4 th Solanaceae Genome Workshop 2007 September 09

4 th Solanaceae Genome Workshop 2007, September 09 th 13 th, Jeju Island, Korea T 0848 TM 15 T 0731 T 0966 TG 438 T 1257 TG 216 CT 54 TM 18 T 1347 T 0676 T 1497 T 1962 T 1414 T 1428 T 1355 T 1112 THE FRENCH CONTRIBUTION TO THE INTERNATIONAL TOMATO GENOME SEQUENCING PROGRAM

Chromosome 7 4 th Solanaceae Genome Workshop 2007, September 09 th 13 th, Jeju Island, Korea T 0848 TM 15 T 0731 T 0966 TG 438 T 1257 TG 216 CT 54 TM 18 T 1347 T 0676 T 1497 T 1962 T 1414 T 1428 T 1355 T 1112 Selection of seed BACs

BAC selection strategy • Selection and sequencing of 100 "seed BACs" to cover the gene rich regions of Chromosome 7 • Check the location of each BAC on K 7 by FISH and/or by polymorphism screening on ILs • Selection of overlapping BACs by in silico approaches or 3 D DNA pool screening (built from 2 BAC libraries). 4 th Solanaceae Genome Workshop 2007, September 09 th 13 th, Jeju Island, Korea

Initial source of Seed BACs : overgo assay • 43 BACs selected from the initial list of "overgo" seed BACs • 20 BAC sequenced • 12 Not on K 7 • 11 Discarded (large overlapping, misannotation…) 27% of the « overgo » seed BAC are not on K 7 4 th Solanaceae Genome Workshop 2007, September 09 th 13 th, Jeju Island, Korea

Initial strategy : FPC contig • Minimal tiling path strategy (FPC, USA and China): • 23 potential BACs selected • 9 sequenced • 11 not on K 7 • 3 discarded (large overlapping, misannotation…) 47% of BACs selected on the basis of FPC contigs could not be used 4 th Solanaceae Genome Workshop 2007, September 09 th 13 th, Jeju Island, Korea

New strategies for "seed BAC" selection: (1) in silico search using genetic markers • Expend 2000 genetic map: 182 markers on group 7 • 27 potential seed BACs selected based on blastn of the marker sequence against BES database • 18 sequenced • 5 not on K 7 • 4 discarded (large overlapping, misannotation…) 18% of seed BACs selected by in silico search are not on K 7 4 th Solanaceae Genome Workshop 2007, September 09 th 13 th, Jeju Island, Korea

New strategies for "seed BAC" selection: (2) Other sources • collaboration / bibliography : – – Australie / David Jones (fusarium) FISH / Steve Stack Sun Locus (Steve Tanksley) Sequences from Database (Genbank) 4 BACs 3 BACs 2 BACs 4 BACs – 13 seed BACs sequenced 4 th Solanaceae Genome Workshop 2007, September 09 th 13 th, Jeju Island, Korea

New strategies for "seed BAC" selection: (3) SYNGENTA markers • 26 gene markers and 48 SSR mapped on K 7: After sequence analysis only 16 genes and 36 SSR were retained (sequences redondancy) • In silico search for BES sequence homology 6 seed BAC selected and sequenced • Macroarray hybridization with remaining gene markers (collab. CNRGV) 4 seed BAC selected and sequenced • The Syngenta SSR markers will be used when all Expend 2000 markers will be tested 4 th Solanaceae Genome Workshop 2007, September 09 th 13 th, Jeju Island, Korea

"seed BAC" and "overlapping BAC" selection (4) 3 D-DNA pool screening The 3 D-DNA pools have been validated for selection of both: • • seed BAC screens by genetic markers and for overlapping BAC selection If markers are not found in the 3 D-DNA pools of Mbo. I and Hind. III libraries, the Eco. RI Macroarray filters are screened 4 th Solanaceae Genome Workshop 2007, September 09 th 13 th, Jeju Island, Korea

Chromosome 7 4 th Solanaceae Genome Workshop 2007, September 09 th 13 th, Jeju Island, Korea T 0848 TM 15 T 0731 T 0966 TG 438 T 1257 TG 216 CT 54 TM 18 T 1347 T 0676 T 1497 T 1962 T 1414 T 1428 T 1355 T 1112 Check BAC location on K 7
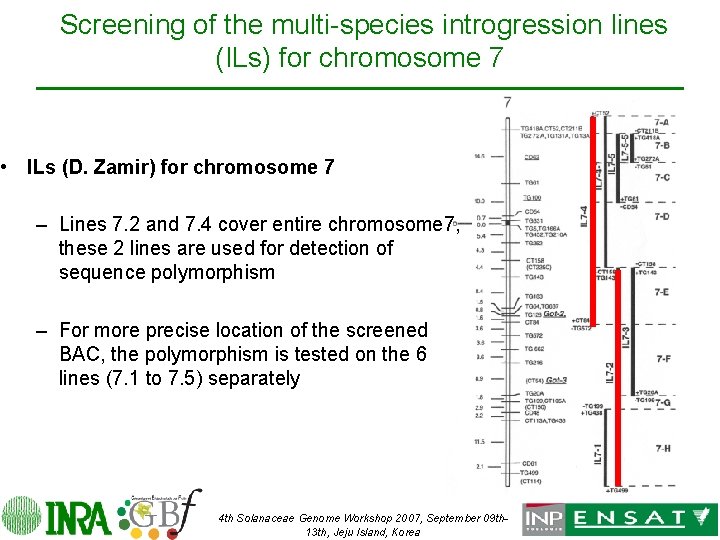
Screening of the multi-species introgression lines (ILs) for chromosome 7 • ILs (D. Zamir) for chromosome 7 – Lines 7. 2 and 7. 4 cover entire chromosome 7, these 2 lines are used for detection of sequence polymorphism – For more precise location of the screened BAC, the polymorphism is tested on the 6 lines (7. 1 to 7. 5) separately 4 th Solanaceae Genome Workshop 2007, September 09 th 13 th, Jeju Island, Korea

ILs screening results Screened BACs 125 BACs validated on K 7 62 % BAC not on K 7 15 % Unsuccessful tests 23 %* *On 28 unusable BAC sequences, 18 BAC gave good result using FISH 78 BACs on chromosome 7 checked by screening the ILs 4 th Solanaceae Genome Workshop 2007, September 09 th 13 th, Jeju Island, Korea

BAC-FISH • BAC-FISH technology on pachyten meiotic chromosomes – The Netherlands (H. de Jong) – USA (Steve Stack / Colorado) – China (Zukan Chen) • BAC-FISH technology on mitotic chromosome – France (INRA-Rennes / Olivier Coriton) 4 th Solanaceae Genome Workshop 2007, September 09 th 13 th, Jeju Island, Korea

BAC-FISH on Chromosome 7 *On 20 unsuccessful BACs, 13 BACs gave good result using ILs 44 "seed BAC" assigned to K 7 by FISH 4 th Solanaceae Genome Workshop 2007, September 09 th 13 th, Jeju Island, Korea

Present state of the progress Septrember 1 st, 2 OO 7 • Total BAC sequenced 120 : 40 complete + 80 in progress (DACs finishing strategy) – "seed BAC": 31 complete + 57 in progress – "overlapping BACs": 9 complete + 23 in progress 4 th Solanaceae Genome Workshop 2007, September 09 th 13 th, Jeju Island, Korea
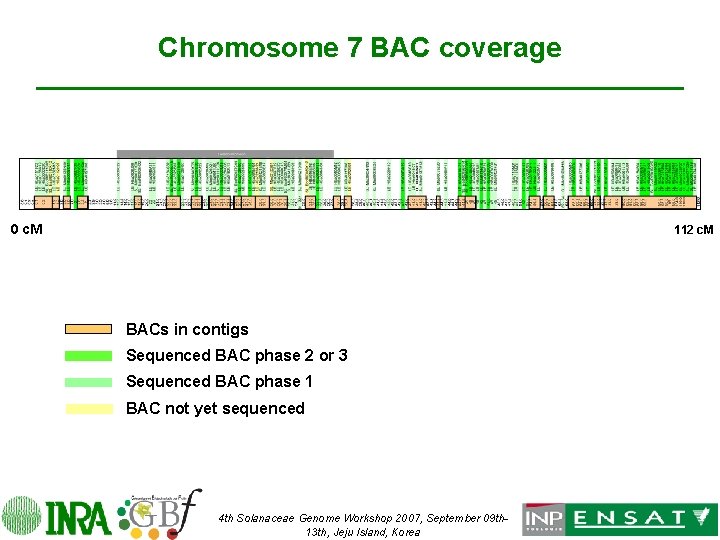
Chromosome 7 BAC coverage 0 c. M 112 c. M BACs in contigs Sequenced BAC phase 2 or 3 Sequenced BAC phase 1 BAC not yet sequenced 4 th Solanaceae Genome Workshop 2007, September 09 th 13 th, Jeju Island, Korea
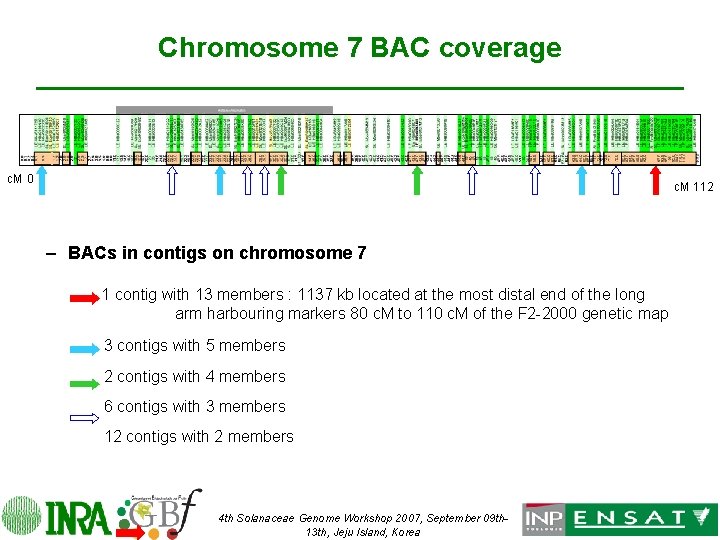
Chromosome 7 BAC coverage c. M 0 c. M 112 – BACs in contigs on chromosome 7 1 contig with 13 members : 1137 kb located at the most distal end of the long arm harbouring markers 80 c. M to 110 c. M of the F 2 -2000 genetic map 3 contigs with 5 members 2 contigs with 4 members 6 contigs with 3 members 12 contigs with 2 members 4 th Solanaceae Genome Workshop 2007, September 09 th 13 th, Jeju Island, Korea

Contigs of BACs on chromosome 7 • The selection of overlapping BACs started only few months ago – 78 BACs are in contigs on chromosome 7 • • • 1 contig with 13 members 1137 kb located at the most distal end of the long arm harbouring markers 80 c. M to 110 c. M of the F 2 -2000 genetic map 3 contigs with 5 members 2 contigs with 4 members 6 contigs with 3 members 12 contigs with 2 members – 42 BAC remain single 4 th Solanaceae Genome Workshop 2007, September 09 th 13 th, Jeju Island, Korea

Chromosome 7 4 th Solanaceae Genome Workshop 2007, September 09 th 13 th, Jeju Island, Korea T 0848 TM 15 T 0731 T 0966 TG 438 T 1257 TG 216 CT 54 TM 18 T 1347 T 0676 T 1497 T 1962 T 1414 T 1428 T 1355 T 1112 Gene content of euchromatic and heterochromatic BACs

Chromosome 7 The heterochromatin region M 21 D 10 H 30 C 22 • FISH mapping on meiotic chromosomes allowed the positioning of two BACs (H 30 C 22 and M 21 D 10) at the borders of a large heterochromatic region on chromosome 7. • 10 BACs lying in heterochromatin and 15 on euchromatin were sequenced. • The sequences were blasted for the presence of sequence repeats (blastn/SGN Uni. Repeats) and genes (blastx/Ref. Seq protein A. th). 4 th Solanaceae Genome Workshop 2007, September 09 th 13 th, Jeju Island, Korea

DNA sequence analysis in Hetero- and Eu-chromatin Heterochromatin Euchromatin nb BAC 10 15 total size (kb) 1124 1287 % repeats 16 1, 3 nb gene / 100 kb 3, 4 8, 1 • heterochromatic BACs contain up to 1/3 of the total putative genes identified in the 25 BACs. • Coverage of heterochromatin might be required if the aim is to retrieve most of the gene-containing regions of chromosome 7. 4 th Solanaceae Genome Workshop 2007, September 09 th 13 th, Jeju Island, Korea

Chromosome 7 4 th Solanaceae Genome Workshop 2007, September 09 th 13 th, Jeju Island, Korea T 0848 TM 15 T 0731 T 0966 TG 438 T 1257 TG 216 CT 54 TM 18 T 1347 T 0676 T 1497 T 1962 T 1414 T 1428 T 1355 T 1112 The built of 3 -D DNA pools for BAC screening

3 D-DNA pool screening • The 3 D pools were generated in collaboration with the French Plant Genomic Resource Centre (CNRGV, INRA-Toulouse) – Half of the Hind. III BAC library: • 168 plates 384 • 64512 clones • 7. 8 X genome equivalent – The entire Mbo. I BAC library : • 144 plates 384 • 52 296 clones • 7. 5 X genome equivalent 4 th Solanaceae Genome Workshop 2007, September 09 th 13 th, Jeju Island, Korea
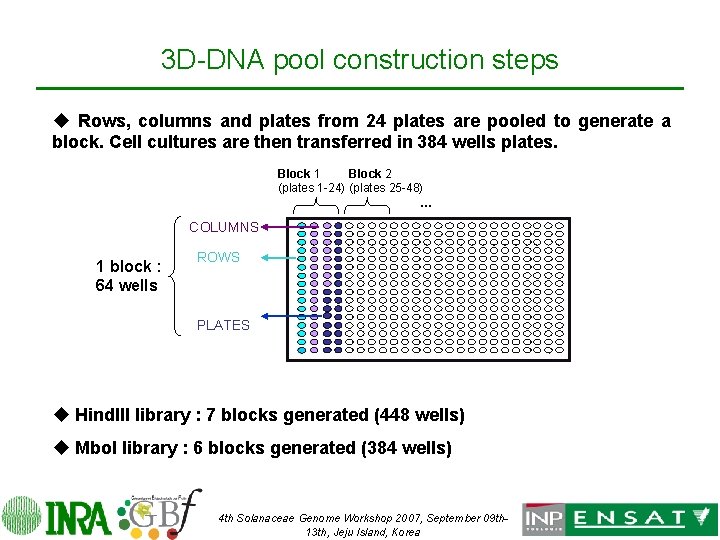
3 D-DNA pool construction steps u Rows, columns and plates from 24 plates are pooled to generate a block. Cell cultures are then transferred in 384 wells plates. Block 1 Block 2 (plates 1 -24) (plates 25 -48) … COLUMNS 1 block : 64 wells ROWS PLATES u Hind. III library : 7 blocks generated (448 wells) u Mbo. I library : 6 blocks generated (384 wells) 4 th Solanaceae Genome Workshop 2007, September 09 th 13 th, Jeju Island, Korea

3 D-DNA pool construction steps u DNA of the pooled cultures is amplified from the blocks using Phi 29 enzyme Dean, F. B. , Genome research, 2001 u After dilution a high amount of material is available for screening http: //cnrgv. toulouse. inra. fr/ 4 th Solanaceae Genome Workshop 2007, September 09 th 13 th, Jeju Island, Korea
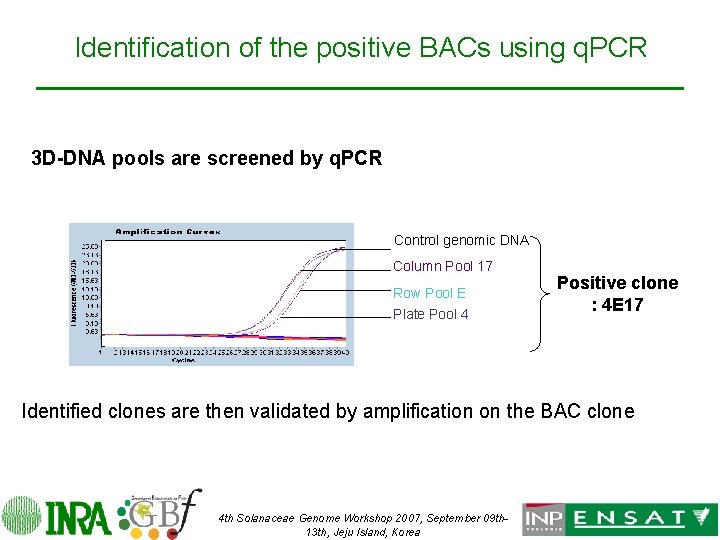
Identification of the positive BACs using q. PCR 3 D-DNA pools are screened by q. PCR Control genomic DNA Column Pool 17 Row Pool E Plate Pool 4 Positive clone : 4 E 17 Identified clones are then validated by amplification on the BAC clone 4 th Solanaceae Genome Workshop 2007, September 09 th 13 th, Jeju Island, Korea
Chromosome 7 4 th Solanaceae Genome Workshop 2007, September 09 th 13 th, Jeju Island, Korea T 0848 TM 15 T 0731 T 0966 TG 438 T 1257 TG 216 CT 54 TM 18 T 1347 T 0676 T 1497 T 1962 T 1414 T 1428 T 1355 T 1112 Macroarray DNA filters

Macroarray filters generated by the French Plant Genomic Resource Centre (CNRGV, INRA-Toulouse) Mbo. I Hind. III Eco. RI 144 plates 336 plates 196 plates spotting 2 macro-arrays 5 macro-arrays 3 macro-arrays 72 plates/filter = 27 648 clones 4 th Solanaceae Genome Workshop 2007, September 09 th 13 th, Jeju Island, Korea

The French Genomic Ressources Center for Plant CNRGV is both service provider and repository of plant genomes and data The missions are : - Centralize and maintain plant genomic resources - Distribute these resources at the international level - Provide high quality research material - Provide genomic tools for studying genomic collections - Propose genomics training court Hélène BERGES : INRA - CNRGV / email : hberges@toulouse. inra. fr http: //cnrgv. toulouse. inra. fr/ CNRGV-INRA Chemin de Borde Rouge/ B. P. 52627 / 31326 Castanet-Tolosan Tél: 05 61 28 52 September 53 / Fax: 0509 th 61 28 55 64 4 th Solanaceae Genome Workshop 2007, 13 th, Jeju Island, Korea

The French Genomic Ressources Center for Plant The CNRGV proposes pooling of genomic libraries. Pools of bacterial clones constitute a powerful tool for the screening of genomic libraries. Bacterial clones are mixed, with the aim of minimizing the number of reactions required to identify a clone containing a sequence of interest. - Production of different pools Plate Pools Row Pools Column Pools Superpool: mix of all of the clones from all the plates Plate Pools : mix of the 384 clones on each plate Row Pools : mix of the clones from each row for all the plates: 16 row pools The tomato Hind. II and Mbo. I BAC libraries pools are available for the community. Column Pools : mix of the clones from each column for all the plates: 24 column pools CNRGV-INRA Chemin de Borde Rouge/ B. P. 52627 / 31326 Castanet-Tolosan Tél: 05 61 28 52 September 53 / Fax: 0509 th 61 28 55 64 4 th Solanaceae Genome Workshop 2007, 13 th, Jeju Island, Korea
- Slides: 30