864 Design and applications of unified controlled vocabularies
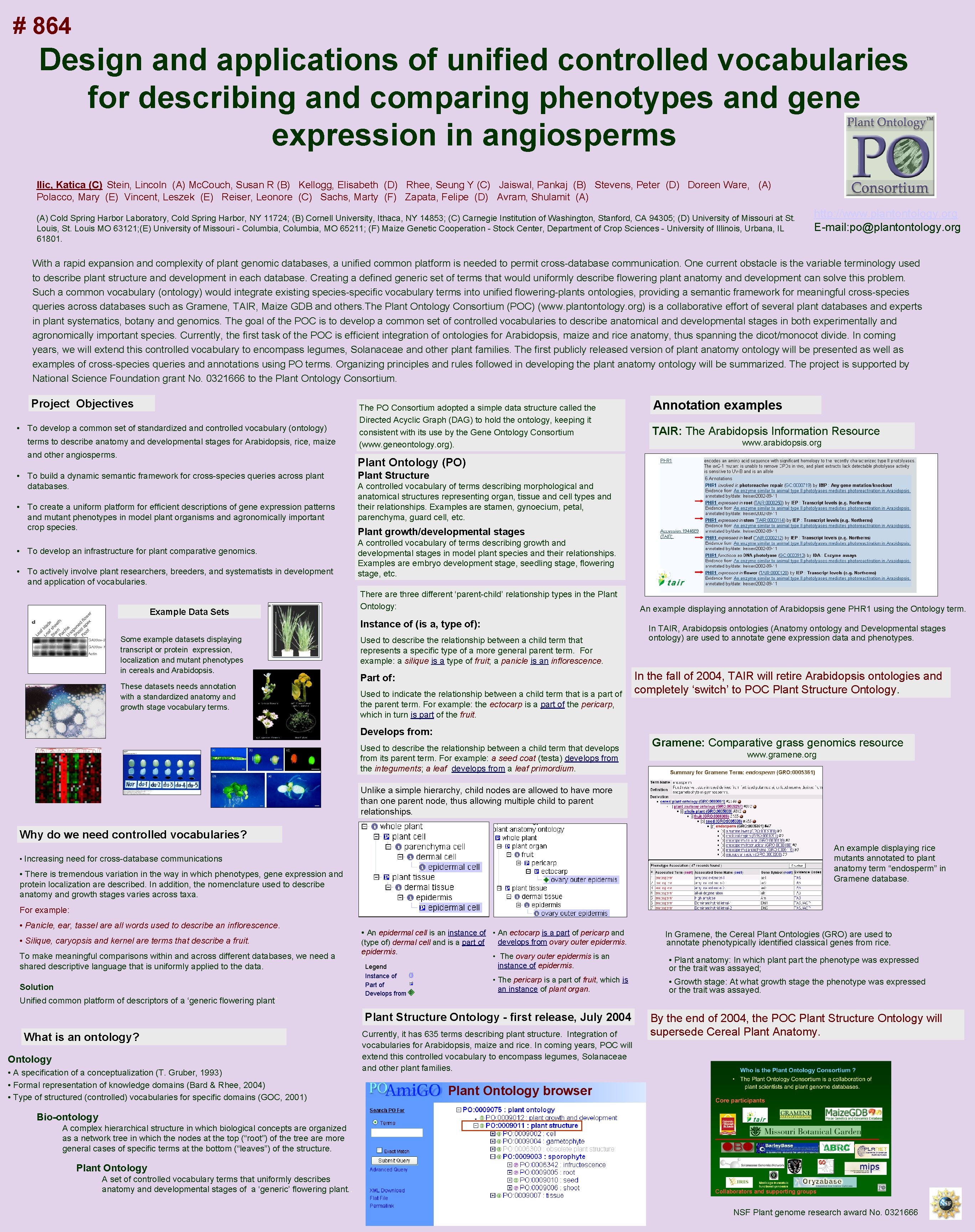
# 864 Design and applications of unified controlled vocabularies for describing and comparing phenotypes and gene expression in angiosperms Ilic, Katica (C) Stein, Lincoln (A) Mc. Couch, Susan R (B) Kellogg, Elisabeth (D) Rhee, Seung Y (C) Jaiswal, Pankaj (B) Stevens, Peter (D) Doreen Ware, (A) Polacco, Mary (E) Vincent, Leszek (E) Reiser, Leonore (C) Sachs, Marty (F) Zapata, Felipe (D) Avram, Shulamit (A) Cold Spring Harbor Laboratory, Cold Spring Harbor, NY 11724; (B) Cornell University, Ithaca, NY 14853; (C) Carnegie Institution of Washington, Stanford, CA 94305; (D) University of Missouri at St. Louis, St. Louis MO 63121; (E) University of Missouri - Columbia, MO 65211; (F) Maize Genetic Cooperation - Stock Center, Department of Crop Sciences - University of Illinois, Urbana, IL 61801. http: //www. plantontology. org E-mail: po@plantontology. org With a rapid expansion and complexity of plant genomic databases, a unified common platform is needed to permit cross-database communication. One current obstacle is the variable terminology used to describe plant structure and development in each database. Creating a defined generic set of terms that would uniformly describe flowering plant anatomy and development can solve this problem. Such a common vocabulary (ontology) would integrate existing species-specific vocabulary terms into unified flowering-plants ontologies, providing a semantic framework for meaningful cross-species queries across databases such as Gramene, TAIR, Maize GDB and others. The Plant Ontology Consortium (POC) (www. plantontology. org) is a collaborative effort of several plant databases and experts in plant systematics, botany and genomics. The goal of the POC is to develop a common set of controlled vocabularies to describe anatomical and developmental stages in both experimentally and agronomically important species. Currently, the first task of the POC is efficient integration of ontologies for Arabidopsis, maize and rice anatomy, thus spanning the dicot/monocot divide. In coming years, we will extend this controlled vocabulary to encompass legumes, Solanaceae and other plant families. The first publicly released version of plant anatomy ontology will be presented as well as examples of cross-species queries and annotations using PO terms. Organizing principles and rules followed in developing the plant anatomy ontology will be summarized. The project is supported by National Science Foundation grant No. 0321666 to the Plant Ontology Consortium. Project Objectives • To develop a common set of standardized and controlled vocabulary (ontology) terms to describe anatomy and developmental stages for Arabidopsis, rice, maize and other angiosperms. The PO Consortium adopted a simple data structure called the Directed Acyclic Graph (DAG) to hold the ontology, keeping it consistent with its use by the Gene Ontology Consortium (www. geneontology. org). Annotation examples TAIR: The Arabidopsis Information Resource www. arabidopsis. org Plant Ontology (PO) • To build a dynamic semantic framework for cross-species queries across plant databases. • To create a uniform platform for efficient descriptions of gene expression patterns and mutant phenotypes in model plant organisms and agronomically important crop species. • To develop an infrastructure for plant comparative genomics. • To actively involve plant researchers, breeders, and systematists in development and application of vocabularies. Example Data Sets Plant Structure A controlled vocabulary of terms describing morphological and anatomical structures representing organ, tissue and cell types and their relationships. Examples are stamen, gynoecium, petal, parenchyma, guard cell, etc. Plant growth/developmental stages A controlled vocabulary of terms describing growth and developmental stages in model plant species and their relationships. Examples are embryo development stage, seedling stage, flowering stage, etc. There are three different ‘parent-child’ relationship types in the Plant Ontology: Instance of (is a, type of): Some example datasets displaying transcript or protein expression, localization and mutant phenotypes in cereals and Arabidopsis. These datasets needs annotation with a standardized anatomy and growth stage vocabulary terms. Used to describe the relationship between a child term that represents a specific type of a more general parent term. For example: a silique is a type of fruit; a panicle is an inflorescence. Part of: Used to indicate the relationship between a child term that is a part of the parent term. For example: the ectocarp is a part of the pericarp, which in turn is part of the fruit. Develops from: Used to describe the relationship between a child term that develops from its parent term. For example: a seed coat (testa) develops from the integuments; a leaf develops from a leaf primordium. An example displaying annotation of Arabidopsis gene PHR 1 using the Ontology term. In TAIR, Arabidopsis ontologies (Anatomy ontology and Developmental stages ontology) are used to annotate gene expression data and phenotypes. In the fall of 2004, TAIR will retire Arabidopsis ontologies and completely ‘switch’ to POC Plant Structure Ontology. Gramene: Comparative grass genomics resource www. gramene. org Unlike a simple hierarchy, child nodes are allowed to have more than one parent node, thus allowing multiple child to parent relationships. Why do we need controlled vocabularies? An example displaying rice mutants annotated to plant anatomy term “endosperm” in Gramene database. • Increasing need for cross-database communications • There is tremendous variation in the way in which phenotypes, gene expression and protein localization are described. In addition, the nomenclature used to describe anatomy and growth stages varies across taxa. For example: • Panicle, ear, tassel are all words used to describe an inflorescence. • Silique, caryopsis and kernel are terms that describe a fruit. To make meaningful comparisons within and across different databases, we need a shared descriptive language that is uniformly applied to the data. Solution Unified common platform of descriptors of a ‘generic flowering plant’ • An epidermal cell is an instance of • An ectocarp is a part of pericarp and (type of) dermal cell and is a part of epidermis. ) Legend Instance of Part of Develops from i P D develops from ovary outer epidermis. • The ovary outer epidermis is an instance of epidermis. • Plant anatomy: In which plant part the phenotype was expressed or the trait was assayed; • The pericarp is a part of fruit, which is an instance of plant organ. • Growth stage: At what growth stage the phenotype was expressed or the trait was assayed. Plant Structure Ontology - first release, July 2004 What is an ontology? Ontology • A specification of a conceptualization (T. Gruber, 1993) • Formal representation of knowledge domains (Bard & Rhee, 2004) • Type of structured (controlled) vocabularies for specific domains (GOC, 2001) In Gramene, the Cereal Plant Ontologies (GRO) are used to annotate phenotypically identified classical genes from rice. Currently, it has 635 terms describing plant structure. . Integration of vocabularies for Arabidopsis, maize and rice. In coming years, POC will extend this controlled vocabulary to encompass legumes, Solanaceae and other plant families. By the end of 2004, the POC Plant Structure Ontology will supersede Cereal Plant Anatomy. Plant Ontology browser Bio-ontology A complex hierarchical structure in which biological concepts are organized as a network tree in which the nodes at the top (“root”) of the tree are more general cases of specific terms at the bottom (“leaves”) of the structure. Plant Ontology A set of controlled vocabulary terms that uniformly describes anatomy and developmental stages of a ‘generic’ flowering plant. . Collaborators and supporting groups NSF Plant genome research award No. 0321666

- Slides: 2