A biodiversityinspired approach to marine ecosystem modelling Jorn

A biodiversity-inspired approach to marine ecosystem modelling Jorn Bruggeman Bas Kooijman Theoretical biology Vrije Universiteit Amsterdam
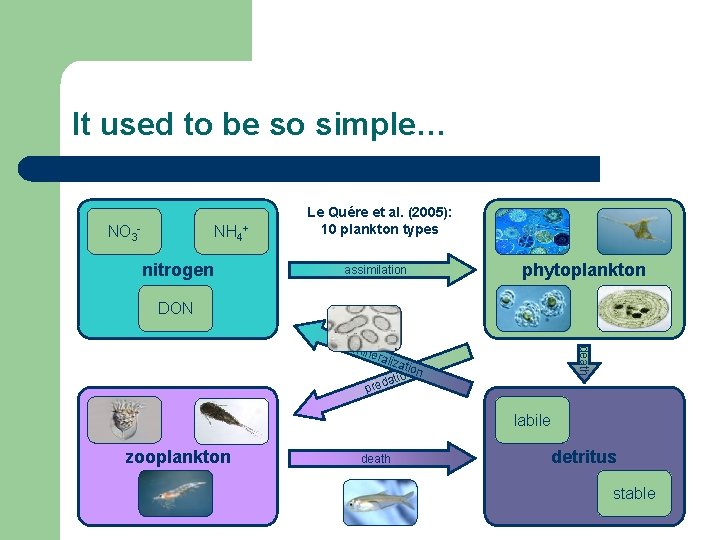
It used to be so simple… NO 3 - NH 4+ nitrogen Le Quére et al. (2005): 10 plankton types assimilation phytoplankton DON death min eral izat ion n o i t a d e r p labile zooplankton death detritus stable
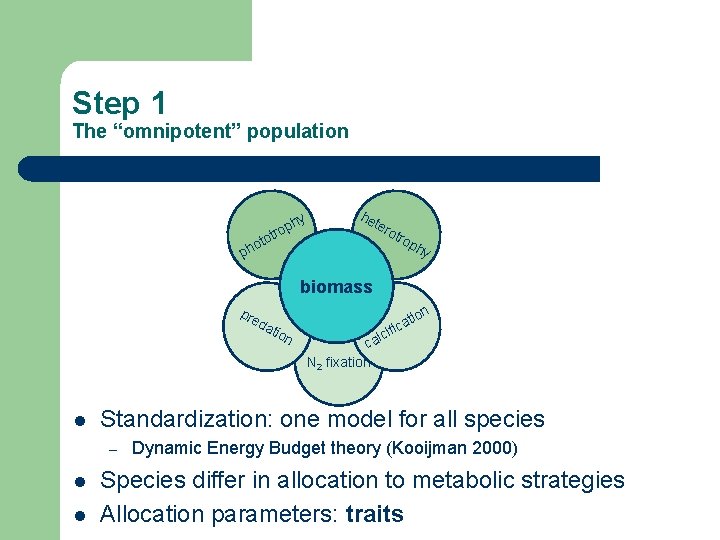
Step 1 The “omnipotent” population t o ph y ph o otr he ter otr op hy biomass pre da l l tio n lc ca N 2 fixation Standardization: one model for all species – l n tio a ific Dynamic Energy Budget theory (Kooijman 2000) Species differ in allocation to metabolic strategies Allocation parameters: traits
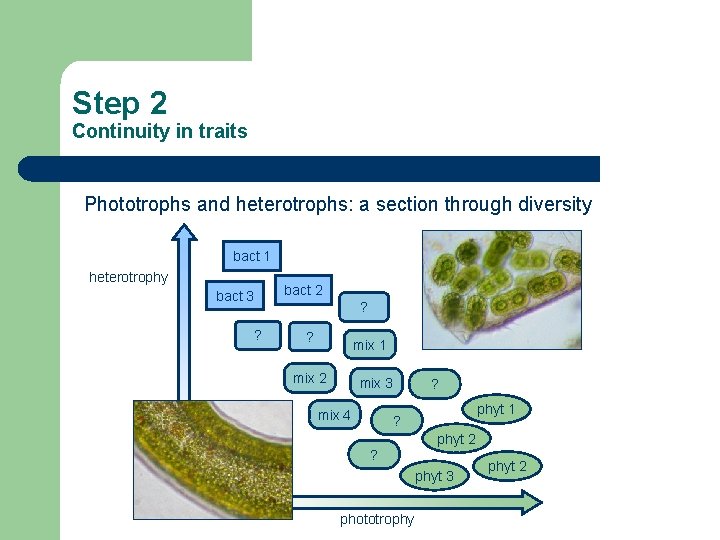
Step 2 Continuity in traits Phototrophs and heterotrophs: a section through diversity bact 1 heterotrophy bact 3 ? bact 2 ? ? mix 1 mix 2 mix 3 mix 4 ? phyt 1 ? phyt 2 ? phyt 3 phototrophy phyt 2

Step 3 “Everything is everywhere; the environment selects” l Every possible species present at all times – – l The environment changes because of – – l implementation: continuous immigration of trace amounts of all species similar to: minimum biomass (Burchard et al. 2006), constant variance of trait distribution (Wirtz & Eckhardt 1996) external forcing, e. g. periodicity of light, mixing ecosystem dynamics, e. g. depletion of nutrients Changing environment drives succession – – – niche presence = time- and space-dependent trait value combinations define species & niche trait distribution will change in space and time
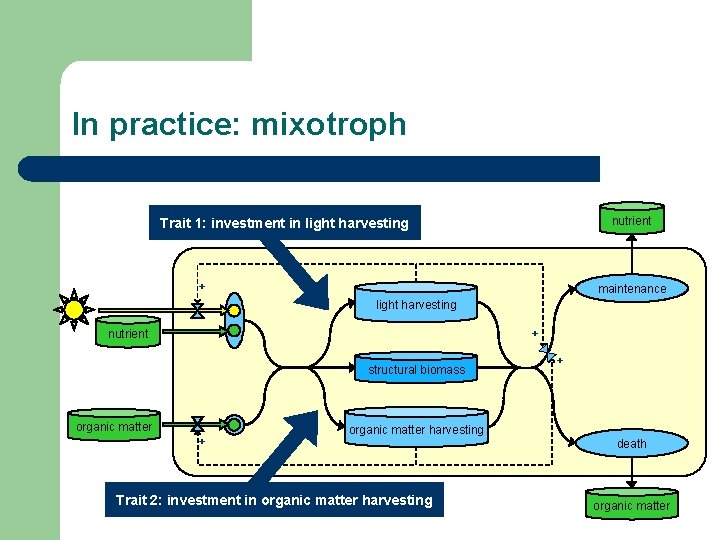
In practice: mixotroph nutrient Trait 1: investment in light harvesting + maintenance light harvesting nutrient + structural biomass organic matter + + organic matter harvesting Trait 2: investment in organic matter harvesting death organic matter

Setting l General Ocean Turbulence Model (GOTM) – – – l Scenario: Bermuda Atlantic Time-series Study (BATS) – – l 1 D water column depth- and time-dependent turbulent diffusivity k-ε turbulence model surface forcing from ERA-40 dataset initial state: observed depth profiles temperature/salinity Parameter fitting – – fitted internal wave parameterization to temperature profiles fitting biological parameters to observed depth profiles of chlorophyll and DIN simultaneously
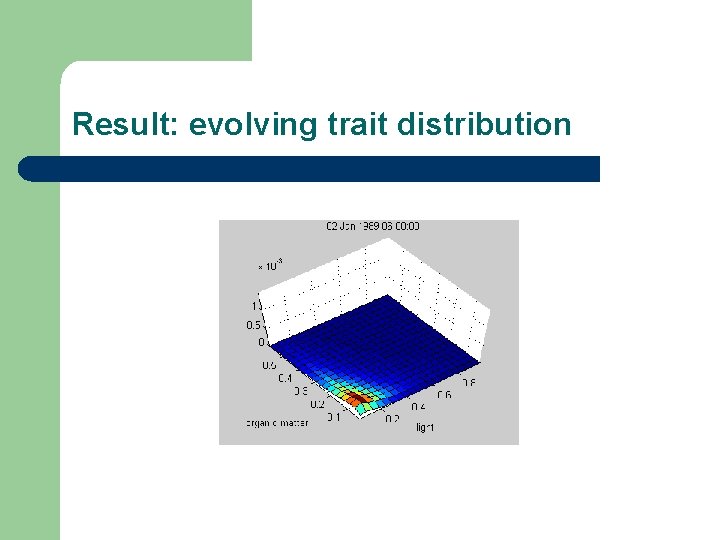
Result: evolving trait distribution
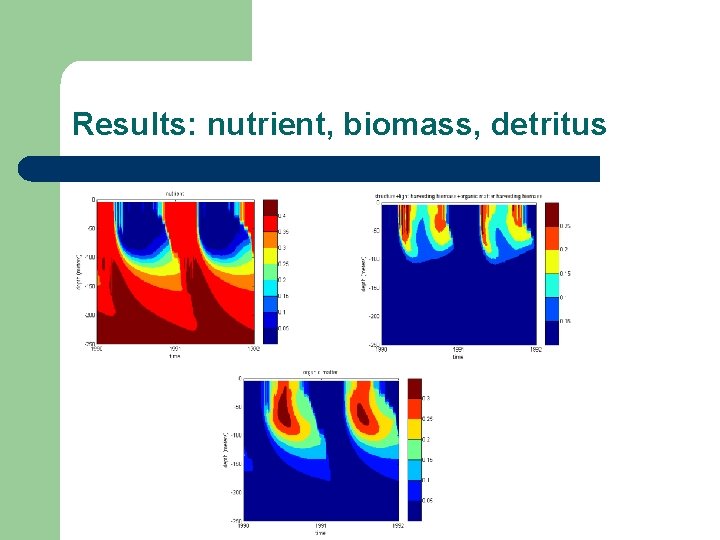
Results: nutrient, biomass, detritus
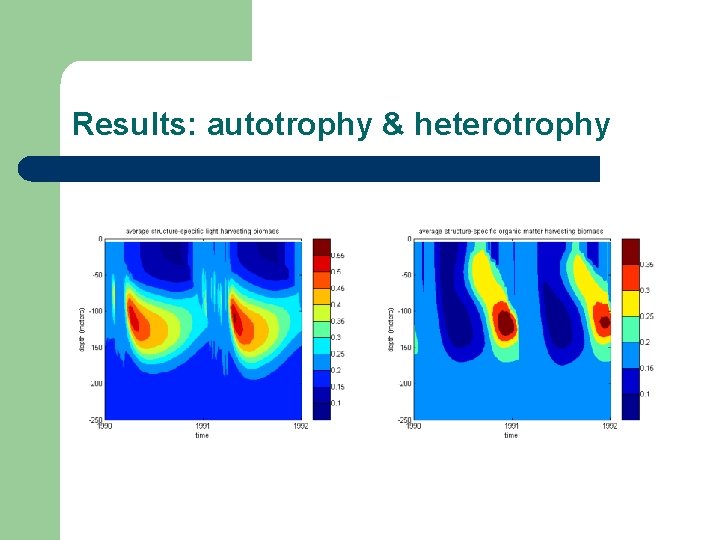
Results: autotrophy & heterotrophy

Simpler trait distributions 1. Before: “brute-force” – – – 2. 2 traits 20 x 20 grid = 400 state variables (‘species’) flexible: any distribution shape (multimodality) possible high computational cost Now: simplify via assumptions on distribution shape characterize trait distribution by moments: mean, variance, etc. 2. express higher moments in terms of first moments (moment closure) 3. evolve first moments E. g. 2 traits 2 x (mean, variance) = 4 state variables 1.
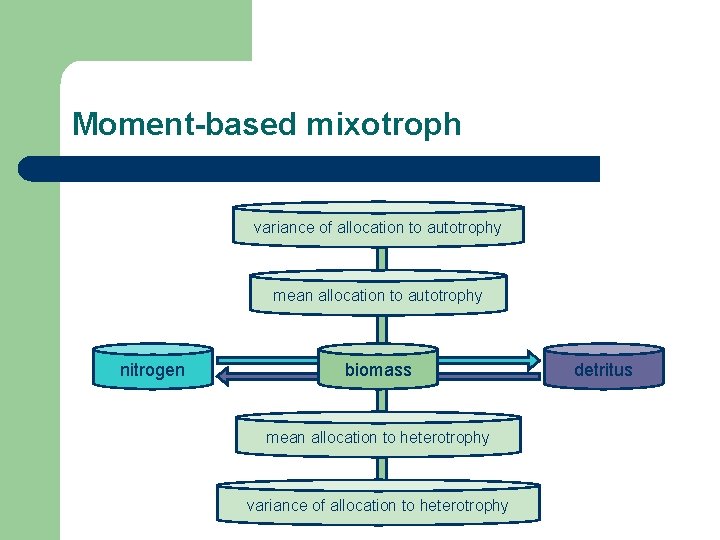
Moment-based mixotroph variance of allocation to autotrophy mean allocation to autotrophy nitrogen biomass mean allocation to heterotrophy variance of allocation to heterotrophy detritus
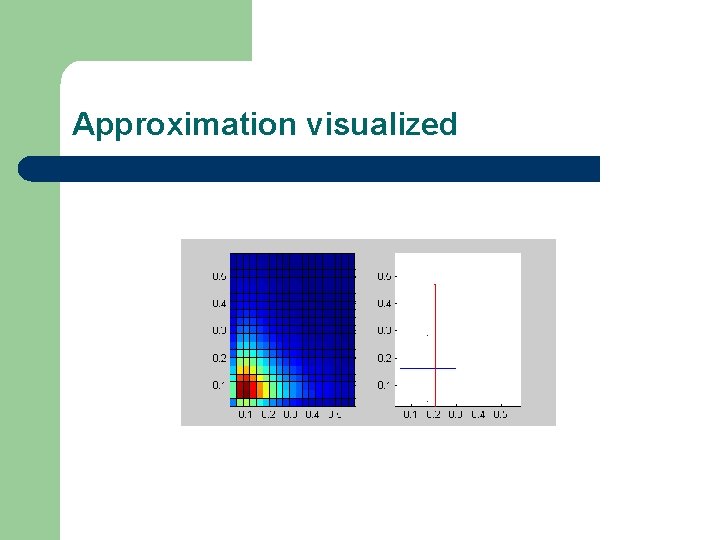
Approximation visualized
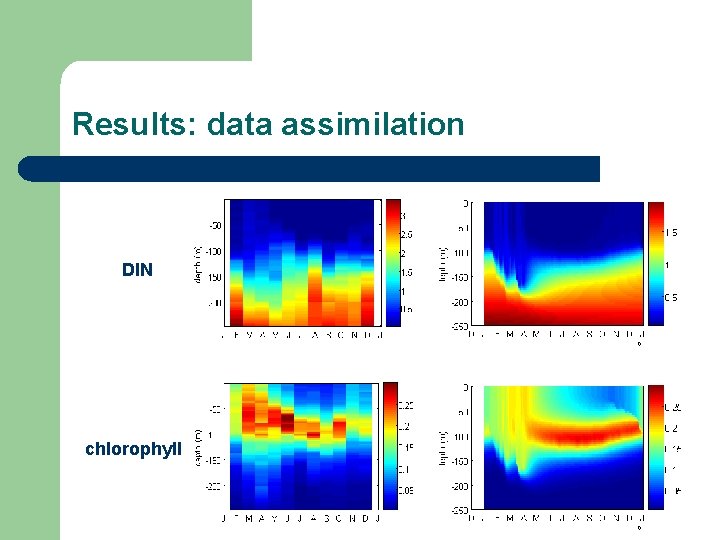
Results: data assimilation DIN chlorophyll

Conclusions l Simple mixotroph + biodiversity model shows – – – l Time-dependent species composition: autotrophic species (e. g. diatoms) replaced by mixotrophic/heterotrophic species (e. g. dinoflagellates) Depth-dependent species composition: subsurface chlorophyll maximum Good description of BATS chlorophyll and DIN Modeled biodiversity adds flexibility “in a good way”: – – Moments represent biodiversity mechanistic derivation, not ad-hoc Direct (measurable) implications for mass- and energy balances

Outlook l Selection of traits, e. g. – – l Biodiversity-based ecosystem models – l Metabolic strategies Individual size Rich dynamics through succession rather than physiological detail Use of biodiversity indicators (variance of traits) – – Effect of biodiversity on ecosystem functioning Effect of external factors (eutrophication, toxicants) on diversity
- Slides: 16