 Courtesy of Kiran Garimella
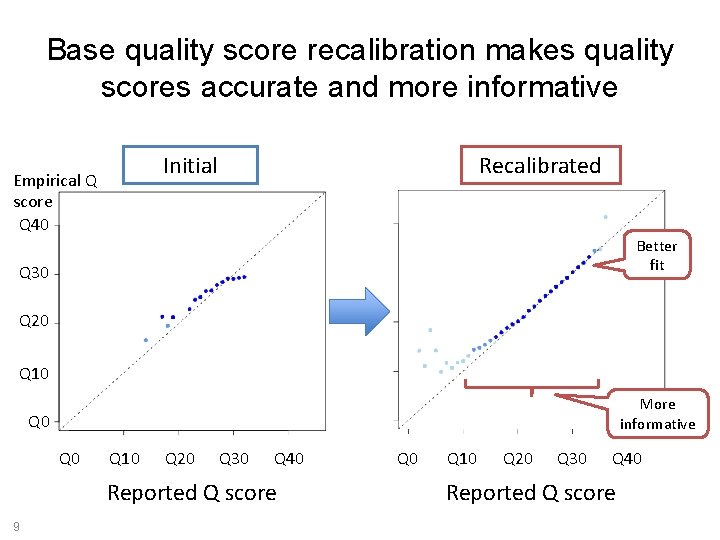
Base quality score recalibration makes quality scores accurate and more informative Initial Empirical Q score Q 40 Recalibrated Better fit Q 30 Q 20 Q 10 More informative Q 0 Q 10 Q 20 Q 30 Q 40 Reported Q score 9 Q 0 Q 10 Q 20 Q 30 Q 40 Reported Q score
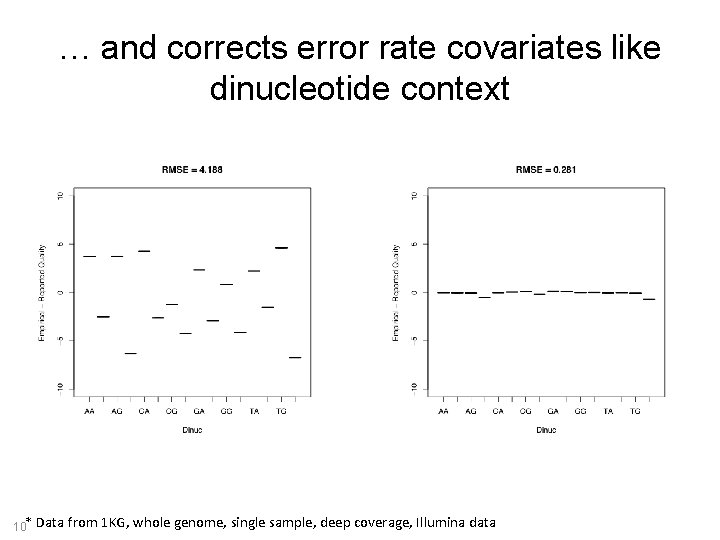
… and corrects error rate covariates like dinucleotide context 10* Data from 1 KG, whole genome, single sample, deep coverage, Illumina data

Recalibration identifies high-quality bases and improves SNP calls 1 KG 454 lane Initial Recalibation No. bases in lanes 80 M Lane wide reported Q 28. 7 28. 2 Lane wide empirical Q 28. 4 17, 554 9, 635 % of true Q 25 bases 89% 95% % of true Q 30 bases 0% 53% RMSE between Q reported and empirical Identifies >50% bases as true Q 30 Results in ~10% more SNP calls at same quality compared to unrecalibrated data
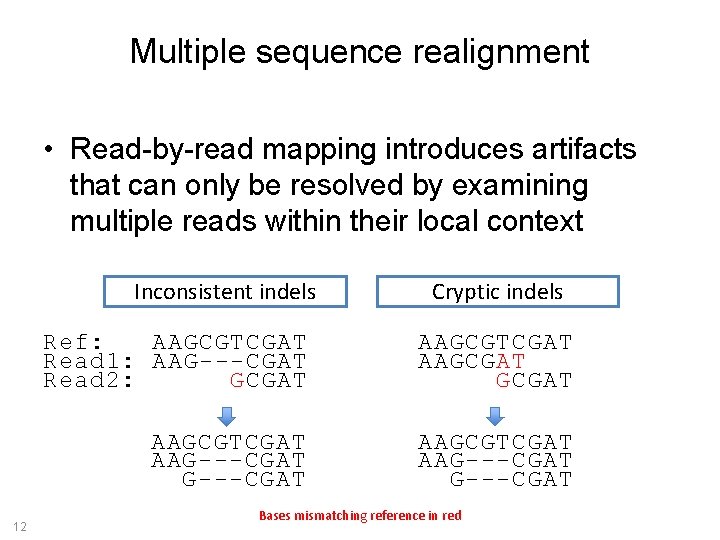
Multiple sequence realignment • Read-by-read mapping introduces artifacts that can only be resolved by examining multiple reads within their local context Inconsistent indels 12 Cryptic indels Ref: AAGCGTCGAT Read 1: AAG---CGAT Read 2: GCGAT AAGCGTCGAT AAGCGAT AAGCGTCGAT AAG---CGAT Bases mismatching reference in red
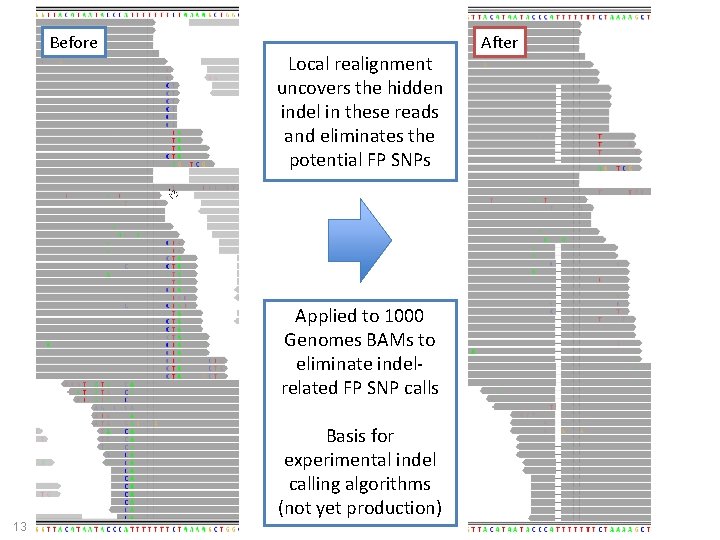
Before Local realignment uncovers the hidden indel in these reads and eliminates the potential FP SNPs Applied to 1000 Genomes BAMs to eliminate indelrelated FP SNP calls Basis for experimental indel calling algorithms (not yet production) 13 After

Analysis-ready BAM file • At this stage, we have taken the machine generated reads and: – Aligned them to the genome – Flagged QC− reads and molecular duplicates – Generated empirically accurate base quality scores – Realigned reads to be locally mutually consistent • These analysis-ready BAM files are then: – Distributed to the project via db. GAP – Sent into downstream variant detection 14
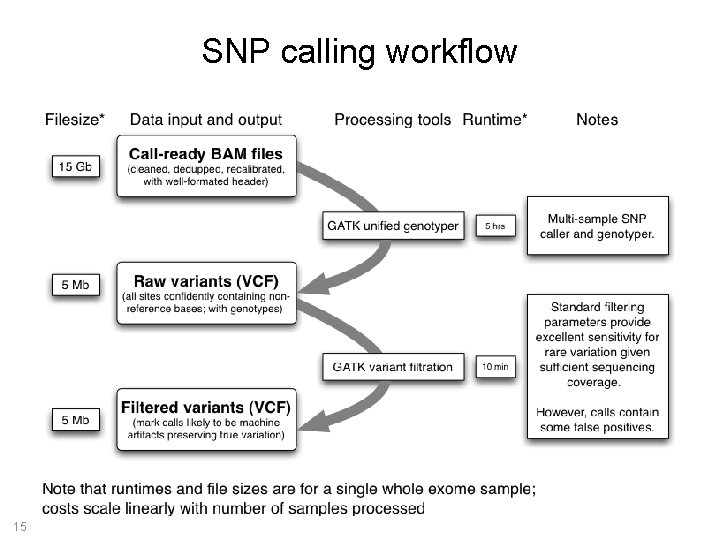
SNP calling workflow 15

GATK single sample genotype likelihoods Bayesian model Likelihood of the Likelihood for the. Prior for the data given the Independent base model genotype • Likelihood of data computed using pileup of bases and associated quality scores at given locus • Only “good bases” are included: those satisfying minimum base quality, mapping read quality, pair mapping quality, NQS • P(b | G) uses platform-specific confusion matrices • L(G|D) computed for all 10 genotypes 16 See http: //www. broadinstitute. org/gsa/wiki/index. php/Unified_genotyper for more information
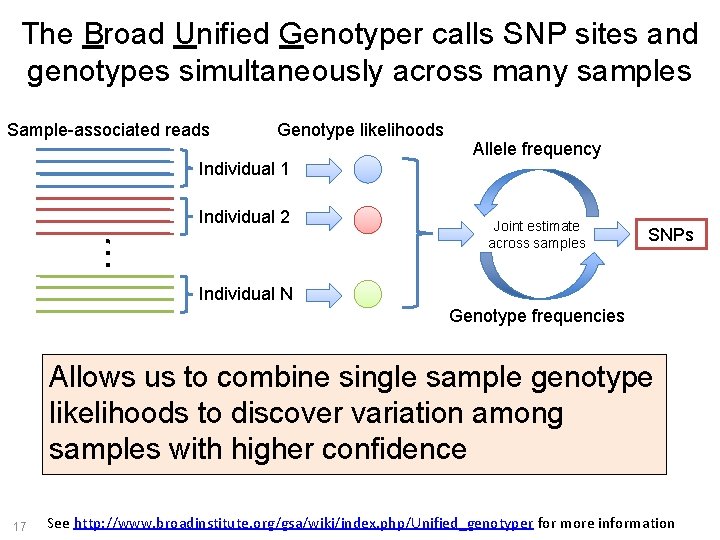
The Broad Unified Genotyper calls SNP sites and genotypes simultaneously across many samples Sample-associated reads Genotype likelihoods Allele frequency Individual 1 Individual 2 Joint estimate across samples SNPs Individual N Genotype frequencies Allows us to combine single sample genotype likelihoods to discover variation among samples with higher confidence 17 See http: //www. broadinstitute. org/gsa/wiki/index. php/Unified_genotyper for more information

Variant call format (VCF) files: calls, genotypes, and calling annotations VCF record for an A/G SNP at 22: 49582364 INFO field 22 49582364. A G 198. 96 0 AB=0. 67; AC No. chromosomescarrying AB AC=3; alt allele AF=0. 50; AN Total no. of chromosomes Hrun AN=6; DP=87; Dels=0. 00; AF Allele frequency MQ HRun=1; DP Depth of coverage MQ 0 MQ=71. 31; MQ 0=22; QD QUAL score over depth SB QD=2. 29; SB=-31. 76 GT: DP: GQ 0/1: 12: 99. 00 0/1: 11: 89. 43 0/1: 28: 37. 78 Allele balance of ref/alt in hets Length of longest contiguous homopolymer RMS MAPQ of all reads No. of MAPQ 0 reads at locus Estimated SB score Heterozygous genotype A/G in all three individuals 18 See http: //www. broadinstitute. org/gsa/wiki/index. php/ for more information

SNP calling artifacts • Machine artifacts, mismapped reads, misaligned indels can result in false SNP calls – Raw novel SNP calls can have ~5 -20% false discovery rate • SNPs with features character of machine artifact are marked as filtered in the VCF – Separates reliable calls from likely errors • Working on methods to calibrate SNP quality scores that incorporate error covariates 19
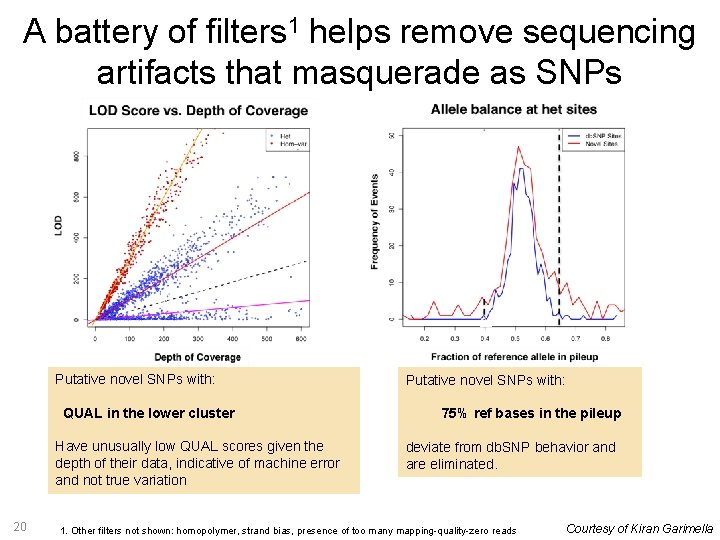
A battery of filters 1 helps remove sequencing artifacts that masquerade as SNPs Putative novel SNPs with: QUAL in the lower cluster Have unusually low QUAL scores given the depth of their data, indicative of machine error and not true variation 20 Putative novel SNPs with: 75% ref bases in the pileup deviate from db. SNP behavior and are eliminated. 1. Other filters not shown: homopolymer, strand bias, presence of too many mapping-quality-zero reads Courtesy of Kiran Garimella
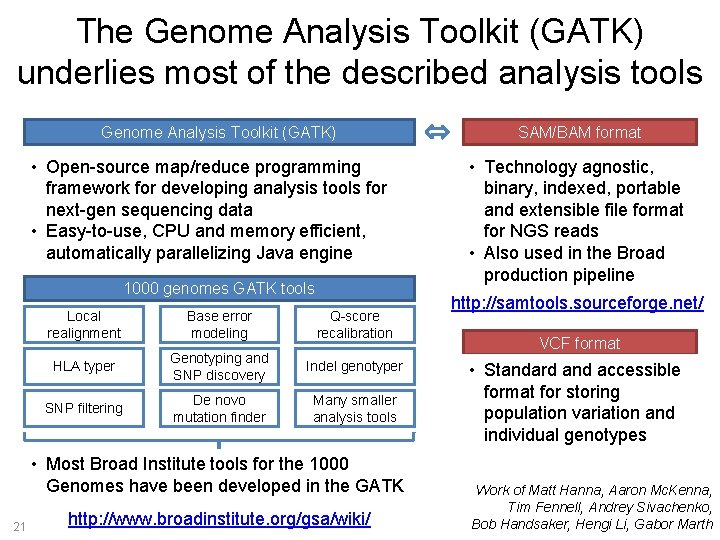
The Genome Analysis Toolkit (GATK) underlies most of the described analysis tools Genome Analysis Toolkit (GATK) • Open-source map/reduce programming framework for developing analysis tools for next-gen sequencing data • Easy-to-use, CPU and memory efficient, automatically parallelizing Java engine 1000 genomes GATK tools Local realignment Base error modeling Q-score recalibration HLA typer Genotyping and SNP discovery Indel genotyper SNP filtering De novo mutation finder Many smaller analysis tools • Most Broad Institute tools for the 1000 Genomes have been developed in the GATK 21 http: //www. broadinstitute. org/gsa/wiki/ SAM/BAM format • Technology agnostic, binary, indexed, portable and extensible file format for NGS reads • Also used in the Broad production pipeline http: //samtools. sourceforge. net/ VCF format • Standard and accessible format for storing population variation and individual genotypes Work of Matt Hanna, Aaron Mc. Kenna, Tim Fennell, Andrey Sivachenko, Bob Handsaker, Hengi Li, Gabor Marth

Develop and apply methods in NGS to medical genetics projects like ESP and 1000 Genome • The Genome Sequencing and Analysis group in Medical and Population Genetics at the Broad Institute is hiring Computational Biologist Ph. D. -level position focused on technology development and medical genetics analysis Bioinformatic Analyst B. A. /M. A. -level position focused on production data processing and analysis Talk to me for more information or email depristo@broadinstitute. org 22

ANALYSIS OF SNP CALLING ACROSS CENTERS AND SAMPLES: KIRAN GARIMELLA

Appendix
- Slides: 24