A BIONETGEN Tutorial Outline Downloading BNG Installing BNG

A BIONETGEN Tutorial

Outline • Downloading BNG • Installing BNG • Running BNG – text-based interface – graphical interface – RULEBUILDER – web-based interface – GETBONNIE • Simple example • Extending the example • Detailed model of TLR 4 signaling
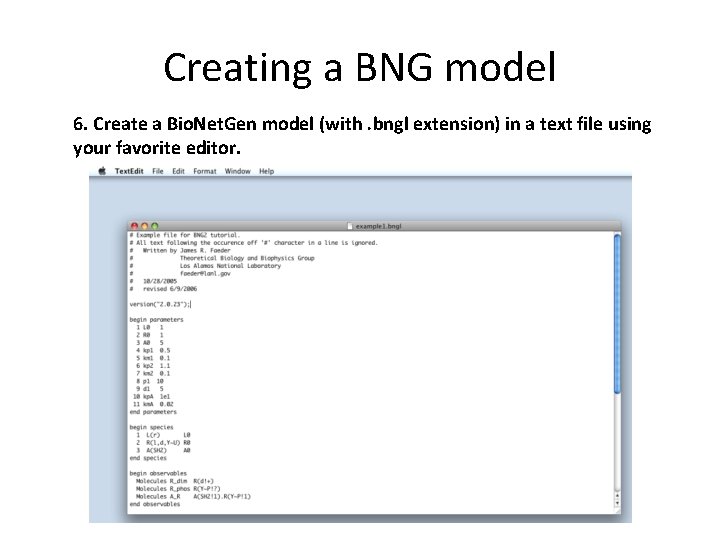
Creating a BNG model 6. Create a Bio. Net. Gen model (with. bngl extension) in a text file using your favorite editor.
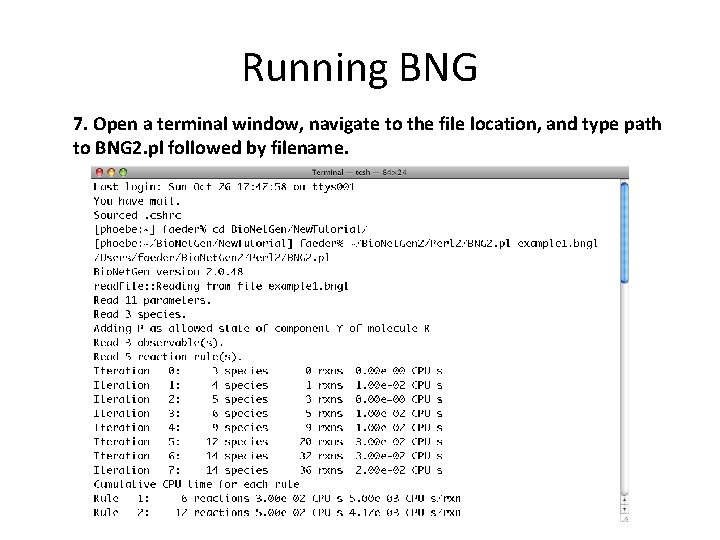
Running BNG 7. Open a terminal window, navigate to the file location, and type path to BNG 2. pl followed by filename.

The BNG logfile 8. (optional) Piping output to a logfile. ~/Bio. Net. Gen 2/Perl 2/BNG 2. pl example 1. bngl > & example 1. log Bio. Net. Gen version 2. 0. 48 read. File: : Reading from file example 1. bngl Read 11 parameters. Read 3 species. Adding P as allowed state of component Y of molecule R Read 3 observable(s). Read 5 reaction rule(s). Iteration 0: 3 species 0 rxns 0. 00 e+00 CPU s Iteration 1: 4 species 1 rxns 1. 00 e-02 CPU s Iteration 2: 5 species 3 rxns 0. 00 e+00 CPU s Iteration 3: 6 species 5 rxns 1. 00 e-02 CPU s Iteration 4: 9 species 9 rxns 1. 00 e-02 CPU s Iteration 5: 12 species 20 rxns 3. 00 e-02 CPU s Iteration 6: 14 species 32 rxns 3. 00 e-02 CPU s Iteration 7: 14 species 36 rxns 2. 00 e-02 CPU s Cumulative CPU time for each rule Rule 1: 6 reactions 3. 00 e-02 CPU s 5. 00 e-03 CPU s/rxn Rule 2: 12 reactions 5. 00 e-02 CPU s 4. 17 e-03 CPU s/rxn Rule 3: 3 reactions 0. 00 e+00 CPU s/rxn Rule 4: 5 reactions 2. 00 e-02 CPU s 4. 00 e-03 CPU s/rxn Rule 5: 10 reactions 1. 00 e-02 CPU s 1. 00 e-03 CPU s/rxn Total : 36 reactions 1. 10 e-01 CPU s 3. 06 e-03 CPU s/rxn Wrote network to example 1. net. CPU TIME: generate_network 0. 1 s. Network simulation using ODEs Running run_network on phoebe
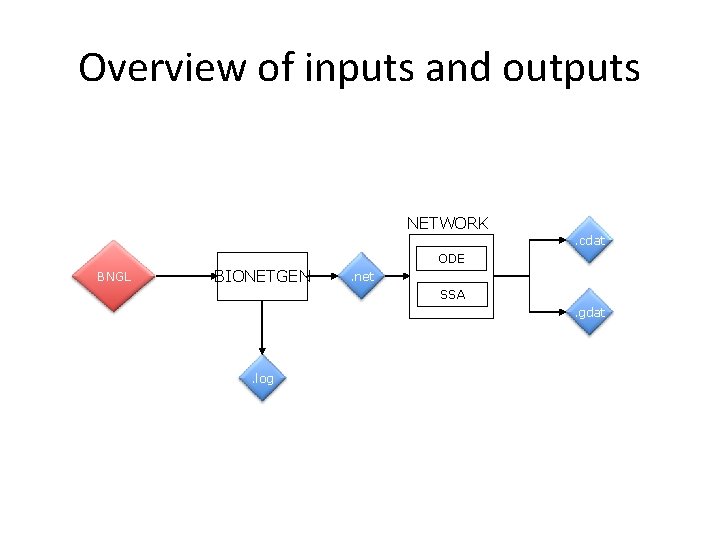
Overview of inputs and outputs NETWORK . cdat ODE BNGL BIONETGEN . net SSA. gdat . log

Elements of a BNGL file • • • parameters molecule types seed species reaction rules observables actions
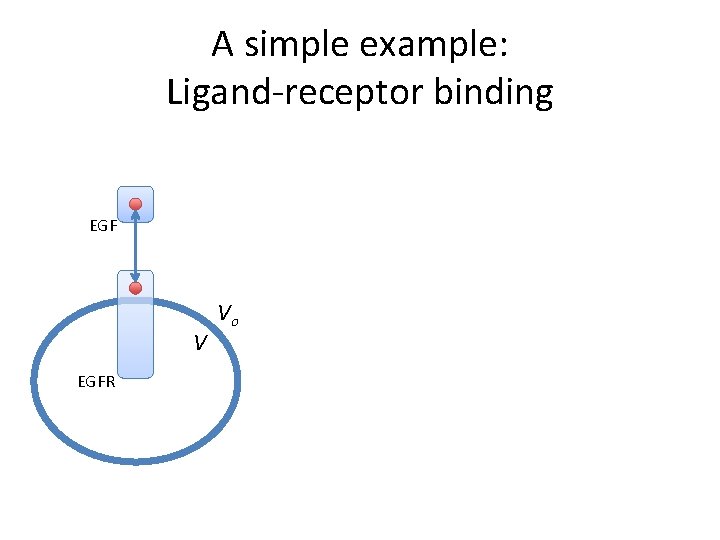
A simple example: Ligand-receptor binding EGF V EGFR Vo
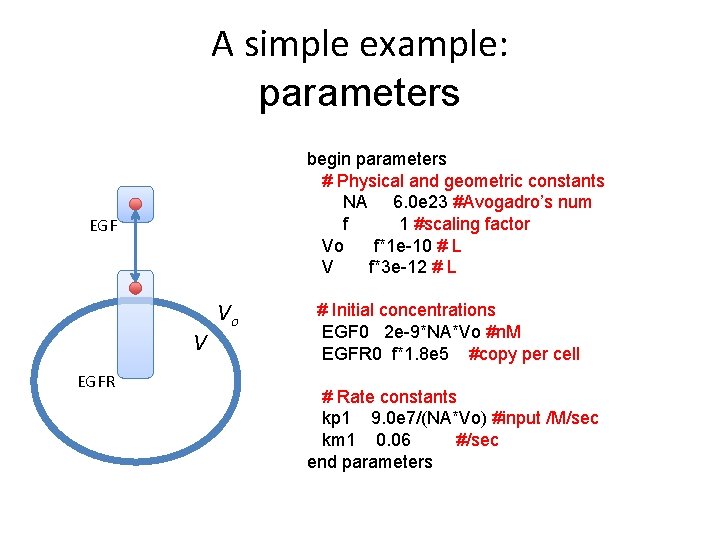
A simple example: parameters begin parameters # Physical and geometric constants NA 6. 0 e 23 #Avogadro’s num f 1 #scaling factor Vo f*1 e-10 # L V f*3 e-12 # L EGF V EGFR Vo # Initial concentrations EGF 0 2 e-9*NA*Vo #n. M EGFR 0 f*1. 8 e 5 #copy per cell # Rate constants kp 1 9. 0 e 7/(NA*Vo) #input /M/sec km 1 0. 06 #/sec end parameters
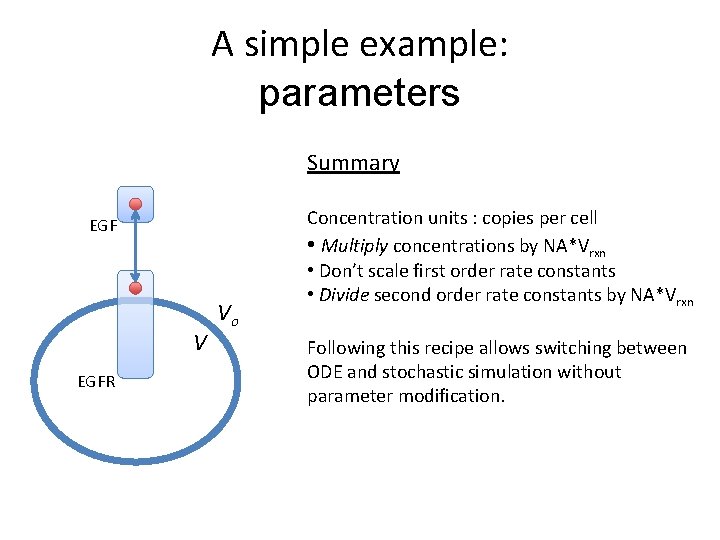
A simple example: parameters Summary EGF V EGFR Vo Concentration units : copies per cell • Multiply concentrations by NA*Vrxn • Don’t scale first order rate constants • Divide second order rate constants by NA*Vrxn Following this recipe allows switching between ODE and stochastic simulation without parameter modification.
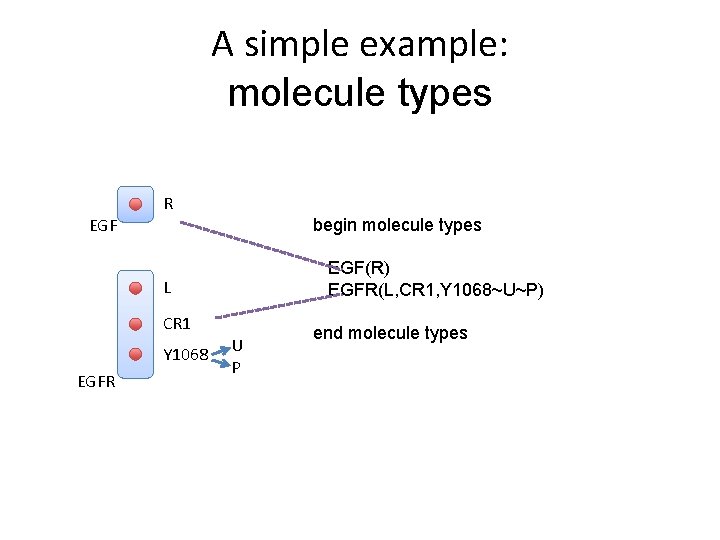
A simple example: molecule types R EGF begin molecule types EGF(R) EGFR(L, CR 1, Y 1068~U~P) L CR 1 Y 1068 EGFR U P end molecule types
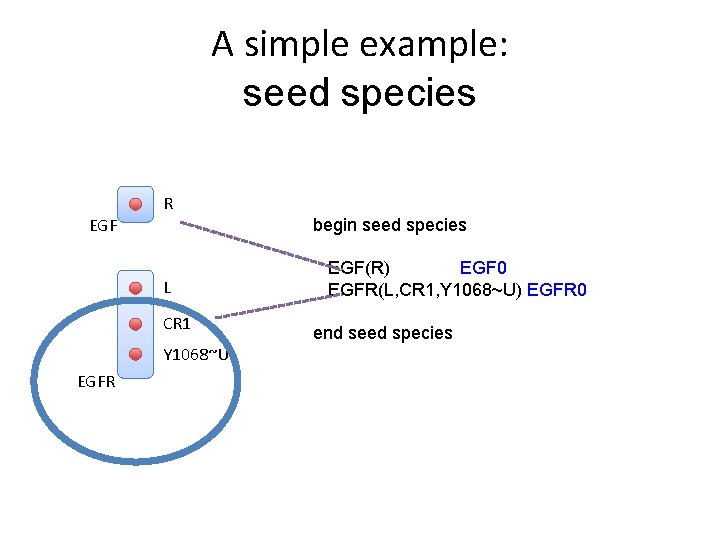
A simple example: seed species R EGF begin seed species L CR 1 Y 1068~U EGFR EGF(R) EGF 0 EGFR(L, CR 1, Y 1068~U) EGFR 0 end seed species

Pattern basics Basic Definition: A set of molecules that may be partially specified and selects a set of species through its allowed mappings. Pattern EGF(R) EGFR(L) EGF(R) EGFR(L, CR 1, Y 1068~U) Mapping Species EGFR(L, CR 1, Y 1068~P)

A simple example: reaction rules + EGFR k+1 k-1 EGFR begin reaction rules EGF(R) + EGFR(L) <-> EGF(R!1). EGFR(L!1) kp 1, km 1 end reaction rules

A simple example: actions begin parameters … end parameters begin molecule types EGF(R) EGFR(L, CR 1, Y 1068~U~P) end molecule types begin seed species EGF(R) EGF 0 EGFR(L, CR 1, Y 1068~U) EGFR 0 end seed species begin reaction rules EGF(R) + EGFR(L) <-> EGF(R!1). EGFR(L!1) kp 1, km 1 end reaction rules # actions generate_network({overwrite=>1}); Generates network of species and reactions by iterative application of rules starting with seed species

A simple example: Application of the forward binding rule Reaction rule EGF(R) + EGFR(L) -> EGF(R!1). EGFR(L!1) kp 1 Add bond Step 1: Match patterns onto reactant species. Step 2: Copy selected species to product side. Step 3: Apply transformation specified by rule.
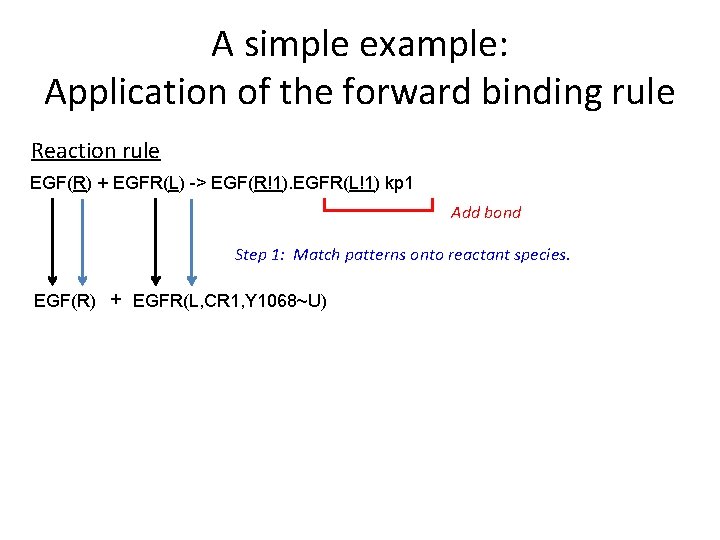
A simple example: Application of the forward binding rule Reaction rule EGF(R) + EGFR(L) -> EGF(R!1). EGFR(L!1) kp 1 Add bond Step 1: Match patterns onto reactant species. EGF(R) + EGFR(L, CR 1, Y 1068~U)
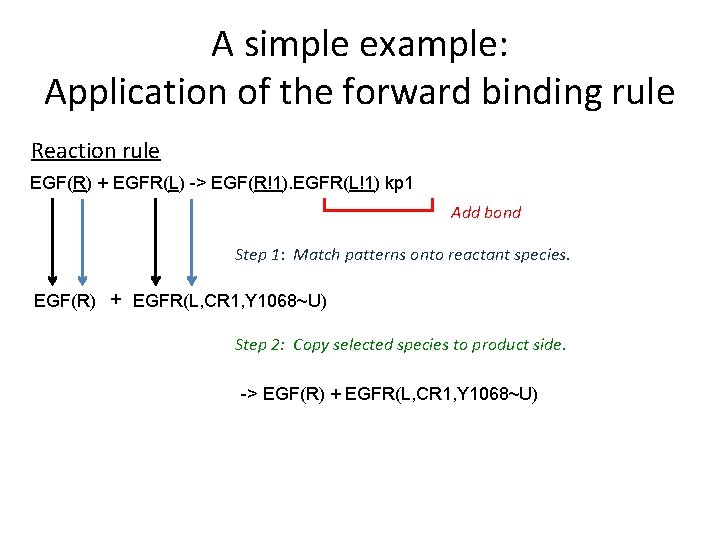
A simple example: Application of the forward binding rule Reaction rule EGF(R) + EGFR(L) -> EGF(R!1). EGFR(L!1) kp 1 Add bond Step 1: Match patterns onto reactant species. EGF(R) + EGFR(L, CR 1, Y 1068~U) Step 2: Copy selected species to product side. -> EGF(R) + EGFR(L, CR 1, Y 1068~U)
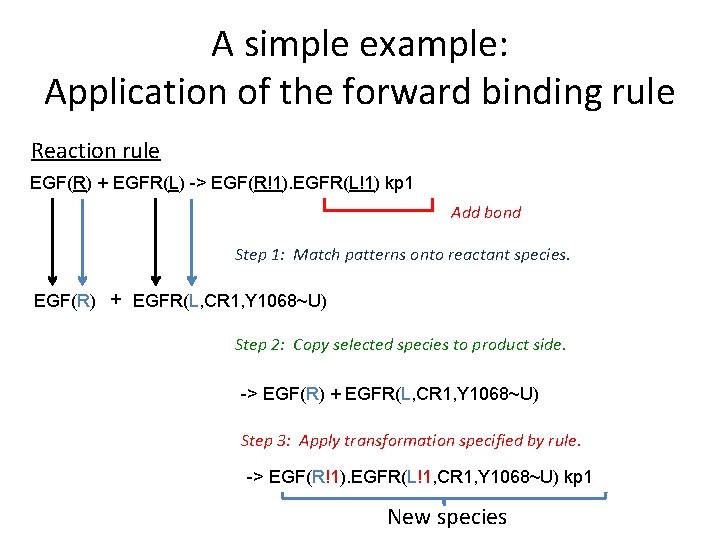
A simple example: Application of the forward binding rule Reaction rule EGF(R) + EGFR(L) -> EGF(R!1). EGFR(L!1) kp 1 Add bond Step 1: Match patterns onto reactant species. EGF(R) + EGFR(L, CR 1, Y 1068~U) Step 2: Copy selected species to product side. -> EGF(R) + EGFR(L, CR 1, Y 1068~U) Step 3: Apply transformation specified by rule. -> EGF(R!1). EGFR(L!1, CR 1, Y 1068~U) kp 1 New species
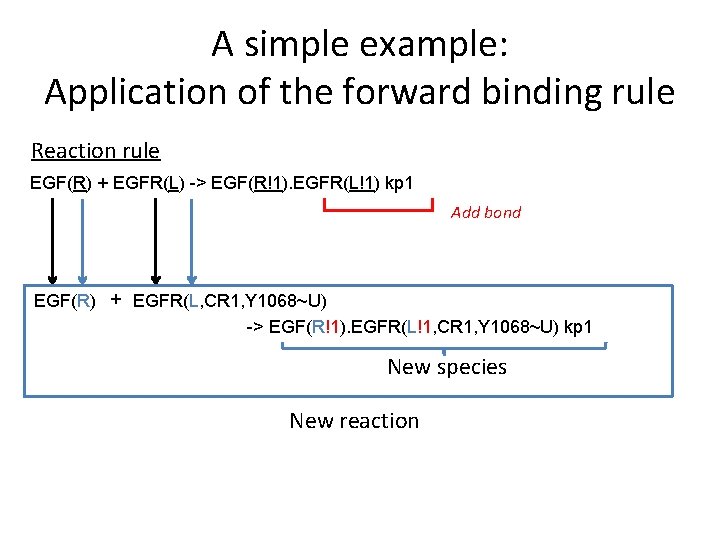
A simple example: Application of the forward binding rule Reaction rule EGF(R) + EGFR(L) -> EGF(R!1). EGFR(L!1) kp 1 Add bond EGF(R) + EGFR(L, CR 1, Y 1068~U) -> EGF(R!1). EGFR(L!1, CR 1, Y 1068~U) kp 1 New species New reaction

A simple example: Application of the reverse binding rule Reaction rule EGF(R!1). EGFR(L!1) -> EGF(R) + EGFR(L) km 1 Delete bond EGF(R!1). EGFR(L!1, CR 1, Y 1068~U)

A simple example: Application of the reverse binding rule Reaction rule EGF(R!1). EGFR(L!1) -> EGF(R) + EGFR(L) km 1 Delete bond EGF(R!1). EGFR(L!1, CR 1, Y 1068~U) -> EGF(R!1). EGFR(L!1, CR 1, Y 1068~U)
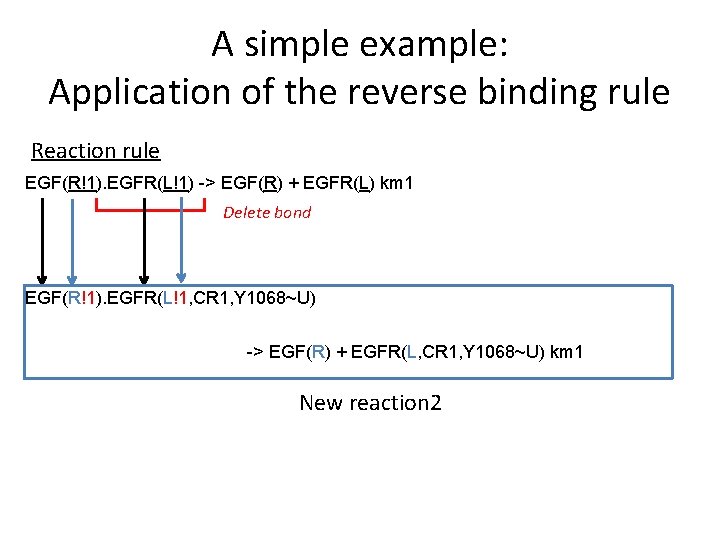
A simple example: Application of the reverse binding rule Reaction rule EGF(R!1). EGFR(L!1) -> EGF(R) + EGFR(L) km 1 Delete bond EGF(R!1). EGFR(L!1, CR 1, Y 1068~U) -> EGF(R) + EGFR(L, CR 1, Y 1068~U) km 1 New reaction 2

A simple example: Overview of generate_network seed species EGF(R) EGFR(L, CR 1, Y 1068~U) Iteration 1: rule 1 f: EGF(R) + EGFR(L) -> EGF(R!1). EGFR(L!1) kp 1

A simple example: Overview of generate_network seed species EGF(R) EGFR(L, CR 1, Y 1068~U) Iteration 1: rule 1 f: EGF(R) + EGFR(L) -> EGF(R!1). EGFR(L!1) kp 1 new reaction: EGF(R) + EGFR(L, CR 1, Y 1068~U) -> EGF(R!1). EGFR(L!1, CR 1, Y 1068~U) kp 1 new species: EGF(R!1). EGFR(L!1, CR 1, Y 1068~U)

A simple example: Overview of generate_network seed species EGF(R) EGFR(L, CR 1, Y 1068~U) Iteration 1: rule 1 f: EGF(R) + EGFR(L) -> EGF(R!1). EGFR(L!1) kp 1 new reaction: EGF(R) + EGFR(L, CR 1, Y 1068~U) -> EGF(R!1). EGFR(L!1, CR 1, Y 1068~U) kp 1 new species: EGF(R!1). EGFR(L!1, CR 1, Y 1068~U) Network after Iteration 1 species 1 EGF(R) 2 EGFR(L, CR 1, Y 1068~U) 3 EGF(R!1). EGFR(L!1, CR 1, Y 1068~U) reactions 1 1, 2 3 kp 1

A simple example: Overview of generate_network From iteration 1 species 1 EGF(R) 2 EGFR(L, CR 1, Y 1068~U) 3 EGF(R!1). EGFR(L!1, CR 1, Y 1068~U) Iteration 2: rule 1 r: EGF(R!1). EGFR(L!1) -> EGF(R) + EGFR(L) km 1 new reaction: EGF(R!1). EGFR(L!1, CR 1, Y 1068~U) -> EGF(R) + EGFR(L, CR 1, Y 1068~U) km 1 Network after Iteration 2 species 1 EGF(R) 2 EGFR(L, CR 1, Y 1068~U) 3 EGF(R!1). EGFR(L!1, CR 1, Y 1068~U) reactions 1 1, 2 3 kp 1 # rule 1 f 2 3 1, 2 km 1 # rule 1 r

A simple example: Overview of generate_network From iteration 1 species 1 EGF(R) 2 EGFR(L, CR 1, Y 1068~U) 3 EGF(R!1). EGFR(L!1, CR 1, Y 1068~U) Iteration 2: rule 1 r: EGF(R!1). EGFR(L!1) -> EGF(R) + EGFR(L) km 1 new reaction: EGF(R!1). EGFR(L!1, CR 1, Y 1068~U) -> EGF(R) + EGFR(L, CR 1, Y 1068~U) km 1 Network after Iteration 2 - final species 1 EGF(R) 2 EGFR(L, CR 1, Y 1068~U) 3 EGF(R!1). EGFR(L!1, CR 1, Y 1068~U) reactions 1 1, 2 3 kp 1 # rule 1 f 2 3 1, 2 km 1 # rule 1 r
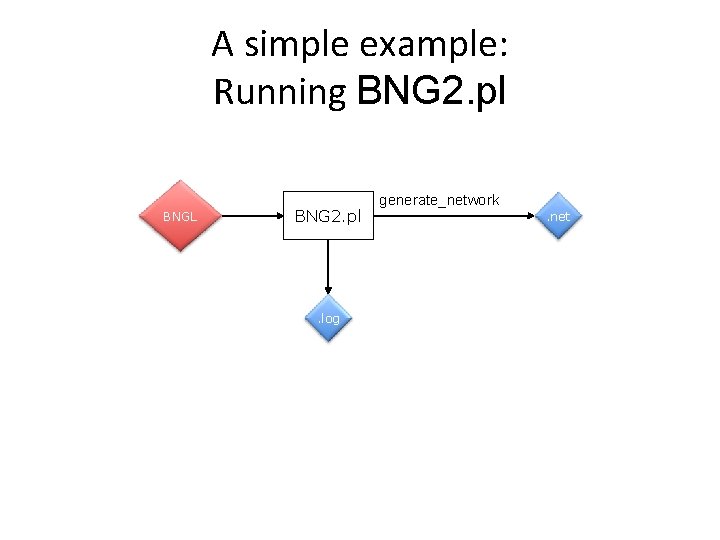
A simple example: Running BNG 2. pl BNGL BNG 2. pl . log generate_network. net
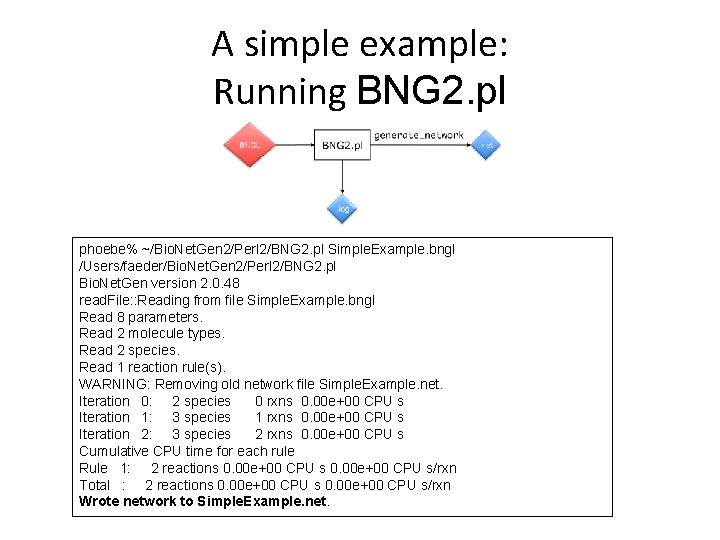
A simple example: Running BNG 2. pl phoebe% ~/Bio. Net. Gen 2/Perl 2/BNG 2. pl Simple. Example. bngl /Users/faeder/Bio. Net. Gen 2/Perl 2/BNG 2. pl Bio. Net. Gen version 2. 0. 48 read. File: : Reading from file Simple. Example. bngl Read 8 parameters. Read 2 molecule types. Read 2 species. Read 1 reaction rule(s). WARNING: Removing old network file Simple. Example. net. Iteration 0: 2 species 0 rxns 0. 00 e+00 CPU s Iteration 1: 3 species 1 rxns 0. 00 e+00 CPU s Iteration 2: 3 species 2 rxns 0. 00 e+00 CPU s Cumulative CPU time for each rule Rule 1: 2 reactions 0. 00 e+00 CPU s/rxn Total : 2 reactions 0. 00 e+00 CPU s/rxn Wrote network to Simple. Example. net.
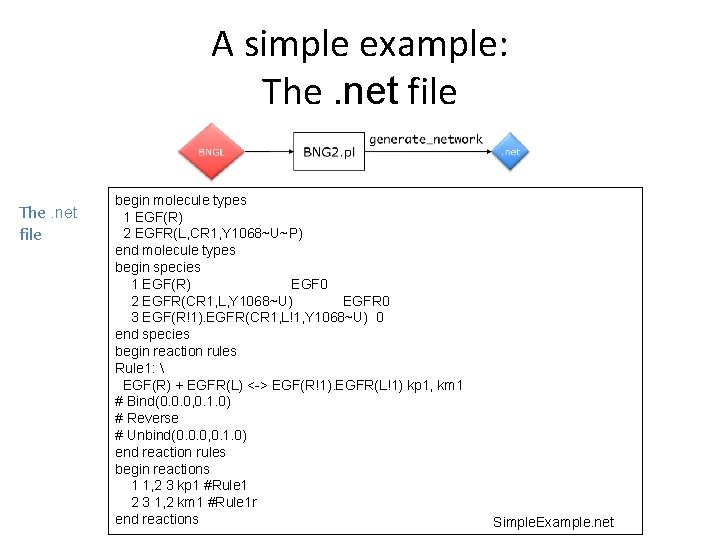
A simple example: The. net file begin molecule types 1 EGF(R) 2 EGFR(L, CR 1, Y 1068~U~P) end molecule types begin species 1 EGF(R) EGF 0 2 EGFR(CR 1, L, Y 1068~U) EGFR 0 3 EGF(R!1). EGFR(CR 1, L!1, Y 1068~U) 0 end species begin reaction rules Rule 1: EGF(R) + EGFR(L) <-> EGF(R!1). EGFR(L!1) kp 1, km 1 # Bind(0. 0. 0, 0. 1. 0) # Reverse # Unbind(0. 0. 0, 0. 1. 0) end reaction rules begin reactions 1 1, 2 3 kp 1 #Rule 1 2 3 1, 2 km 1 #Rule 1 r end reactions Simple. Example. net
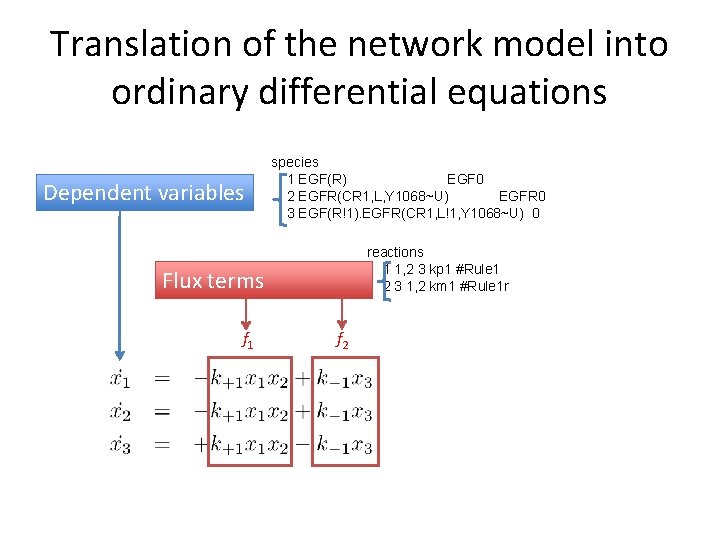
Translation of the network model into ordinary differential equations Dependent variables species 1 EGF(R) EGF 0 2 EGFR(CR 1, L, Y 1068~U) EGFR 0 3 EGF(R!1). EGFR(CR 1, L!1, Y 1068~U) 0 reactions 1 1, 2 3 kp 1 #Rule 1 2 3 1, 2 km 1 #Rule 1 r Flux terms f 1 f 2
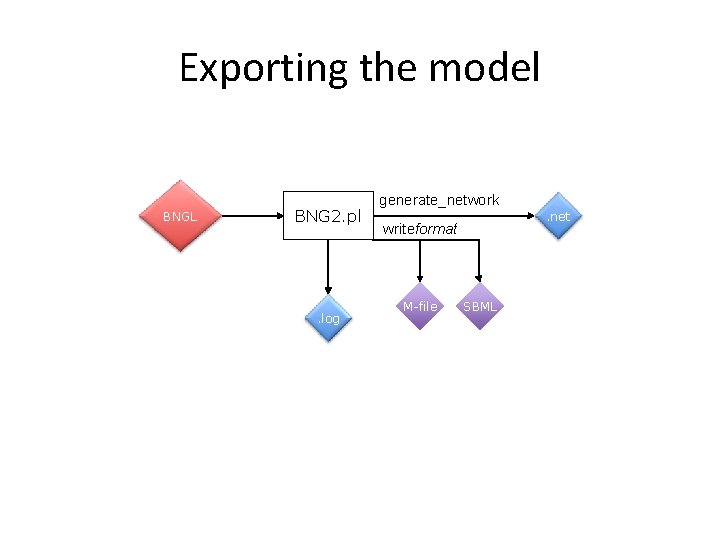
Exporting the model BNGL BNG 2. pl . log generate_network. net writeformat M-file SBML

Exporting model equations to Matlab: write. Mfile begin parameters … end parameters begin molecule types EGF(R) EGFR(L, CR 1, Y 1068~U~P) end molecule types begin seed species EGF(R) EGF 0 EGFR(L, CR 1, Y 1068~U) EGFR 0 end seed species begin reaction rules EGF(R) + EGFR(L) <-> EGF(R!1). EGFR(L!1) kp 1, km 1 end reaction rules # actions generate_network({overwrite=>1}); write. Mfile();

Ordinary differential equations in the M-file Simple. Example. m % M-file for model Simple. Example created by Bio. Net. Gen 2. 0. 48 function [t_out, obs_out, x_out]= Simple. Example(tend) Nspecies=3; Nreactions=2; obs_out=zeros(1); % Parameters NA=6. 0 e 23; f=1; Vo=f*1 e-10; V=f*3 e-12; EGF 0=(2 e-9*NA)*Vo; EGFR 0=f*1. 8 e 5; kp 1=3. 0 e 6/(NA*V); km 1=0. 06; % Intial concentrations x 0= [ EGF 0; EGFR 0; 0; ]; % Reaction flux function f= flux(t, x) f(1)=kp 1*x(1)*x(2); f(2)=km 1*x(3); end % Derivative function d= xdot(t, x) f=flux(t, x); d(1, 1)= -f(1) +f(2); d(2, 1)= -f(1) +f(2); d(3, 1)= +f(1) -f(2); end snames={'EGF(R)', 'EGFR(CR 1, L, Y 1068~U)', 'EGF(R!1). EGFR(CR 1, L!1, Y 1068~U)'}; % Integrate ODEs [t_out, x_out]= ode 15 s(@xdot, [0 tend], x 0); % plot species concentrations plot(t_out, x_out); legend(snames); end
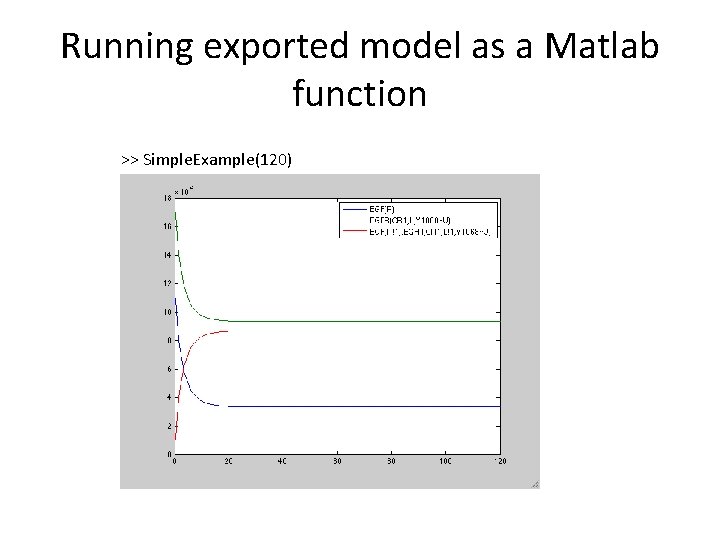
Running exported model as a Matlab function >> Simple. Example(120)

Export to the Systems Biology Markup Language (SBML) • SBML is an XML specification for biological models • Many simulation tools read and write SBML • Supports standard reaction network models • generate_network must be invoked prior to export • Support for rule-based models will appear in Level 3 (current level is 2+) • Website: http: //sbml. org
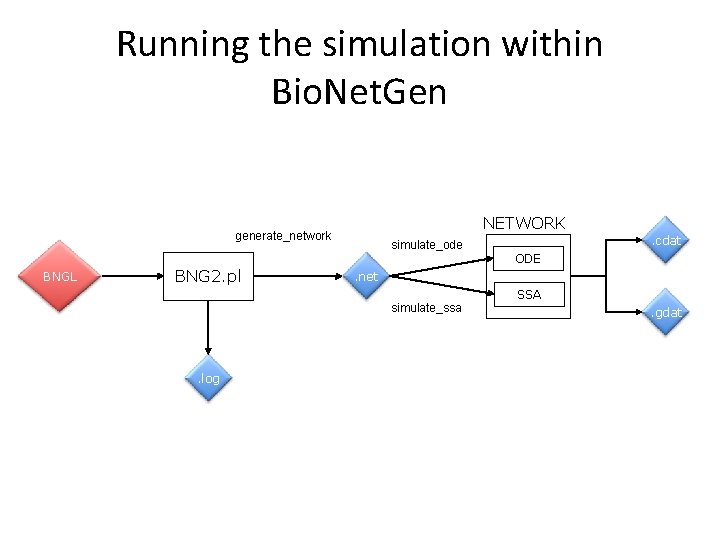
Running the simulation within Bio. Net. Gen NETWORK generate_network BNGL BNG 2. pl simulate_ode ODE . net simulate_ssa . log . cdat SSA. gdat

Running the simulation within Bio. Net. Gen begin parameters … end parameters begin molecule types EGF(R) EGFR(L, CR 1, Y 1068~U~P) end molecule types begin seed species EGF(R) EGF 0 EGFR(L, CR 1, Y 1068~U) EGFR 0 end seed species begin reaction rules EGF(R) + EGFR(L) <-> EGF(R!1). EGFR(L!1) kp 1, km 1 end reaction rules # actions generate_network({overwrite=>1}); save. Concentrations(); simulate_ode({suffix=>ode, t_end=>120, n_steps=>50});

Running the simulation within Bio. Net. Gen begin parameters … end parameters begin molecule types EGF(R) EGFR(L, CR 1, Y 1068~U~P) end molecule types begin seed species EGF(R) EGF 0 EGFR(L, CR 1, Y 1068~U) EGFR 0 end seed species BNG, like many other systems biology begin reaction rules applications, uses the CVODE library for EGF(R) + EGFR(L) <-> EGF(R!1). EGFR(L!1) kp 1, solving km 1 ODE’s. end reaction rules # actions generate_network({overwrite=>1}); save. Concentrations(); Sparse option can be used to solve systems up to about 50, 000 equations. simulate_ode({suffix=>ode, t_end=>120, n_steps=>50});

The. cdat file Simple. Example_ode. cdat # time 0. 00000000 e+00 2. 399999999 e+00 4. 799999998 e+00 7. 199999993 e+00 1 2 1. 200000000 e+05 6. 9387285118773594 e+04 5. 0363939566816996 e+04 4. 1819631802026437 e+04 3 1. 800000000 e+05 1. 2938728511877365 e+05 1. 1036393956681706 e+05 1. 0181963180202652 e+05 0. 00000000 e+00 5. 0612714881226493 e+04 6. 9636060433183069 e+04 7. 8180368197973614 e+04 Your favorite plotting package Some useful options • Phi. BPlot – Java-based (JFree. Chart) plotting utility provided with BNG • xmgrace – Unix / X windows graphing package great for XY plots • Matlab, Mathematica, gnuplot, Excel, etc.
The. cdat file in Phi. BPlot
The. cdat file in Phi. BPlot These numbers correspond to indices of species in. net file

The. cdat file in Phi. BPlot These numbers correspond to indices of species in. net file begin species 1 EGF(R) EGF 0 2 EGFR(CR 1, L, Y 1068~U) EGFR 0 3 EGF(R!1). EGFR(CR 1, L!1, Y 1068~U) 0 end species

Stochastic simulations in Bio. Net. Gen • Network simulation engine uses Gillespie Direct method • Adequate performance for networks with up to ~50, 000 species. • May be used in ‘on-the-fly’ mode, which generates species and reactions as needed
Stochastic simulations in Bio. Net. Gen • Network simulation engine uses Gillespie Direct method • Adequate performance for networks with up to ~50, 000 species. • May be used in ‘on-the-fly’ mode, which generates species and reactions as needed

Comparison between ODE and stochastic simulations begin parameters … end parameters begin molecule types EGF(R) EGFR(L, CR 1, Y 1068~U~P) end molecule types begin seed species EGF(R) EGF 0 EGFR(L, CR 1, Y 1068~U) EGFR 0 end seed species begin reaction rules EGF(R) + EGFR(L) <-> EGF(R!1). EGFR(L!1) kp 1, km 1 end reaction rules # actions generate_network({overwrite=>1}); save. Concentrations(); simulate_ode({suffix=>ode, t_end=>120, n_steps=>50}); reset. Concentrations(); simulate_ssa({suffix=>ssa, t_end=>120, n_steps=>50});
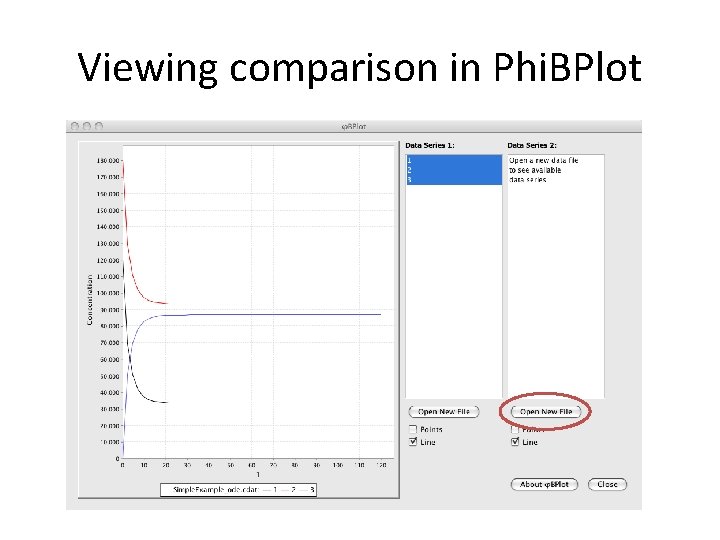
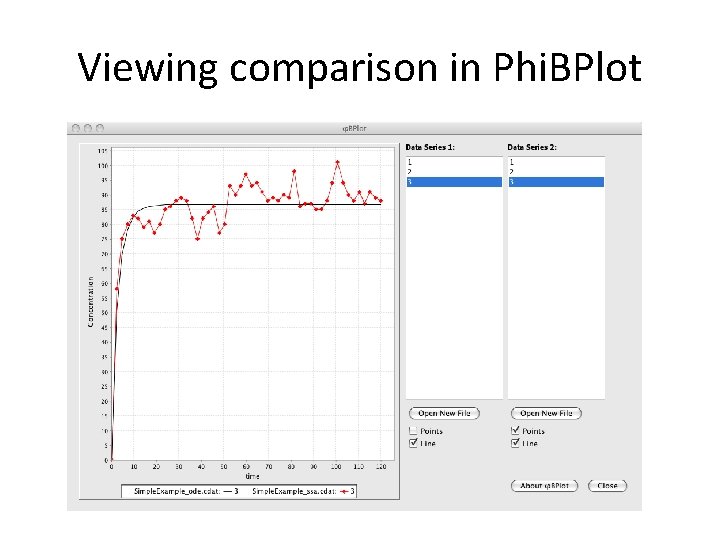
Viewing comparison in Phi. BPlot
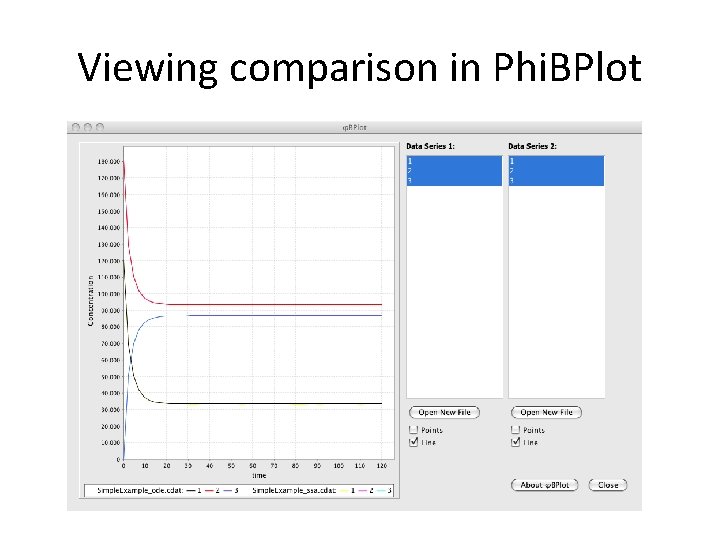
Viewing comparison in Phi. BPlot
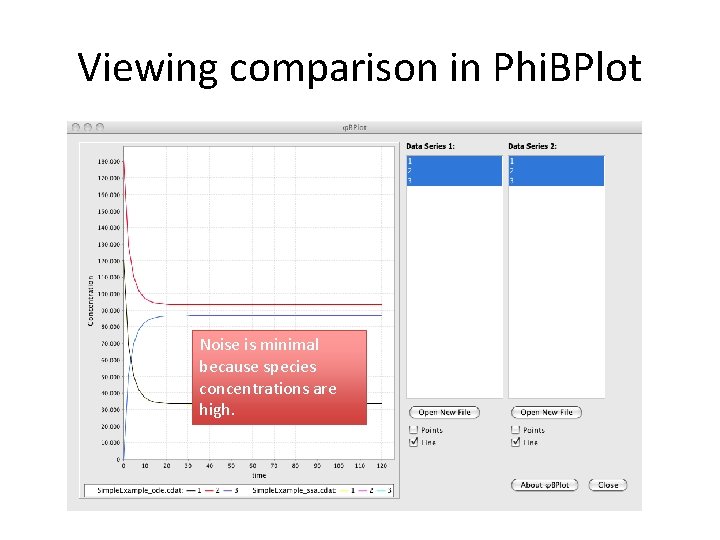
Viewing comparison in Phi. BPlot Noise is minimal because species concentrations are high.
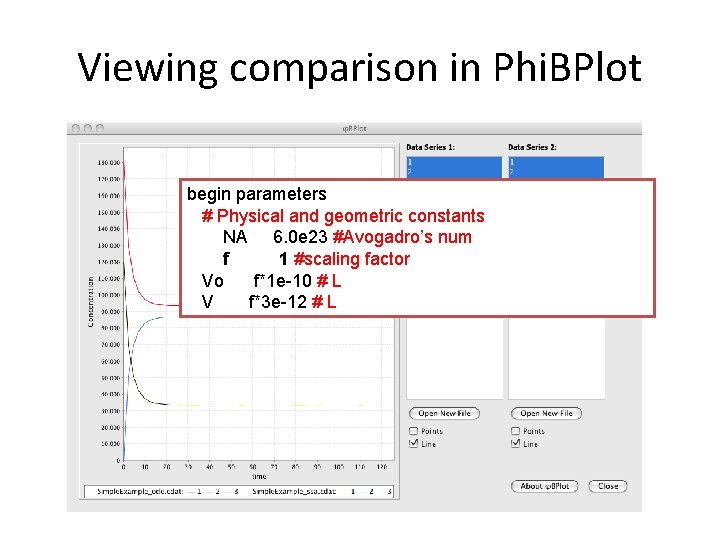
Viewing comparison in Phi. BPlot begin parameters # Physical and geometric constants NA 6. 0 e 23 #Avogadro’s num f 1 #scaling factor Vo f*1 e-10 # L V f*3 e-12 # L
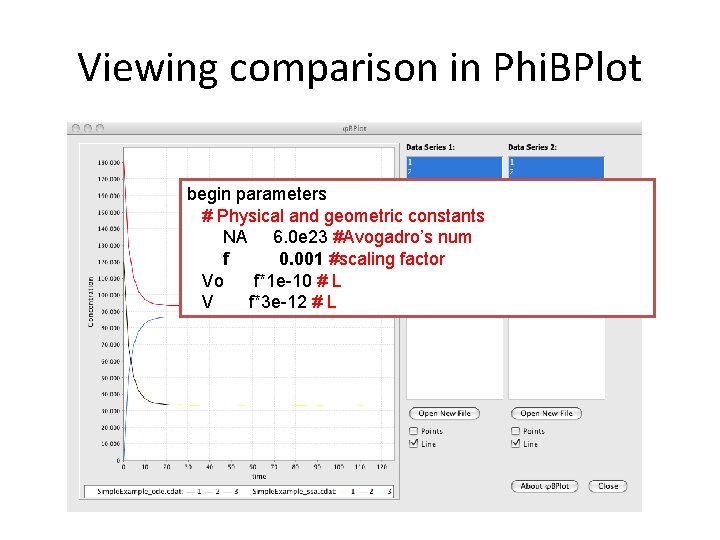
Viewing comparison in Phi. BPlot begin parameters # Physical and geometric constants NA 6. 0 e 23 #Avogadro’s num f 0. 001 #scaling factor Vo f*1 e-10 # L V f*3 e-12 # L
Viewing comparison in Phi. BPlot
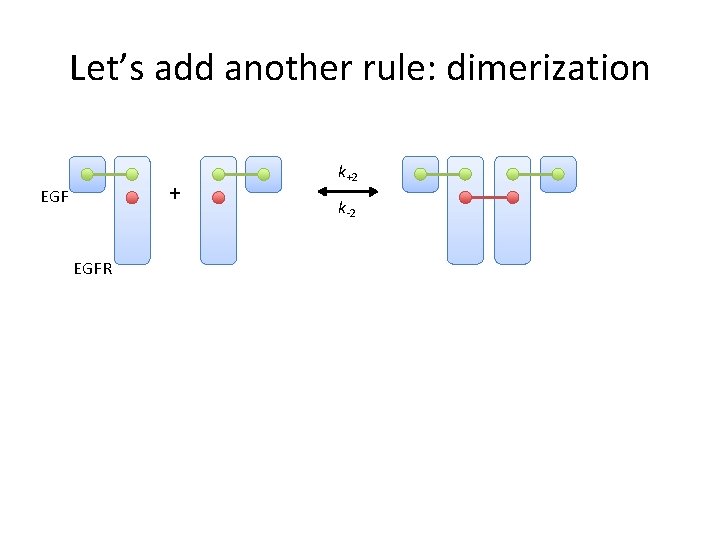
Let’s add another rule: dimerization + EGFR k+2 k-2
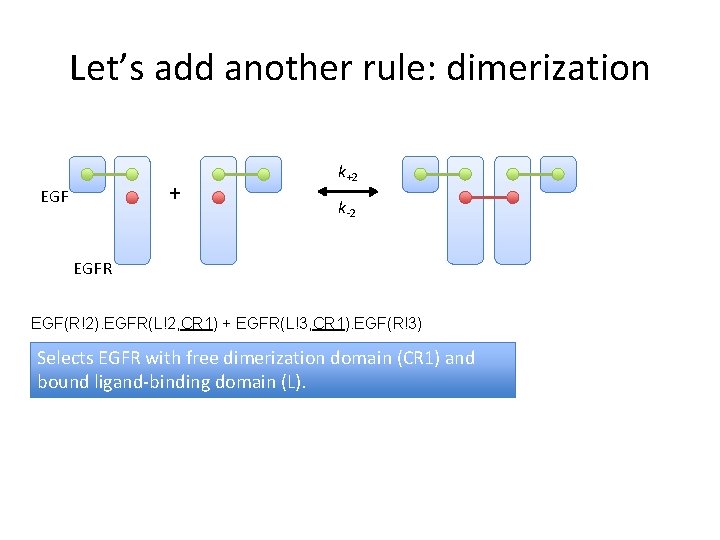
Let’s add another rule: dimerization + EGF k+2 k-2 EGFR EGF(R!2). EGFR(L!2, CR 1) + EGFR(L!3, CR 1). EGF(R!3) Selects EGFR with free dimerization domain (CR 1) and bound ligand-binding domain (L).
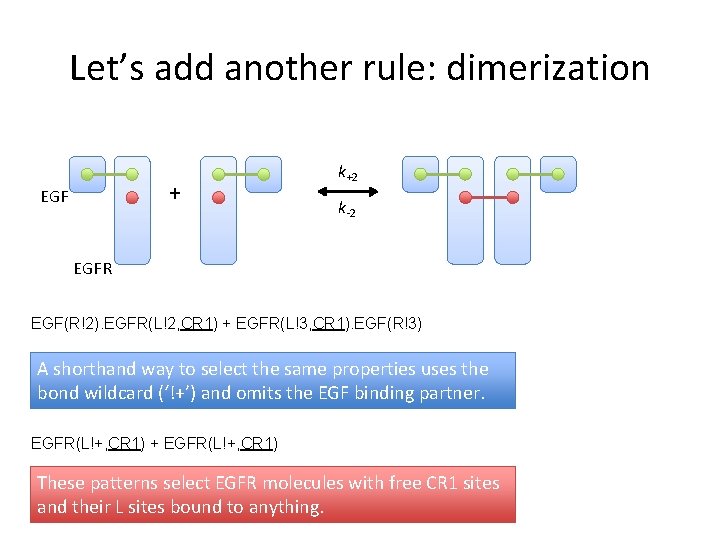
Let’s add another rule: dimerization + EGF k+2 k-2 EGFR EGF(R!2). EGFR(L!2, CR 1) + EGFR(L!3, CR 1). EGF(R!3) A shorthand way to select the same properties uses the bond wildcard (‘!+’) and omits the EGF binding partner. EGFR(L!+, CR 1) + EGFR(L!+, CR 1) These patterns select EGFR molecules with free CR 1 sites and their L sites bound to anything.
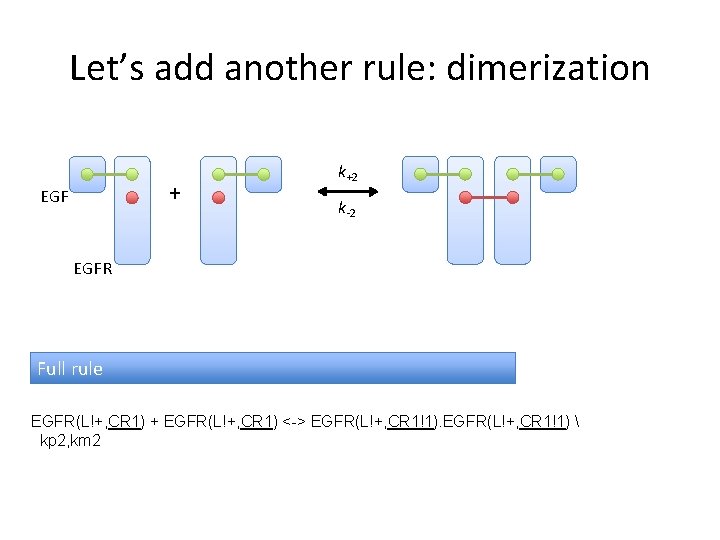
Let’s add another rule: dimerization + EGF k+2 k-2 EGFR Full rule EGFR(L!+, CR 1) + EGFR(L!+, CR 1) <-> EGFR(L!+, CR 1!1) kp 2, km 2
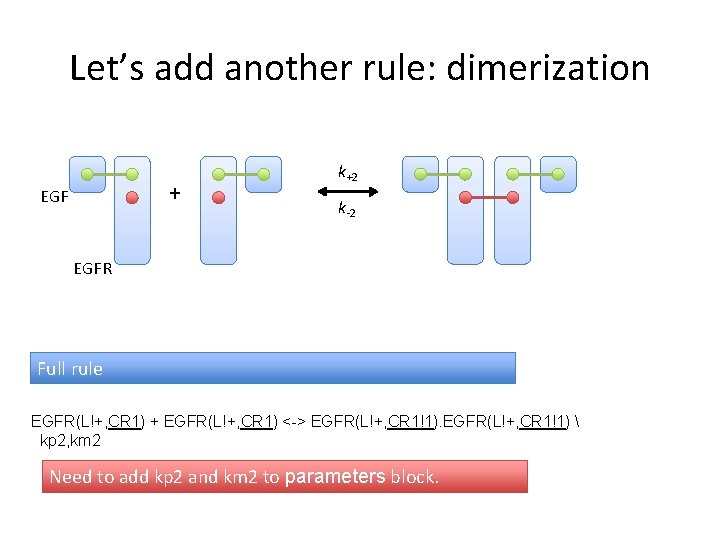
Let’s add another rule: dimerization + EGF k+2 k-2 EGFR Full rule EGFR(L!+, CR 1) + EGFR(L!+, CR 1) <-> EGFR(L!+, CR 1!1) kp 2, km 2 Need to add kp 2 and km 2 to parameters block.
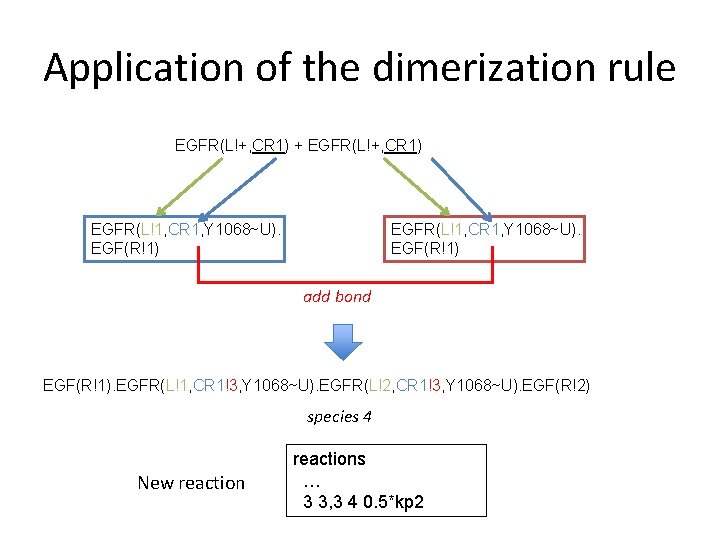
Application of the dimerization rule EGFR(L!+, CR 1) + EGFR(L!+, CR 1) EGFR(L!1, CR 1, Y 1068~U). EGF(R!1) add bond EGF(R!1). EGFR(L!1, CR 1!3, Y 1068~U). EGFR(L!2, CR 1!3, Y 1068~U). EGF(R!2) species 4 New reactions … 3 3, 3 4 0. 5*kp 2
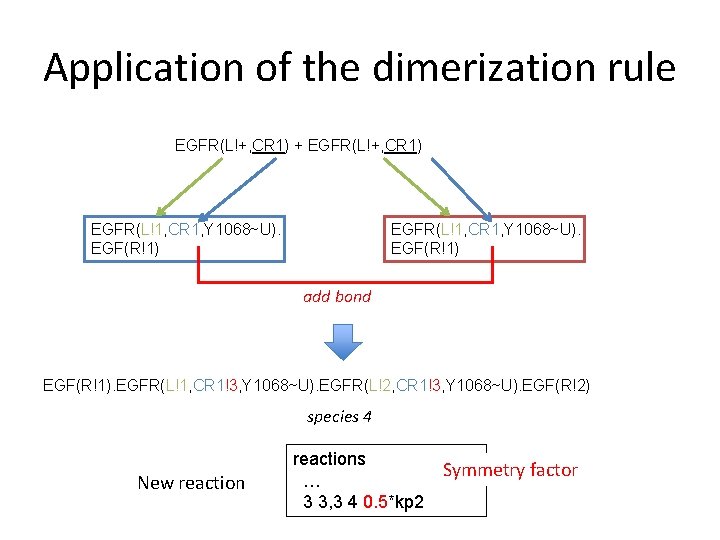
Application of the dimerization rule EGFR(L!+, CR 1) + EGFR(L!+, CR 1) EGFR(L!1, CR 1, Y 1068~U). EGF(R!1) add bond EGF(R!1). EGFR(L!1, CR 1!3, Y 1068~U). EGFR(L!2, CR 1!3, Y 1068~U). EGF(R!2) species 4 New reactions … 3 3, 3 4 0. 5*kp 2 Symmetry factor
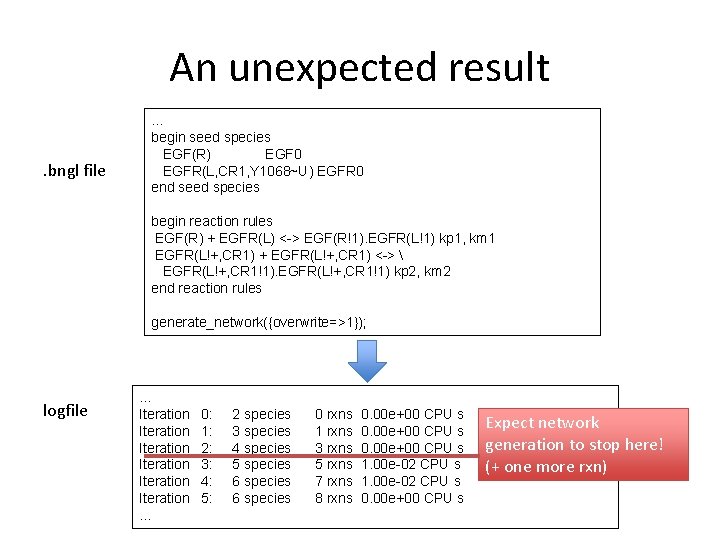
An unexpected result. bngl file … begin seed species EGF(R) EGF 0 EGFR(L, CR 1, Y 1068~U) EGFR 0 end seed species begin reaction rules EGF(R) + EGFR(L) <-> EGF(R!1). EGFR(L!1) kp 1, km 1 EGFR(L!+, CR 1) + EGFR(L!+, CR 1) <-> EGFR(L!+, CR 1!1) kp 2, km 2 end reaction rules generate_network({overwrite=>1}); logfile … Iteration Iteration … 0: 1: 2: 3: 4: 5: 2 species 3 species 4 species 5 species 6 species 0 rxns 1 rxns 3 rxns 5 rxns 7 rxns 8 rxns 0. 00 e+00 CPU s 1. 00 e-02 CPU s 0. 00 e+00 CPU s Expect network generation to stop here! (+ one more rxn)
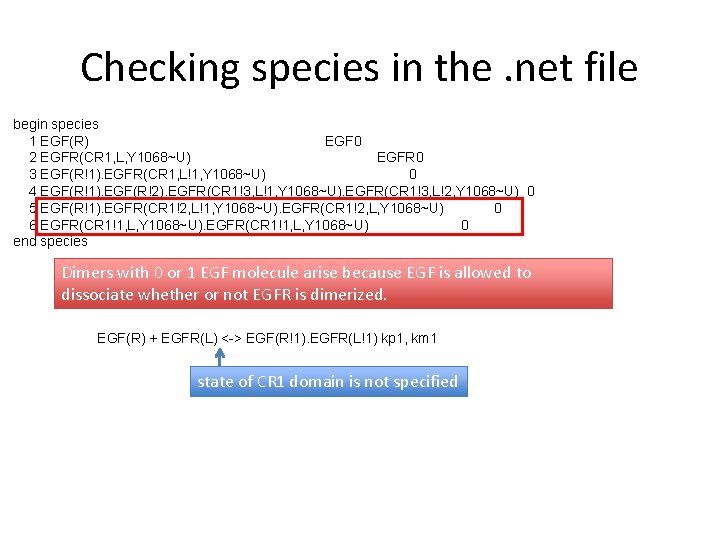
Checking species in the. net file begin species 1 EGF(R) EGF 0 2 EGFR(CR 1, L, Y 1068~U) EGFR 0 3 EGF(R!1). EGFR(CR 1, L!1, Y 1068~U) 0 4 EGF(R!1). EGF(R!2). EGFR(CR 1!3, L!1, Y 1068~U). EGFR(CR 1!3, L!2, Y 1068~U) 0 5 EGF(R!1). EGFR(CR 1!2, L!1, Y 1068~U). EGFR(CR 1!2, L, Y 1068~U) 0 6 EGFR(CR 1!1, L, Y 1068~U) 0 end species Dimers with 0 or 1 EGF molecule arise because EGF is allowed to dissociate whether or not EGFR is dimerized. EGF(R) + EGFR(L) <-> EGF(R!1). EGFR(L!1) kp 1, km 1 state of CR 1 domain is not specified

Checking species in the. net file begin species 1 EGF(R) EGF 0 2 EGFR(CR 1, L, Y 1068~U) EGFR 0 3 EGF(R!1). EGFR(CR 1, L!1, Y 1068~U) 0 4 EGF(R!1). EGF(R!2). EGFR(CR 1!3, L!1, Y 1068~U). EGFR(CR 1!3, L!2, Y 1068~U) 0 5 EGF(R!1). EGFR(CR 1!2, L!1, Y 1068~U). EGFR(CR 1!2, L, Y 1068~U) 0 6 EGFR(CR 1!1, L, Y 1068~U) 0 end species Dimers with 0 or 1 EGF molecule arise because EGF is allowed to dissociate whether or not EGFR is dimerized. EGF(R) + EGFR(L) <-> EGF(R!1). EGFR(L!1) kp 1, km 1 state of CR 1 domain is not specified “Expected” behavior can be recovered by modifying the rule: EGF(R) + EGFR(L, CR 1) <-> EGF(R!1). EGFR(L!1, CR 1) kp 1, km 1 Only EGFR with free CR 1 will bind and release ligand.

Creating an observable: Receptors in dimers Like rules, Observables select species based on pattern matching. EGFR(CR 1!+) matches receptors in dimers EGF(R!1). EGF(R!2). EGFR(CR 1!3, L!1, Y 1068~U). EGFR(CR 1!3, L!2, Y 1068~U) The concentrations of the matching species are stored in a variable with a specified name, computed by summing the concentrations of matching species. begin observables Molecules Rdim EGFR(CR 1!+) end observables
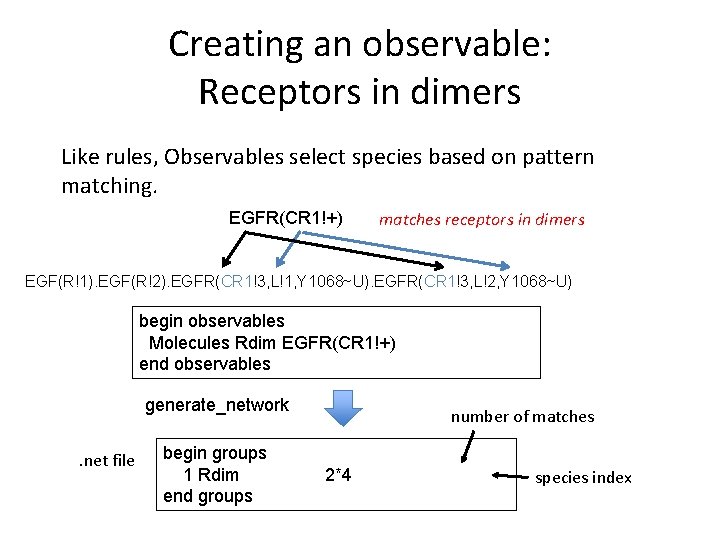
Creating an observable: Receptors in dimers Like rules, Observables select species based on pattern matching. EGFR(CR 1!+) matches receptors in dimers EGF(R!1). EGF(R!2). EGFR(CR 1!3, L!1, Y 1068~U). EGFR(CR 1!3, L!2, Y 1068~U) begin observables Molecules Rdim EGFR(CR 1!+) end observables generate_network. net file begin groups 1 Rdim end groups number of matches 2*4 species index

The. gdat file # time 0. 0000 e+00 2. 40000000 e+00 4. 80000000 e+00 … Rdim 0. 0000 e+00 1. 32882737 e+00 5. 16753986 e+00
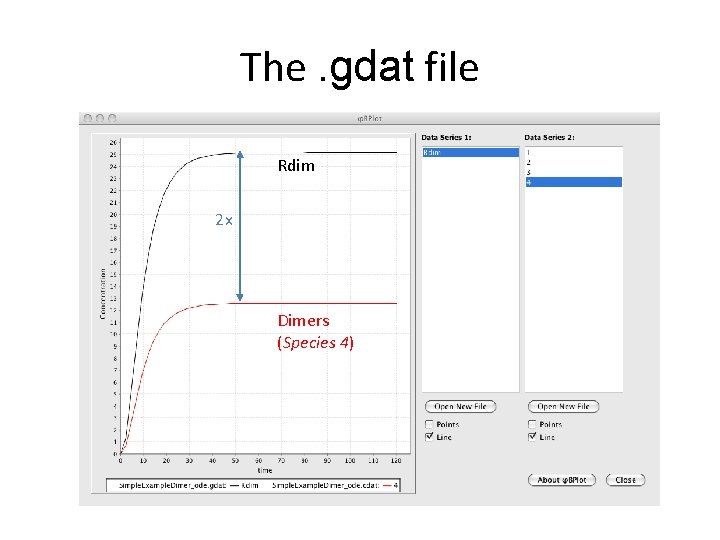
The. gdat file Rdim 2 x Dimers (Species 4)
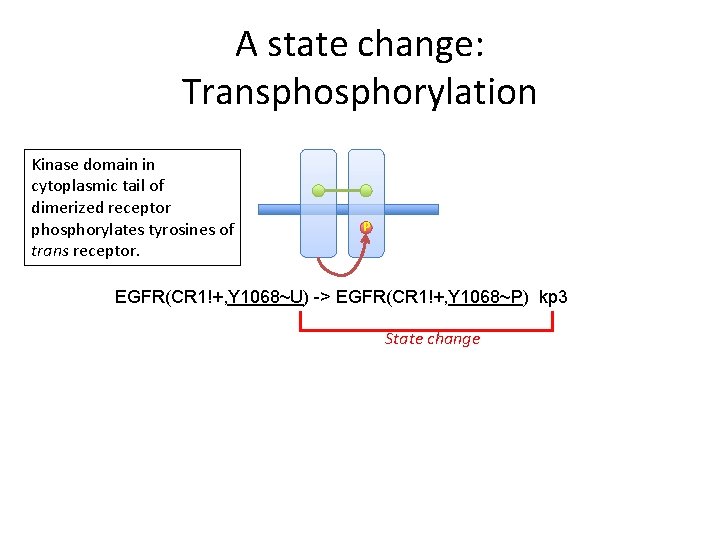
A state change: Transphorylation Kinase domain in cytoplasmic tail of dimerized receptor phosphorylates tyrosines of trans receptor. P EGFR(CR 1!+, Y 1068~U) -> EGFR(CR 1!+, Y 1068~P) kp 3 State change
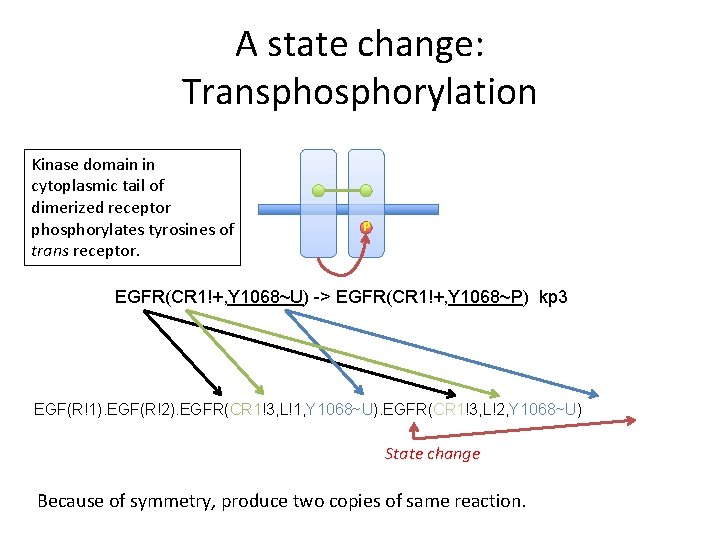
A state change: Transphorylation Kinase domain in cytoplasmic tail of dimerized receptor phosphorylates tyrosines of trans receptor. P EGFR(CR 1!+, Y 1068~U) -> EGFR(CR 1!+, Y 1068~P) kp 3 EGF(R!1). EGF(R!2). EGFR(CR 1!3, L!1, Y 1068~U). EGFR(CR 1!3, L!2, Y 1068~U) State change Because of symmetry, produce two copies of same reaction.

Reaction multiplicity Transphorylation rule generates EGF(R!1). EGF(R!2). EGFR(CR 1!3, L!1, Y 1068~U). EGFR(CR 1!3, L!2, Y 1068~U) Species 4 kp 3 EGF(R!1). EGF(R!2). EGFR(CR 1!3, L!1, Y 1068~P). EGFR(CR 1!3, L!2, Y 1068~U) Species 5 and EGF(R!1). EGF(R!2). EGFR(CR 1!3, L!1, Y 1068~U). EGFR(CR 1!3, L!2, Y 1068~U) Species 4 kp 3 EGF(R!1). EGF(R!2). EGFR(CR 1!3, L!1, Y 1068~U). EGFR(CR 1!3, L!2, Y 1068~P) Species 5 which are combined into reactions … 5 4 5 2*kp 3 #Rule 3

Dephosphorylation Phosphatase excess gives rise to first order dephosphorylation process. P U EGFR(Y 1068~P) -> EGFR(Y 1068~U) km 3 Note the convention that a component without a specified bonding state is required to be unbound. Dephosphorylation can occur only if the site is unbound. Binding of another protein to the site protects it from dephosphorylation.

Monitoring site phosphorylation Correct way to specify observable for receptor phosphorylation observables Molecules RP EGFR(Y 1068~P!? ) selects site in state ‘P’ wildcard allows site to be bound or unbound

Monitoring site phosphorylation Correct way to specify observable for receptor phosphorylation observables Molecules RP EGFR(Y 1068~P!? ) Incorrect way to specify observable for receptor phosphorylation observables Molecules RP EGFR(Y 1068~P) Only monitors sites that are phosphorylated and unbound
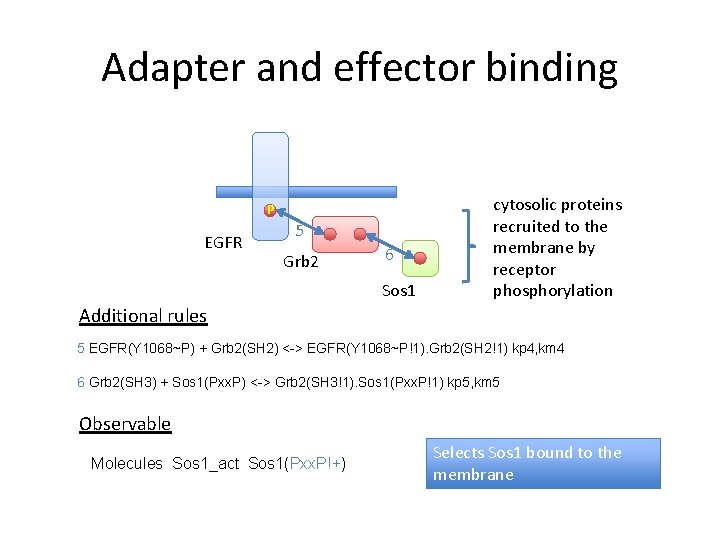
Adapter and effector binding P EGFR 5 Grb 2 6 Sos 1 cytosolic proteins recruited to the membrane by receptor phosphorylation Additional rules 5 EGFR(Y 1068~P) + Grb 2(SH 2) <-> EGFR(Y 1068~P!1). Grb 2(SH 2!1) kp 4, km 4 6 Grb 2(SH 3) + Sos 1(Pxx. P) <-> Grb 2(SH 3!1). Sos 1(Pxx. P!1) kp 5, km 5 Observable Molecules Sos 1_act Sos 1(Pxx. P!+) Selects Sos 1 bound to the membrane
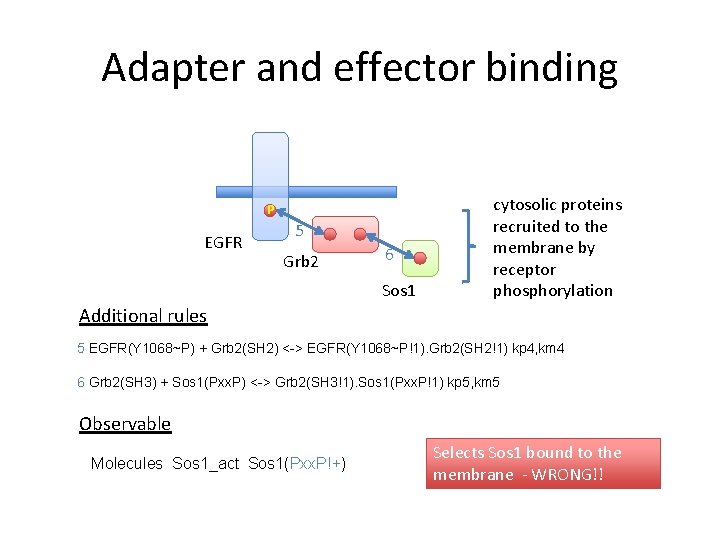
Adapter and effector binding P EGFR 5 Grb 2 6 Sos 1 cytosolic proteins recruited to the membrane by receptor phosphorylation Additional rules 5 EGFR(Y 1068~P) + Grb 2(SH 2) <-> EGFR(Y 1068~P!1). Grb 2(SH 2!1) kp 4, km 4 6 Grb 2(SH 3) + Sos 1(Pxx. P) <-> Grb 2(SH 3!1). Sos 1(Pxx. P!1) kp 5, km 5 Observable Molecules Sos 1_act Sos 1(Pxx. P!+) Selects Sos 1 bound to the membrane - WRONG!!
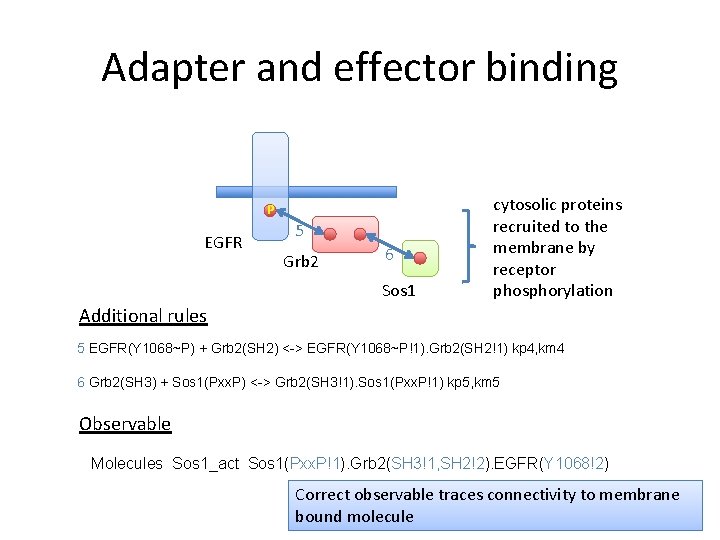
Adapter and effector binding P EGFR 5 Grb 2 6 Sos 1 cytosolic proteins recruited to the membrane by receptor phosphorylation Additional rules 5 EGFR(Y 1068~P) + Grb 2(SH 2) <-> EGFR(Y 1068~P!1). Grb 2(SH 2!1) kp 4, km 4 6 Grb 2(SH 3) + Sos 1(Pxx. P) <-> Grb 2(SH 3!1). Sos 1(Pxx. P!1) kp 5, km 5 Observable Molecules Sos 1_act Sos 1(Pxx. P!1). Grb 2(SH 3!1, SH 2!2). EGFR(Y 1068!2) Correct observable traces connectivity to membrane bound molecule

A degradation reaction begin molecule types … Null() … begin seed species … $Null Simple receptor degradation begin reaction rules … EGFR() -> Null kdeg Deletes any species containing EGFR molecule, including any attached molecules.

A degradation reaction begin molecule types … Null() … begin seed species … $Null Selective degradation of ligand receptor begin reaction rules … EGF(R!1). EGF(R!2). EGFR(L!1, CR 1!3). EGFR(L!2, CR 1!3) -> Trash() kdeg Delete. Molecules This rule causes just the ligand-bound receptor dimer to be degraded, recycling any components that are bound to the cytoplasmic domains of the receptor.
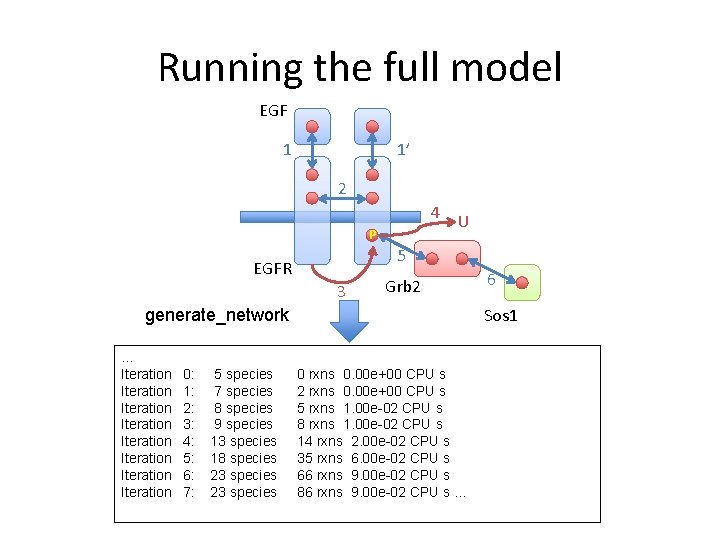
Running the full model EGF 1 1’ 2 4 U P 5 EGFR 3 Grb 2 Sos 1 generate_network … Iteration Iteration 0: 1: 2: 3: 4: 5: 6: 7: 5 species 7 species 8 species 9 species 13 species 18 species 23 species 6 0 rxns 0. 00 e+00 CPU s 2 rxns 0. 00 e+00 CPU s 5 rxns 1. 00 e-02 CPU s 8 rxns 1. 00 e-02 CPU s 14 rxns 2. 00 e-02 CPU s 35 rxns 6. 00 e-02 CPU s 66 rxns 9. 00 e-02 CPU s 86 rxns 9. 00 e-02 CPU s …
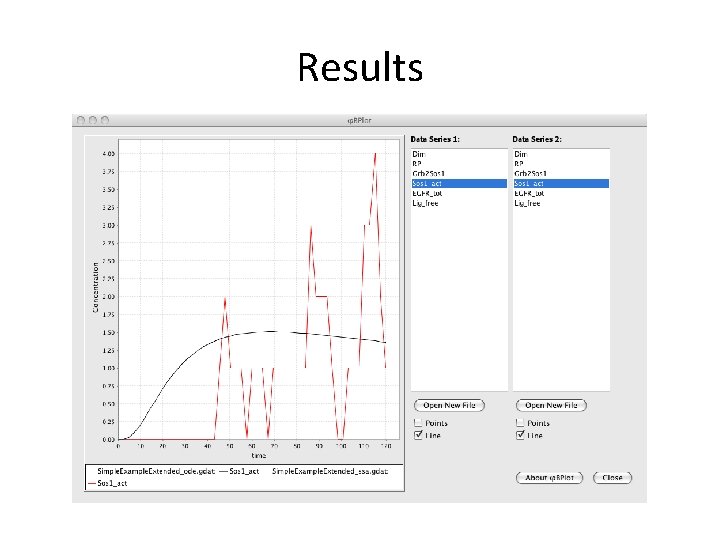
Results

Equilibrating Grb 2 -Sos 1 binding prior to ligand addition Grb 2 -Sos 1 interaction is constitutive Grb 2 6 Sos 1

Equilibrating Grb 2 -Sos 1 binding prior to ligand addition Grb 2 -Sos 1 interaction is constitutive Grb 2 6 Sos 1 Combination of actions can be used to pre-equilibrate prior to ligand addition set. Concentration("EGF(R)", 0); simulate_ode({suffix=>equil, t_end=>10000, n_steps=>50, steady_state=>1, sparse=>1}); set. Concentration("EGF(R)", EGF 0);

Complete summary of actions Action/parameter Type generate_network Description Default Generate species and reactions through iterative application of rules to seed species max_agg int Maximum number of molecules in one species 1 e 99 max_iter int Maximum number of iterations of rule application 100 max_stoich hash Maximum number of molecules of specified type in one species - overwrite 0/1 Overwrite existing. net file 0 (off) print_iter 0/1 Print. net file after each iteration 0 prefixa string Set basename c of. net file to string basename of. bngl file suffixa string Append _string to basename of. net file - simulate_ode / simulate_ssa Simulate current model/network t_end float End time for simulation required t_start float Start time for simulation 0 n_steps int Number of times after t=0 at which to report concentrations/observables 1 sample_times array Times at which to report concentrations/observables (supercedes t_end, n_steps) - netfile string Name of. net file used for simulation - atolb float Absolute error tolerance for ODE’s 1 e-8 rtolb float Relative error tolerance for ODE’s 1 e-8 steady_stateb 0/1 Perform steady state check on species concentrations 0 sparseb 0/1 Use sparse Jacobian / iterative solver (GMRES) in CVODE 0 read. File file Read a. bngl or a. net file string Name of file to read required write. NET / write. SBML / write. Mfile Write current model/network in specified format set. Concentration(species, value) Set concentration of species to value. set. Parameter(parameter, value) Set parameter to value. save. Concentrations() Store current species concentrations. reset. Concentratons() Restore species concentrations to value at point of last save. Concentrations command. see http: //bionetgen. org for Bio. Net. Gen Chapter

Simulation results with and without equilibration Higher initial level of Grb 2 -Sos 1 complex No effect on Sos 1 recruitment

More topics • Parameter scans • Alternate rate laws • Labels – Fluorescent labeling – Carbon-fate maps • Oligomerization – simulate_nf New • Compartments • GETBONNIE

Detailed model of TLR 4 signaling • Main goals – Represent known players and interactions in TLR 4 mediated signaling – Does model qualitatively reproduce observed tolerance in response to repeated stimulation (“preconditioning”)? – Investigate molecular mechanisms leading to tolerance G. An and J. Faeder, Math. Biosci. , to appear.

Approach • Dynamic Knowledge Representation by nonmathematician • Relatively Detailed Representation of Components • Mechanistic/causal emphasis • Known source and stimulus for production of compound (particularly inhibitors) • Abstraction of complex molecular events if output relatively straightforward • Model parameters (component numbers and reaction rates) qualitatively set based on “realistic” levels

Detailed model of TLR 4 signaling
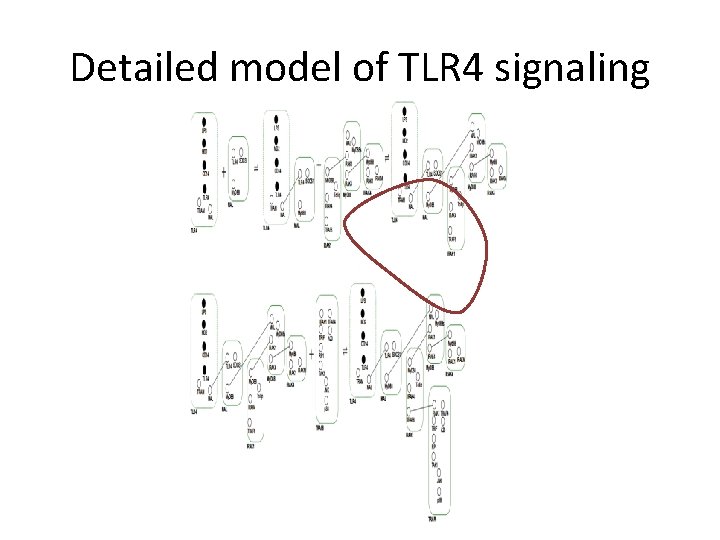
Detailed model of TLR 4 signaling
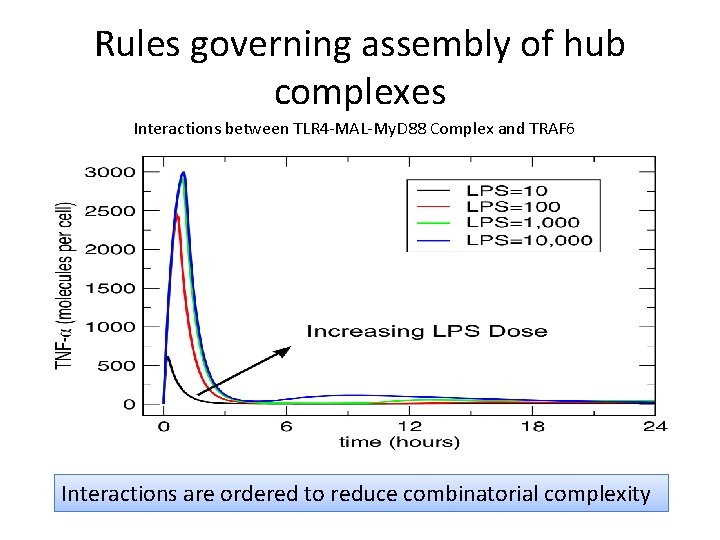
Rules governing assembly of hub complexes Interactions between TLR 4 -MAL-My. D 88 Complex and TRAF 6 Interactions are ordered to reduce combinatorial complexity

Regulation of A 20 expression Binding of NFk. B in the nucleus to A 20 promoter NFk. B(Transcription~No, Activation~Yes, Location~Nucleus)+DNA(A 20) <-> NFk. B(Transcription~Yes!1, Activation~Yes, Location~Nucleus). DNA(A 20!1) NFk. B_DNA_A 20_Bind, NFk. B_DNA_A 20_Unbind Transcription of A 20 gene DNA(A 20!0). NFk. B(Transcription~Yes!0, Activation~Yes, Location~Nucleus) -> DNA(A 20!0). NFk. B(Transcription~Yes!0, Activation~Yes, Location~Nucleus) + A 20 m. RNA(Translation~On) A 20_Transcription_Execute Translation of A 20 m. RNA(Translation~On) -> A 20 m. RNA(Translation~Off) + A 20(TRAF 6) A 20_Translation_Execute Degradation of A 20(TRAF 6)->Trash(c) A 20_Degrade

Generating the model /Users/faeder/shared/Projects/Bio. Net. Gen/Perl 2/BNG 2. pl Bio. Net. Gen version 2. 0. 47+ read. File: : Reading from file TLR 4_V 15_PC. bngl Read 97 parameters. Read 31 molecule types. Read 19 species. Read 41 reaction rule(s). Read 19 observable(s). Iteration 0: 19 species 0 rxns 0. 00 e+00 CPU s Iteration 1: 24 species 5 rxns 4. 00 e-02 CPU s Iteration 2: 27 species 12 rxns 3. 00 e-02 CPU s Iteration 3: 28 species 17 rxns 3. 00 e-02 CPU s Iteration 4: 29 species 19 rxns 1. 00 e-02 CPU s Iteration 5: 31 species 22 rxns 6. 00 e-02 CPU s Iteration 6: 38 species 32 rxns 1. 00 e-01 CPU s Iteration 7: 46 species 58 rxns 3. 60 e-01 CPU s Iteration 8: 53 species 96 rxns 5. 40 e-01 CPU s Iteration 9: 57 species 128 rxns 6. 00 e-01 CPU s Iteration 10: 59 species 145 rxns 4. 30 e-01 CPU s Iteration 11: 62 species 148 rxns 1. 00 e-02 CPU s Iteration 12: 65 species 153 rxns 2. 00 e-02 CPU s Iteration 13: 71 species 168 rxns 5. 00 e-02 CPU s Iteration 14: 76 species 192 rxns 8. 00 e-02 CPU s Iteration 15: 76 species 202 rxns 5. 00 e-02 CPU s
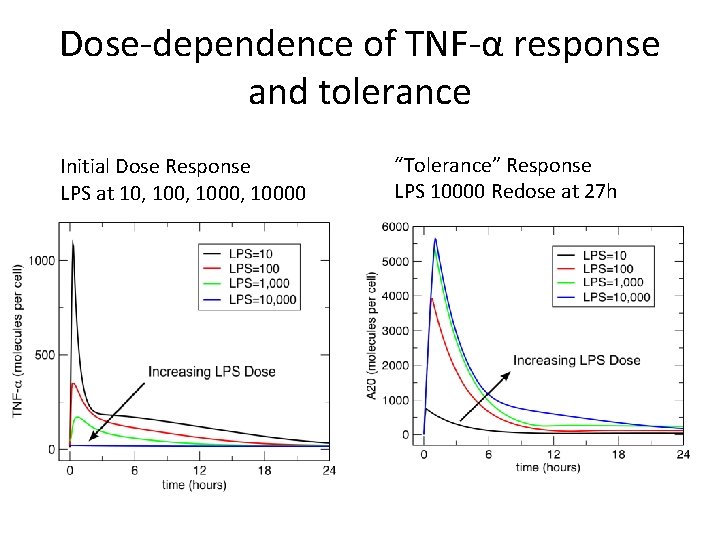
Dose-dependence of TNF-α response and tolerance Initial Dose Response LPS at 10, 1000, 10000 “Tolerance” Response LPS 10000 Redose at 27 h
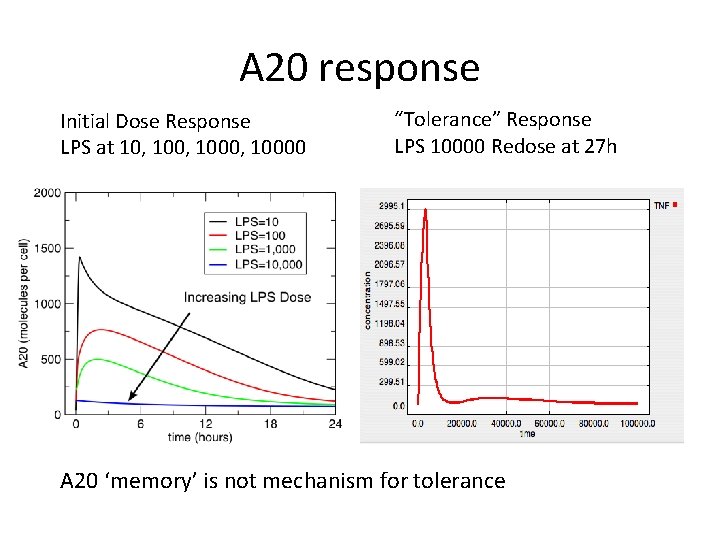
A 20 response Initial Dose Response LPS at 10, 1000, 10000 “Tolerance” Response LPS 10000 Redose at 27 h A 20 ‘memory’ is not mechanism for tolerance
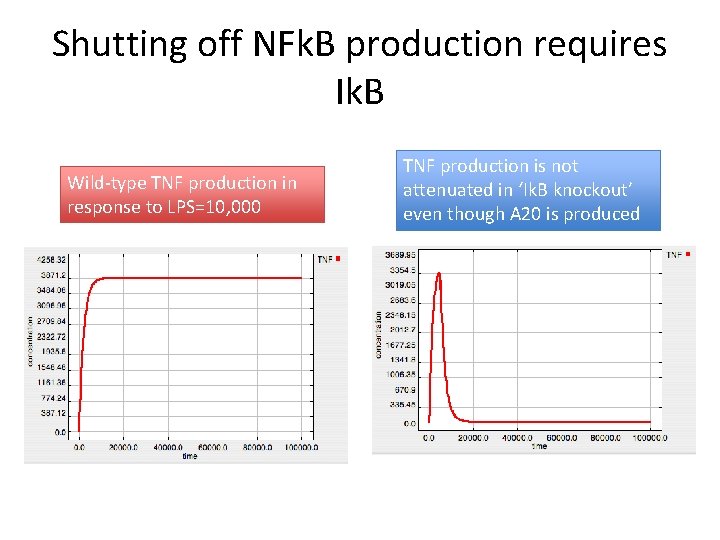
Shutting off NFk. B production requires Ik. B Wild-type TNF production in response to LPS=10, 000 TNF production is not attenuated in ‘Ik. B knockout’ even though A 20 is produced
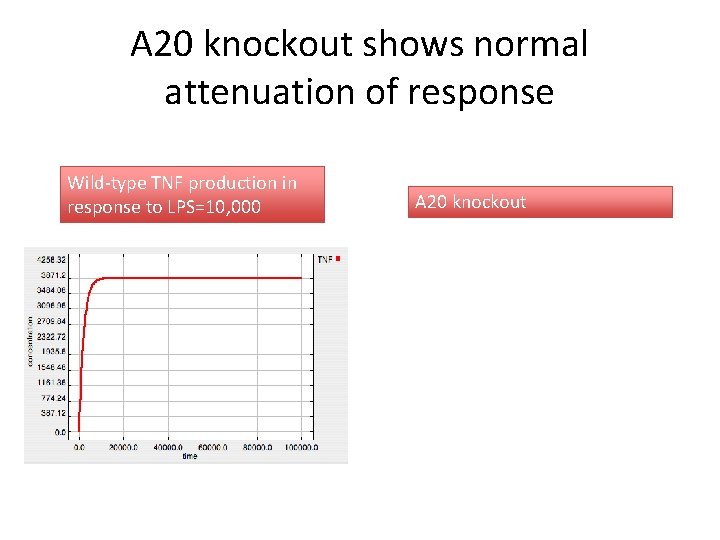
A 20 knockout shows normal attenuation of response Wild-type TNF production in response to LPS=10, 000 A 20 knockout
- Slides: 96