A A 2016 2017 CORSO DI BIOINFORMATICA 2

A. A. 2016 -2017 CORSO DI BIOINFORMATICA 2 per il CLM in BIOLOGIA EVOLUZIONISTICA Scuola di Scienze, Università di Padova Docente: Prof. Stefania Bortoluzzi

Outline • Transcriptomics today • RNA-seq features and advantages • Transcriptome sequencing goals • Analysis pipelines

Transcriptome definition and dynamics QUALITY QUANTITY The set of all RNA molecules Amount/concentration information produced in one cell, or in a is needed in addition to molecule population of cells, at a given time identification Identification of expressed RNA molecules Expression level The transcriptome continuosly changes • Development • Differentiation • Response to environment • Disease Transcriptome variation depends on: • Genetic/epigenetic changes of the genome • Emergent properties of the transcriptome itself as a complex system • Many regulatory RNAs • Many layers of genes expression regulation
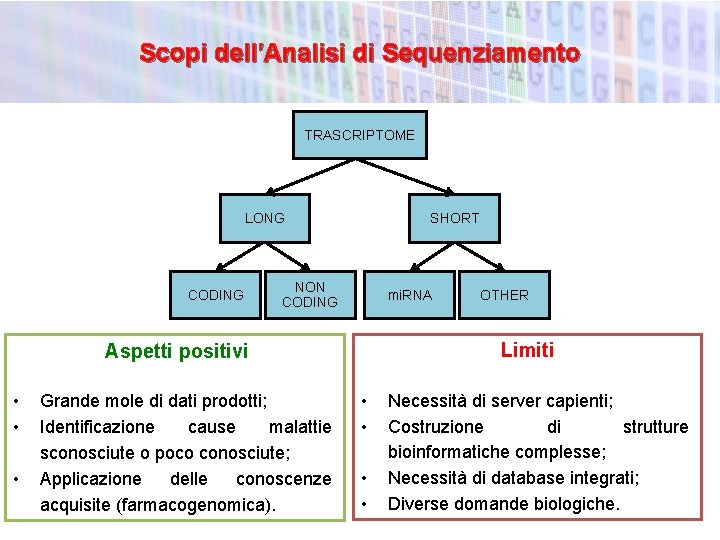
Scopi dell’Analisi di Sequenziamento TRASCRIPTOME LONG CODING SHORT NON CODING mi. RNA Limiti Aspetti positivi • • • Grande mole di dati prodotti; Identificazione cause malattie sconosciute o poco conosciute; Applicazione delle conoscenze acquisite (farmacogenomica). OTHER • • Necessità di server capienti; Costruzione di strutture bioinformatiche complesse; Necessità di database integrati; Diverse domande biologiche.

Genoma • < 3% del genoma umano codifica proteine • Evidenze recenti ottenute con genomic-tiling array e sequenziamento del trascrittoma hanno mostrato che >70% del genoma è trascritto in maniera pervasiva in • RNA codificante ( m. RNA) • Moltissimi prodotti trascrizionali sono RNA, piccoli e lunghi, con scarsissimo potenziale codificante La maggior parte del DNA eucariotico trascritto è non codificante • La production phase di ENCODE ha mostrato che >80% del genoma è biologicamente attivo e funzionale (ruolo regolativo per la maggior parte delle sequenze)

Il trascrittoma non-codificante • RNA non codificanti noti da molto tempo: r. RNA e t. RNA nella traduzione § sn. RNA e sno. RNA nel processamento degli m. RNA § § ribozimi • Ipotesi del “mondo ad RNA” u Le molecole di RNA possono contemporaneamente contenere informazione di sequenza e possedere plasticità strutturale u Gli RNA possono sia interagire con DNA ed altri RNA per appaiamento delle basi complementari, sia fornire siti di legame per proteine
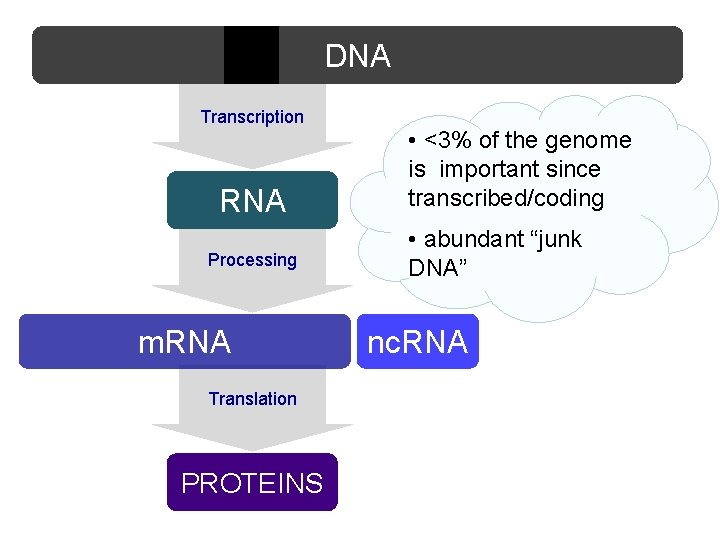
DNA Transcription RNA Processing m. RNA Translation PROTEINS • <3% of the genome is important since transcribed/coding • abundant “junk DNA” nc. RNA
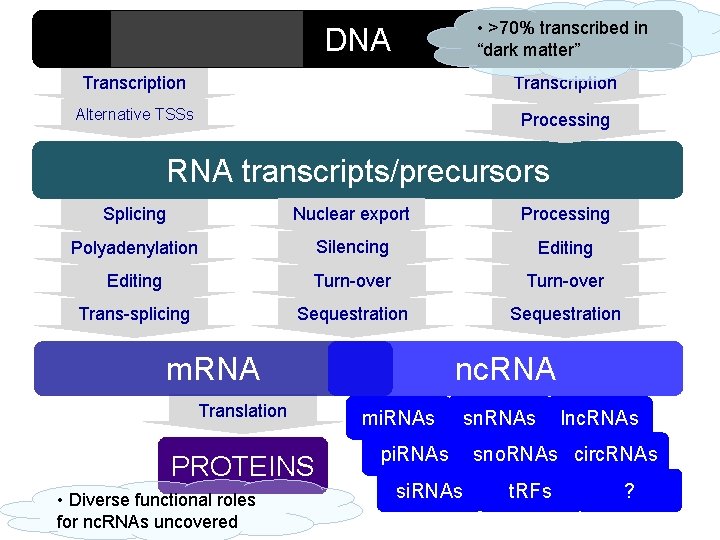
• >70% transcribed in “dark matter” DNA Transcription Alternative TSSs Processing RNA transcripts/precursors Splicing Nuclear export Processing Polyadenylation Silencing Editing Turn-over Trans-splicing Sequestration m. RNA Translation PROTEINS • Diverse functional roles for nc. RNAs uncovered nc. RNA mi. RNAs pi. RNAs sn. RNAs lnc. RNAs sno. RNAs circ. RNAs t. RFs ?
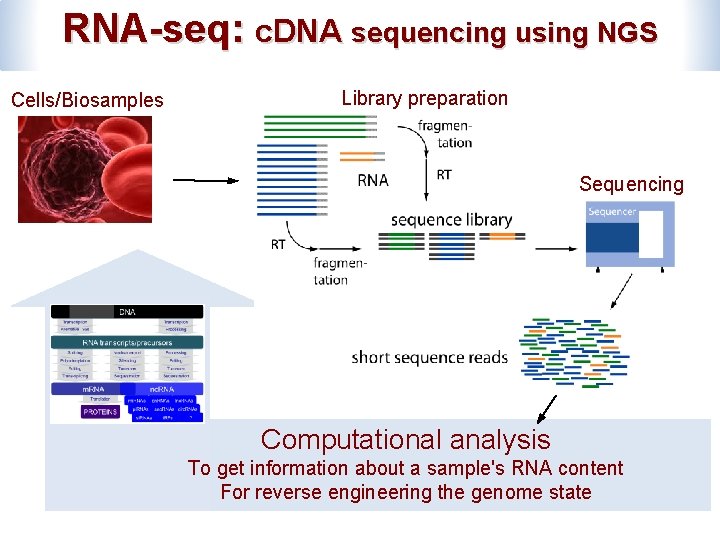
RNA-seq: c. DNA sequencing using NGS Cells/Biosamples Library preparation Sequencing Computational analysis To get information about a sample's RNA content For reverse engineering the genome state
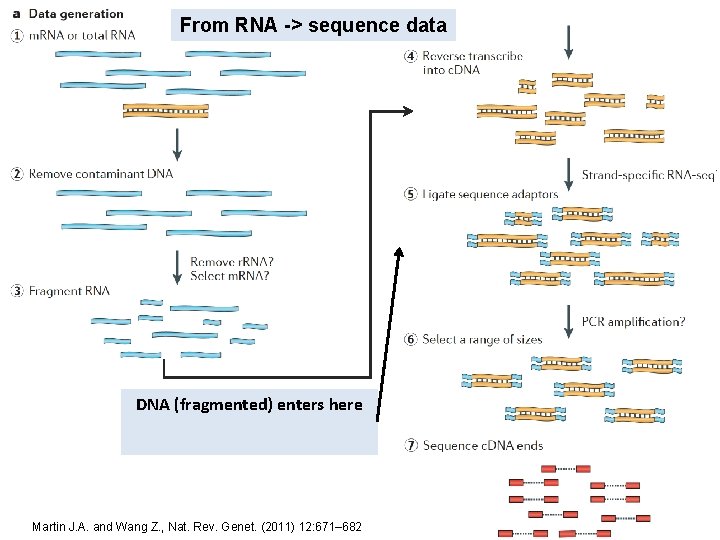
From RNA -> sequence data DNA (fragmented) enters here Martin J. A. and Wang Z. , Nat. Rev. Genet. (2011) 12: 671– 682
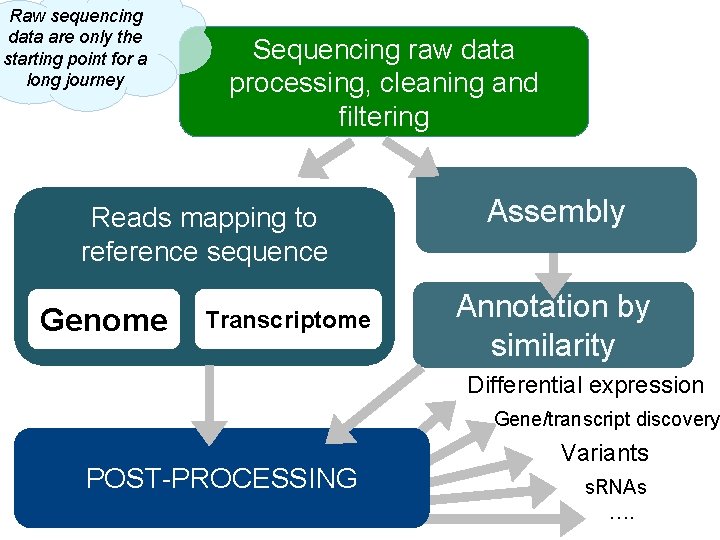
Raw sequencing data are only the starting point for a long journey Sequencing raw data processing, cleaning and filtering Reads mapping to reference sequence Genome Transcriptome Assembly Annotation by similarity Differential expression Gene/transcript discovery POST-PROCESSING Variants s. RNAs ….
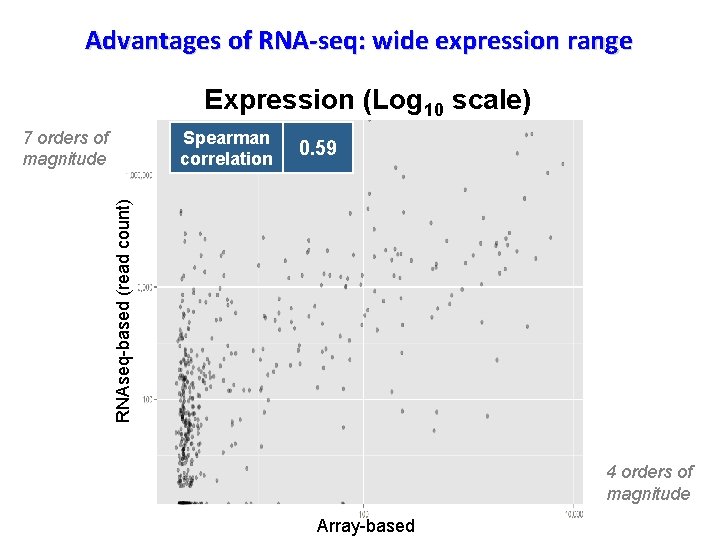
Advantages of RNA-seq: wide expression range Expression (Log 10 scale) Spearman correlation 0. 59 RNAseq-based (read count) 7 orders of magnitude 4 orders of magnitude Array-based
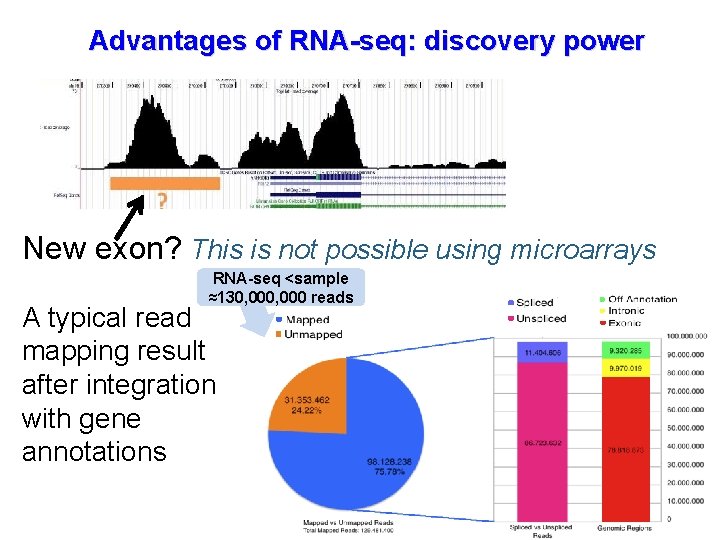
Advantages of RNA-seq: discovery power New exon? This is not possible using microarrays RNA-seq <sample ≈130, 000 reads A typical read mapping result after integration with gene annotations
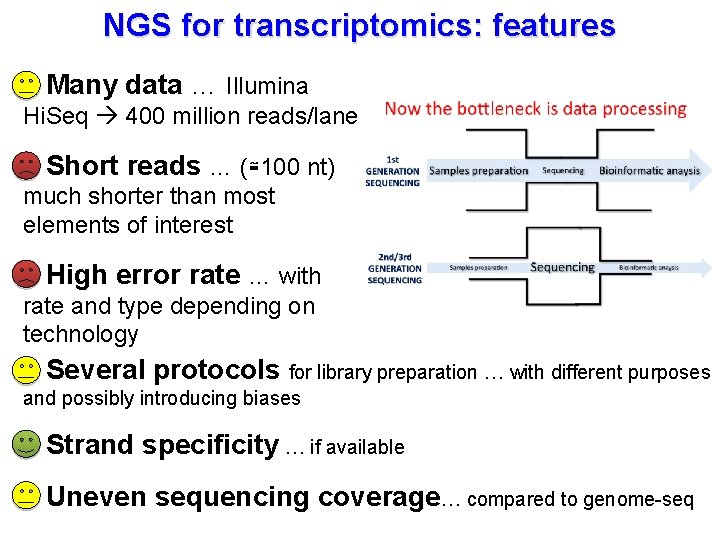
NGS for transcriptomics: features Many data … Illumina Hi. Seq 400 million reads/lane Short reads … (≅100 nt) much shorter than most elements of interest High error rate … with rate and type depending on technology Several protocols for library preparation … with different purposes and possibly introducing biases Strand specificity … if available Uneven sequencing coverage… compared to genome-seq

RNA-seq comes in different flavours • Experimental Design • Poly(A) enrichment or ribosomal RNA depletion? • Single-end or Paired end? • Stranded or not? • How much sequencing data to collect?

Poly(A) enrichment or ribosomal RNA depletion? • Which RNA entities you are interested in… • Transcriptome assembly: remove all ribosomal RNA (and maybe enrich for only poly. A+ transcripts) • Differential gene expression: enrich for Poly(A) • EXCEPTION – If you are aiming to obtain information about long non-coding RNAs • Metatranscriptomics: remove all the host materials • Remove r. RNA by molecular methods prior to sequencing • Remove host m. RNA by computational methods postsequencing

• • Technical replicates • Illumina has low technical variation unlike microarrays • Technical replicates are unnecessary Batch effects • Best to sequence everything for an experiment at the same time • If you are preparing the libraries, be consistent & make them simultaneously Biological replicates • This is essential for your experiment to have any statistical power • At least 3, but the more the better For transcriptome assembly • • RNA can be pooled from various sources to ensure the most robust transcriptome • Pooling can also be done after sequencing, but before assembly For differential gene expression • Pooling RNA from multiple biological replicates is usually not advisable • Only do so if you have multiple pools from each experimental condition
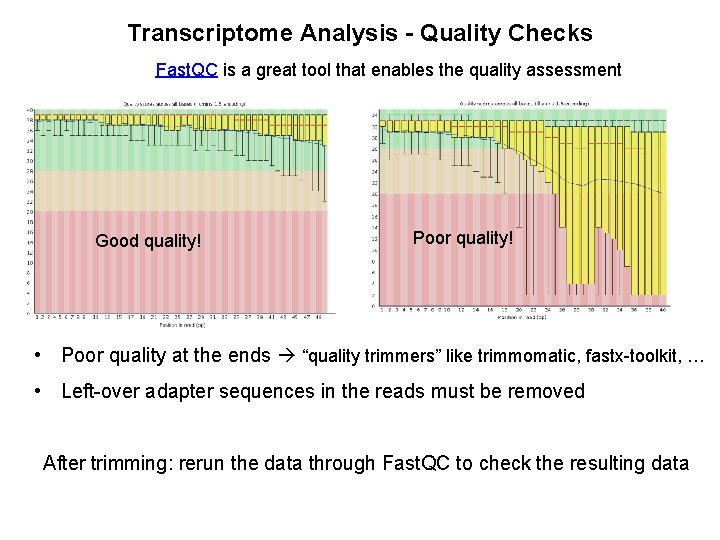
Transcriptome Analysis - Quality Checks Fast. QC is a great tool that enables the quality assessment Good quality! Poor quality! • Poor quality at the ends “quality trimmers” like trimmomatic, fastx-toolkit, … • Left-over adapter sequences in the reads must be removed After trimming: rerun the data through Fast. QC to check the resulting data
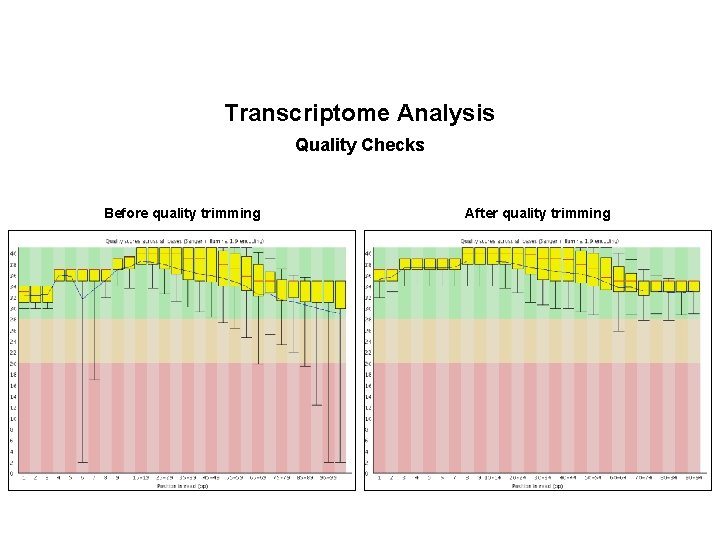
Transcriptome Analysis Quality Checks Before quality trimming After quality trimming
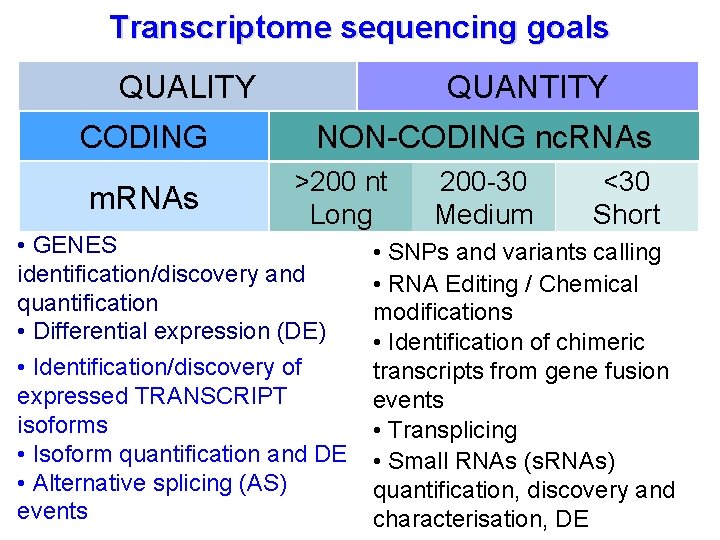
Transcriptome sequencing goals QUALITY CODING m. RNAs QUANTITY NON-CODING nc. RNAs >200 nt Long • GENES identification/discovery and quantification • Differential expression (DE) • Identification/discovery of expressed TRANSCRIPT isoforms • Isoform quantification and DE • Alternative splicing (AS) events 200 -30 Medium <30 Short • SNPs and variants calling • RNA Editing / Chemical modifications • Identification of chimeric transcripts from gene fusion events • Transplicing • Small RNAs (s. RNAs) quantification, discovery and characterisation, DE
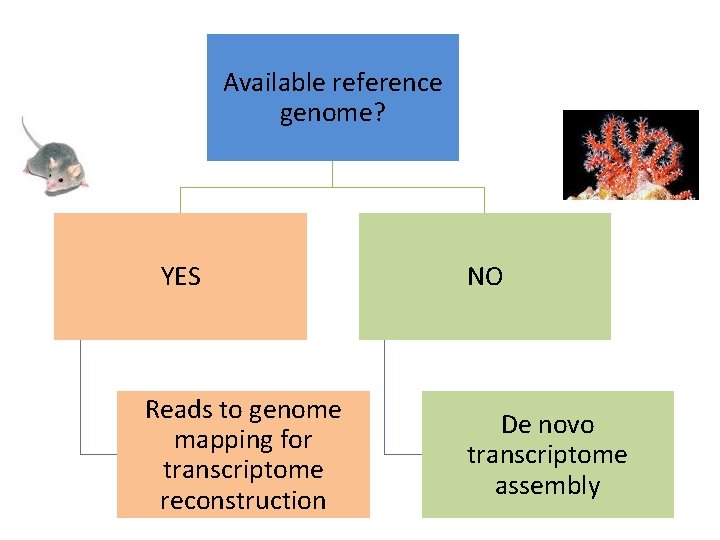
Available reference genome? YES Reads to genome mapping for transcriptome reconstruction NO De novo transcriptome assembly
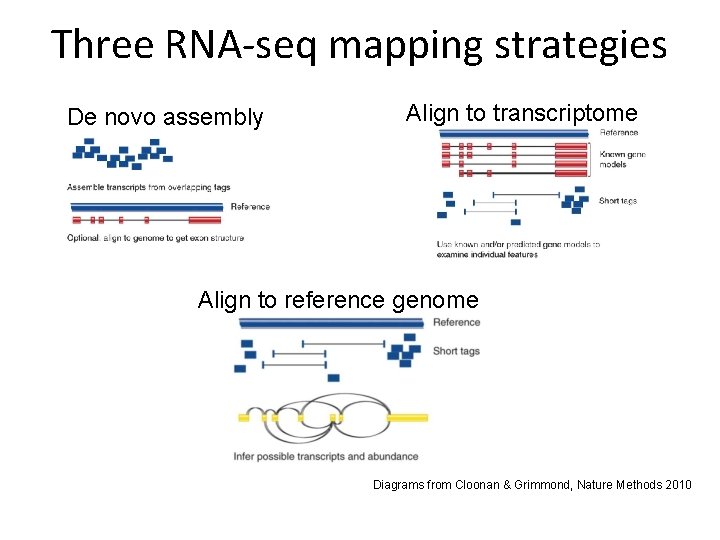
Three RNA-seq mapping strategies De novo assembly Align to transcriptome Align to reference genome Diagrams from Cloonan & Grimmond, Nature Methods 2010
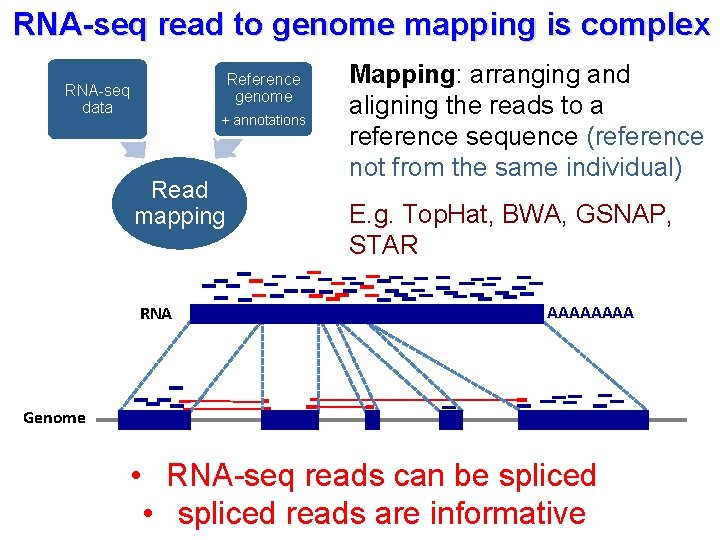
RNA-seq read to genome mapping is complex Reference genome RNA-seq data + annotations Read mapping RNA Mapping: arranging and aligning the reads to a reference sequence (reference not from the same individual) E. g. Top. Hat, BWA, GSNAP, STAR AAAA Genome • RNA-seq reads can be spliced • spliced reads are informative
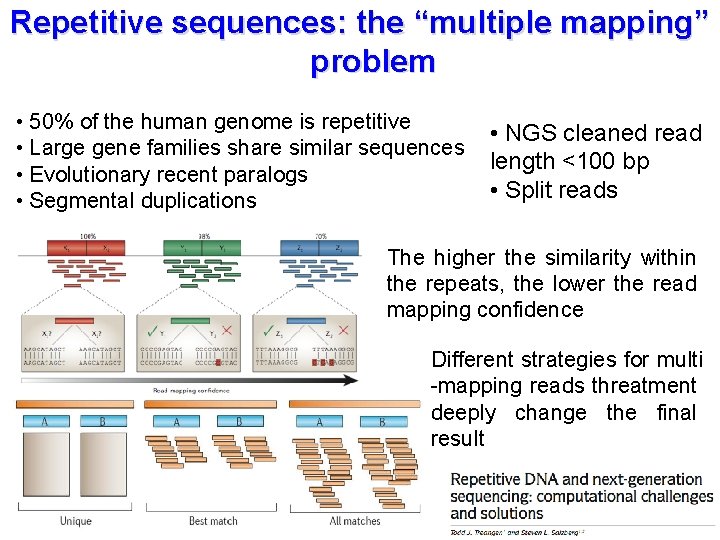
Repetitive sequences: the “multiple mapping” problem • 50% of the human genome is repetitive • Large gene families share similar sequences • Evolutionary recent paralogs • Segmental duplications • NGS cleaned read length <100 bp • Split reads The higher the similarity within the repeats, the lower the read mapping confidence Different strategies for multi -mapping reads threatment deeply change the final result
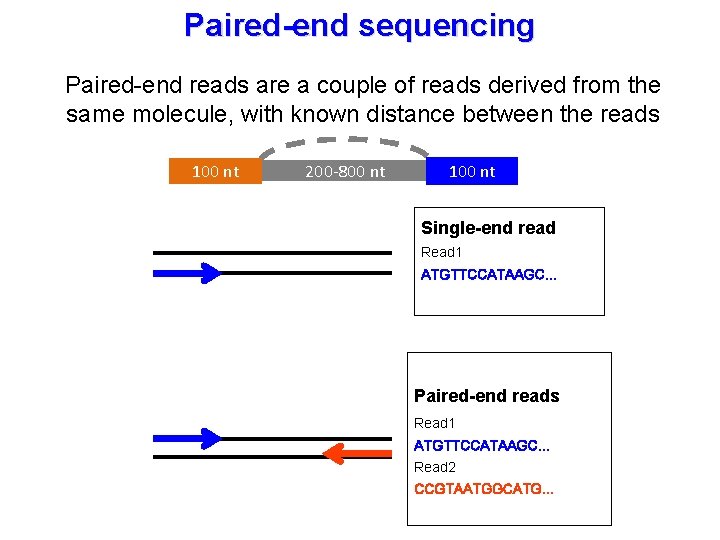
Paired-end sequencing Paired-end reads are a couple of reads derived from the same molecule, with known distance between the reads 100 nt 200 -800 nt 100 nt Single-end read Read 1 ATGTTCCATAAGC… Paired-end reads Read 1 ATGTTCCATAAGC… Read 2 CCGTAATGGCATG…
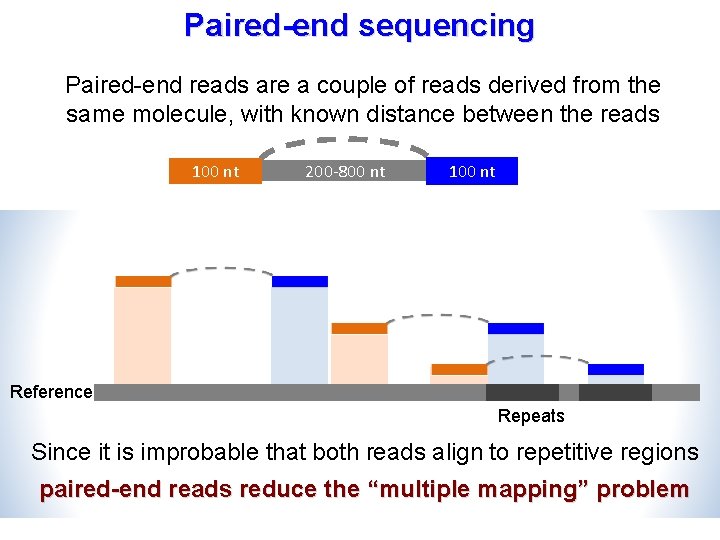
Paired-end sequencing Paired-end reads are a couple of reads derived from the same molecule, with known distance between the reads 100 nt 200 -800 nt 100 nt Reference Repeats Since it is improbable that both reads align to repetitive regions paired-end reads reduce the “multiple mapping” problem
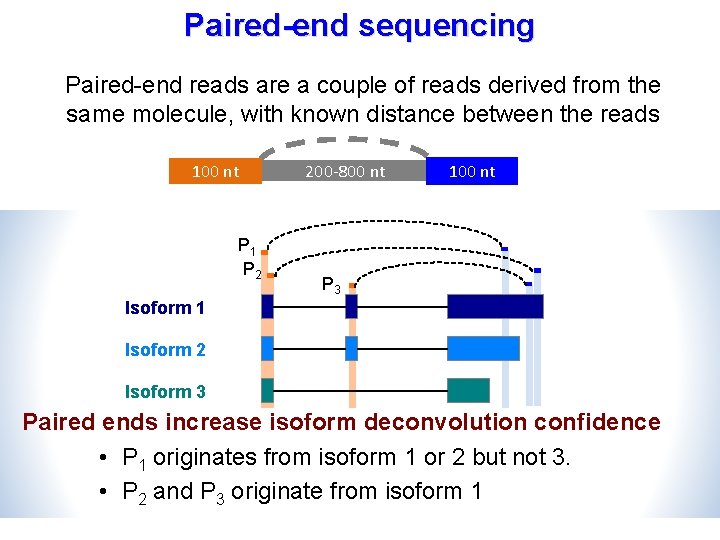
Paired-end sequencing Paired-end reads are a couple of reads derived from the same molecule, with known distance between the reads 100 nt P 1 P 2 200 -800 nt 100 nt P 3 Isoform 1 Isoform 2 Isoform 3 Paired ends increase isoform deconvolution confidence • P 1 originates from isoform 1 or 2 but not 3. • P 2 and P 3 originate from isoform 1
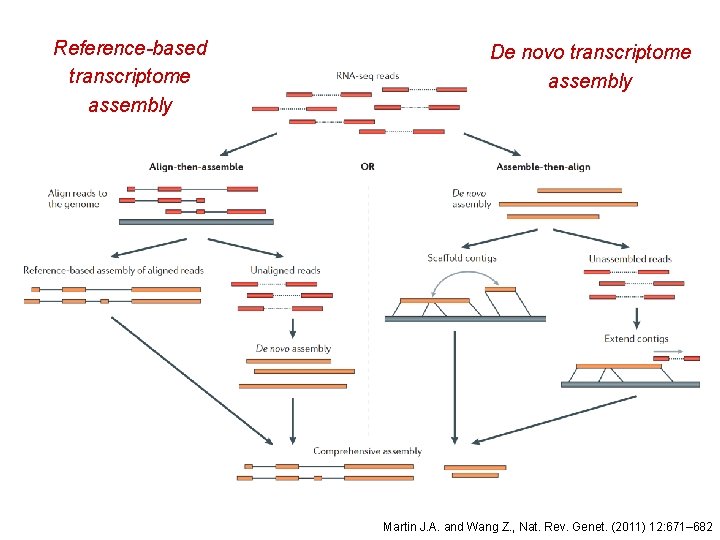
Reference-based transcriptome assembly De novo transcriptome assembly Martin J. A. and Wang Z. , Nat. Rev. Genet. (2011) 12: 671– 682

Reference-based transcriptome assembly • Used when the genome sequence is known • Transcriptome data are not available • Transcriptome information is available but not good enough, • i. e. missing isoforms of genes, or unknown noncoding regions • The existing transcriptome information is for a different tissue type • Cufflinks and Scripture are two reference-based transcriptome assemblers
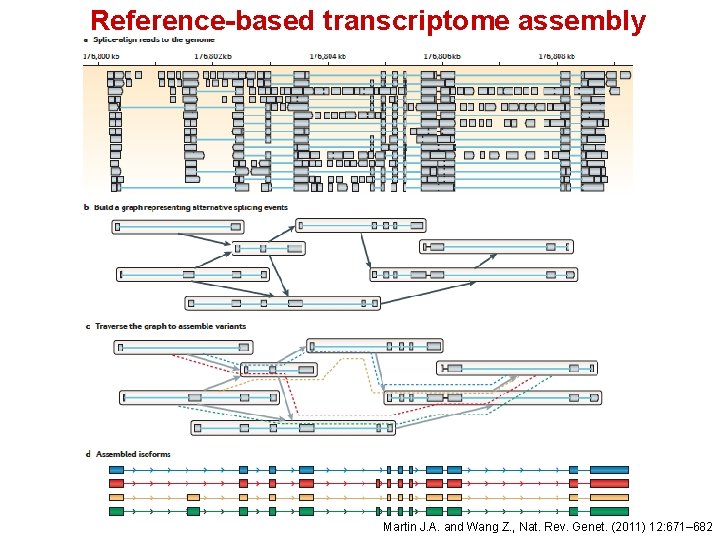
Reference-based transcriptome assembly Martin J. A. and Wang Z. , Nat. Rev. Genet. (2011) 12: 671– 682

Cufflinks assembles transcripts, estimates their abundances, and tests for differential expression in RNA-Seq samples 1. Aligned RNA-Seq reads are assembled into a parsimonious set of transcripts. 2. The relative abundances of transcripts are estimated based on how many reads support each one 3. Sequence-specific biases, introduced during the library preparation, that challenge the assumption of uniform coverage, are corrected 4. Multiple mapping of reads is taken into account

De novo transcriptome assembly • Used when very little information is available for the genome • Often the first step in putting together information about an unknown genome • Amount of data needed for a good de novo assembly is higher than what is needed for a reference-based assembly • Can be used for genome annotation, once the genome is assembled • Trinity, Oases, Trans. ABy. SS, are examples of well-regarded transcriptome assemblers
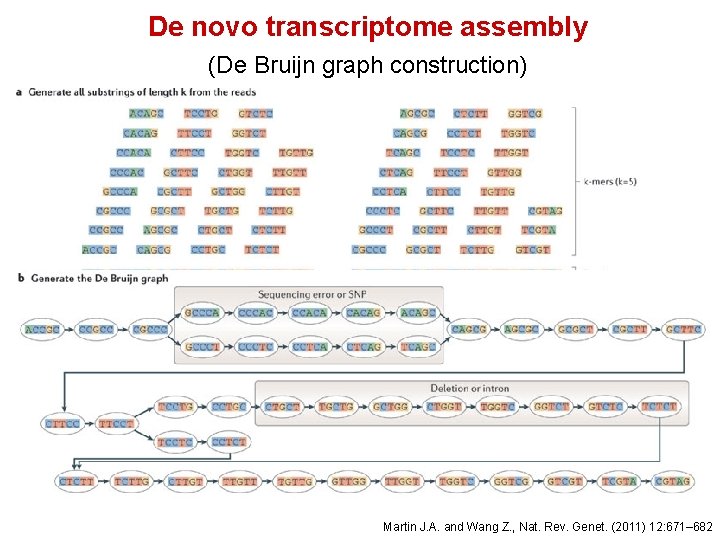
De novo transcriptome assembly (De Bruijn graph construction) Martin J. A. and Wang Z. , Nat. Rev. Genet. (2011) 12: 671– 682
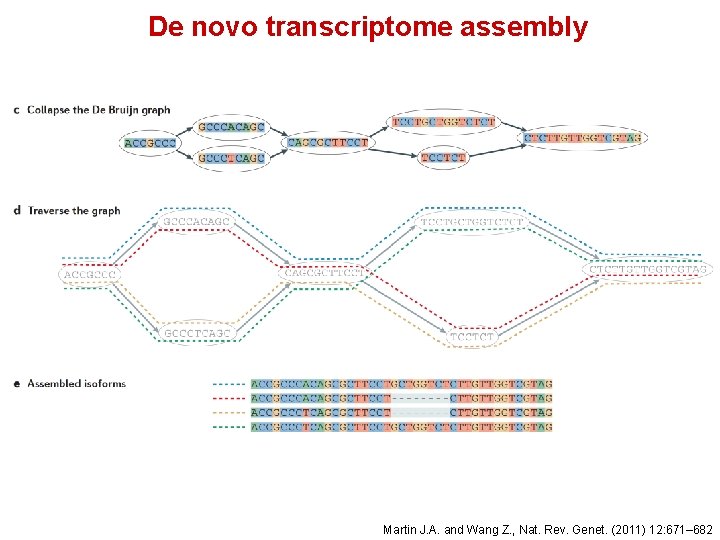
De novo transcriptome assembly Martin J. A. and Wang Z. , Nat. Rev. Genet. (2011) 12: 671– 682
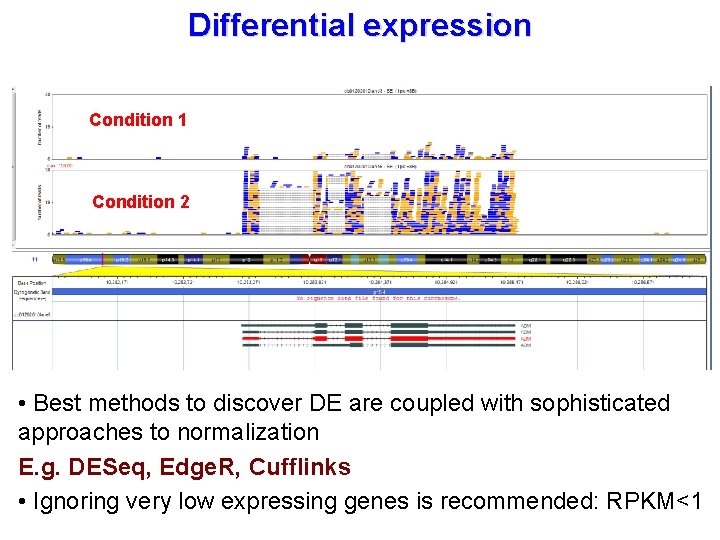
Differential expression Condition 1 Condition 2 • Best methods to discover DE are coupled with sophisticated approaches to normalization E. g. DESeq, Edge. R, Cufflinks • Ignoring very low expressing genes is recommended: RPKM<1

Count-based gene expression quantification • Longer transcripts have higher counts Length bias, an intrasample bias • Different experiments have different yields inter-samples bias. • Normalization is required: Reads Per Kilobase of exonic sequence per Million mapped reads (Mortazavi et al Nature methods 2008) • Non uniform coverage in different exons • Multiple mapping problem • Reads sharing among transcripts • Overlapping genes

Differential expression What genes/transcripts are differentially expressed in various test conditions? • The first step is proper normalization of the data • In general, RNA-Seq data do not follow a normal (Poisson) distribution, but follow a negative binomial distribution –> need of a statistical program that makes the correct assumptions • Edge. R, DESeq, other R/Bioconductor packages for DE detection

Cuff. Diff Uses the Cufflinks transcript quantification engine to calculate gene and transcript expression levels in more than one condition and test them for significant differences • The observed log-fold-change in gene/transcript expression is tested against the null hypothesis of no change • The null distribution of Lg. FC is estimated using the beta negative binomial model for each transcript in each condition

Cuff. Differential Expression tests isoform_exp. diff Transcript differential FPKM. Gene differential FPKM (using summed FPKM of transcripts of the same gene) gene_exp. diff Primary transcript differential FPKM (Tests differences in the summed FPKM of transcripts sharing TSS) tss_group_exp. diff Coding sequence differential FPKM (using summed FPKM of transcripts sharing protein_id) cds_exp. diff Differential splicing tests For each primary transcript, how much differential splicing exists between isoforms processed from a single primary transcript. splicing. diff Differential coding output Differential use of promoters
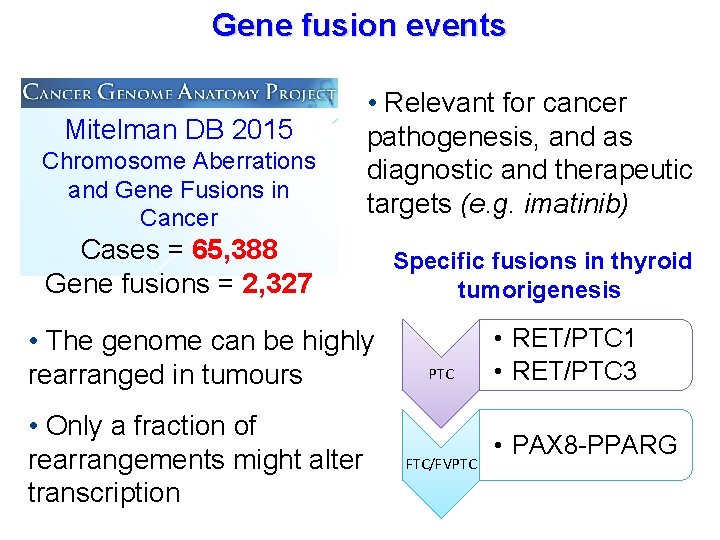
Gene fusion events Mitelman DB 2015 Chromosome Aberrations and Gene Fusions in Cancer • Relevant for cancer pathogenesis, and as diagnostic and therapeutic targets (e. g. imatinib) Cases = 65, 388 Gene fusions = 2, 327 • The genome can be highly rearranged in tumours • Only a fraction of rearrangements might alter transcription Specific fusions in thyroid tumorigenesis PTC FTC/FVPTC • RET/PTC 1 • RET/PTC 3 • PAX 8 -PPARG
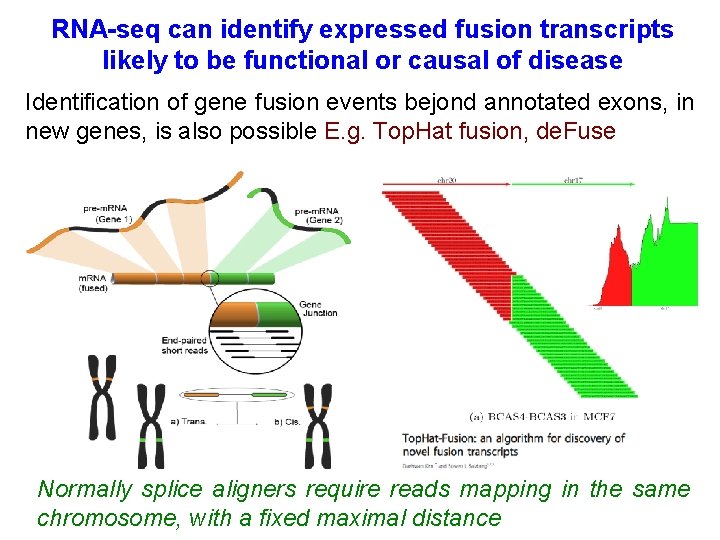
RNA-seq can identify expressed fusion transcripts likely to be functional or causal of disease Identification of gene fusion events bejond annotated exons, in new genes, is also possible E. g. Top. Hat fusion, de. Fuse Normally splice aligners require reads mapping in the same chromosome, with a fixed maximal distance
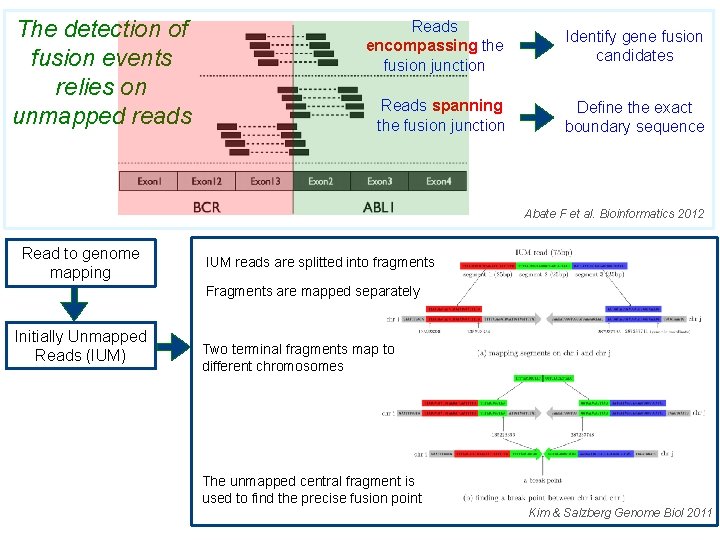
The detection of fusion events relies on unmapped reads Reads encompassing the fusion junction Identify gene fusion candidates Reads spanning the fusion junction Define the exact boundary sequence Abate F et al. Bioinformatics 2012 Read to genome mapping IUM reads are splitted into fragments Fragments are mapped separately Initially Unmapped Reads (IUM) Two terminal fragments map to different chromosomes The unmapped central fragment is used to find the precise fusion point Kim & Salzberg Genome Biol 2011
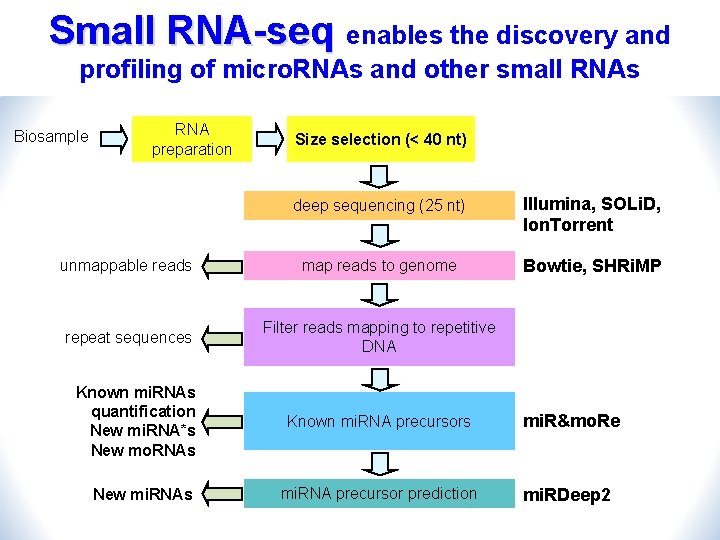
Small RNA-seq enables the discovery and profiling of micro. RNAs and other small RNAs Biosample RNA preparation Size selection (< 40 nt) deep sequencing (25 nt) Illumina, SOLi. D, Ion. Torrent unmappable reads map reads to genome Bowtie, SHRi. MP repeat sequences Filter reads mapping to repetitive DNA Known mi. RNAs quantification New mi. RNA*s New mo. RNAs New mi. RNAs Known mi. RNA precursors mi. R&mo. Re mi. RNA precursor prediction mi. RDeep 2
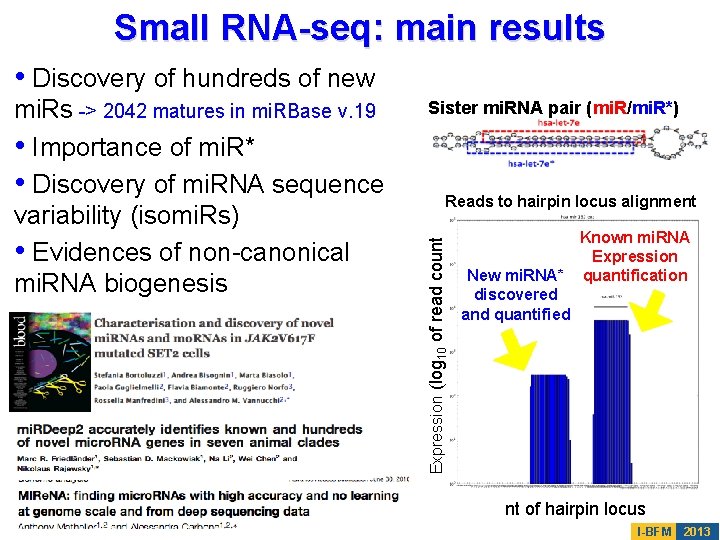
Small RNA-seq: main results • Discovery of hundreds of new Sister mi. RNA pair (mi. R/mi. R*) Reads to hairpin locus alignment Expression (log 10 of read count mi. Rs -> 2042 matures in mi. RBase v. 19 • Importance of mi. R* • Discovery of mi. RNA sequence variability (isomi. Rs) • Evidences of non-canonical mi. RNA biogenesis Known mi. RNA Expression New mi. RNA* quantification discovered and quantified nt of hairpin locus I-BFM 2013

A large-scale small RNA-seq study recently doubled the mi. RNA repertoire • Support of novel mi. RNAs by a Dicer KO experiment • Other validations

Thousands new mi. RNAs validated 2013 2015 NUMBER OF MIRNAS ≈2000 ≈4000 • Many are seed-paralogues of known mirbase mi. RNAs • Many are specific to the Hominidae family of Primates • Weakly expressed • Expressed with tissue-, development-, differentiationspecific patterns

mo. RNA (mi. RNA-offset RNA) discovery u ~20 nt long RNAs derived from tends of pre-mi. RNAs u Novel type of mi. RNA-related small RNAs, discovered by RNA-seq in the ascidian Ciona intestinalis, then found in human cells, in solid tumours and in herpes viruses infected cells. u Some are enriched in the nuclear fraction of RNAs, function unknown u One locus -> 4 products! mi. RNA mo. RNA
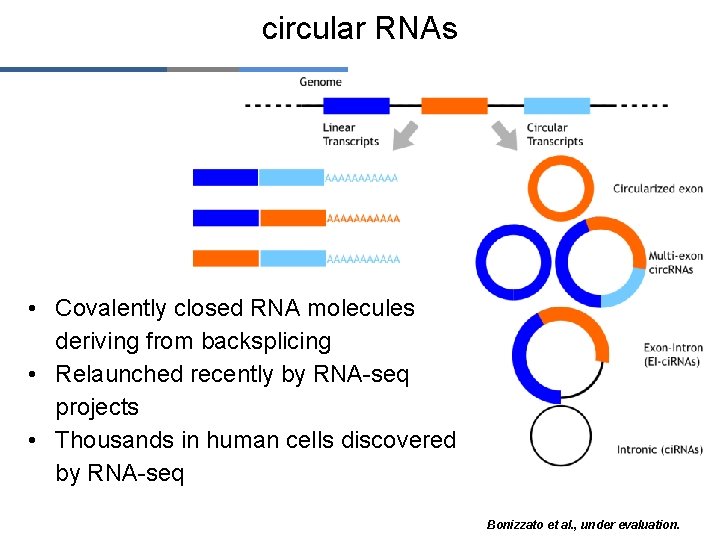
circular RNAs • Covalently closed RNA molecules deriving from backsplicing • Relaunched recently by RNA-seq projects • Thousands in human cells discovered by RNA-seq Bonizzato et al. , under evaluation.

Circ. RNAs play important functions and have distinctive features Evolutionarily conserved Ubiquitary and highly expressed Tissue and developmental stage specificity Highly stable Detectable in body fluids Disease cancer markers Bonizzato et al. , under evaluation. Competitors of m. RNA splicing
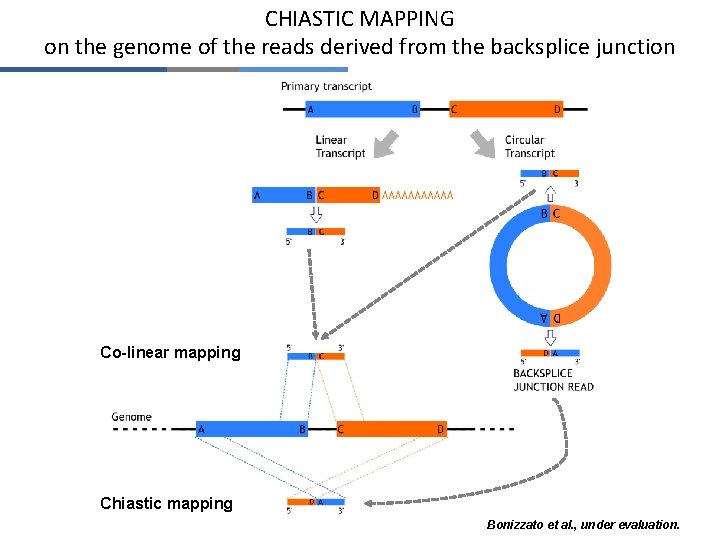
CHIASTIC MAPPING on the genome of the reads derived from the backsplice junction Co-linear mapping Chiastic mapping Bonizzato et al. , under evaluation.
- Slides: 50