4 1 Linkage basic diploid eukaryotic chromosome mapping

4. 1 Linkage: basic diploid eukaryotic chromosome mapping

Linkage discovery • • Bateson and Punnet (early XX c. ): Purple flowers and long pollen PPLL Red flowers and round pollen ppll F 2 – Purple long (P_L_) 4831 – Purple round (P_ll ) 390 – Red long (pp. L_ ) 393 – Red round (ppll ) 1338 • TOTAL 6952 Expected 9 3991 3 1303 1 435

Linkage in Drosophila • T. H. Morgan – pr/pr +: purple/red – vg/vg+ : vestigial winds/normal • pr pr vg vg X pr+ vg+ • Test cross pr pr vg vg X pr+ pr vg+ vg Expected porportion 1: 1: 1: 1 pr + vg pr vg+ pr vg pr + vg + TOTAL 151 154 1195 1339 2839 Repulsion Coupling

Linkage in Drosophila (II) Crossing with alternated alleles: – pr+ vg vg X pr pr vg+ – F 1 pr+ pr vg+ vg X pr pr vg vg pr + vg pr vg + pr vg pr + vg + TOTAL 965 1067 146 157 2335

Linked alleles tend to be inherited together

Crossing over produces new allelic combinations

Linkage detection

For linked genes, recombinant frequencies are less than 50 percent
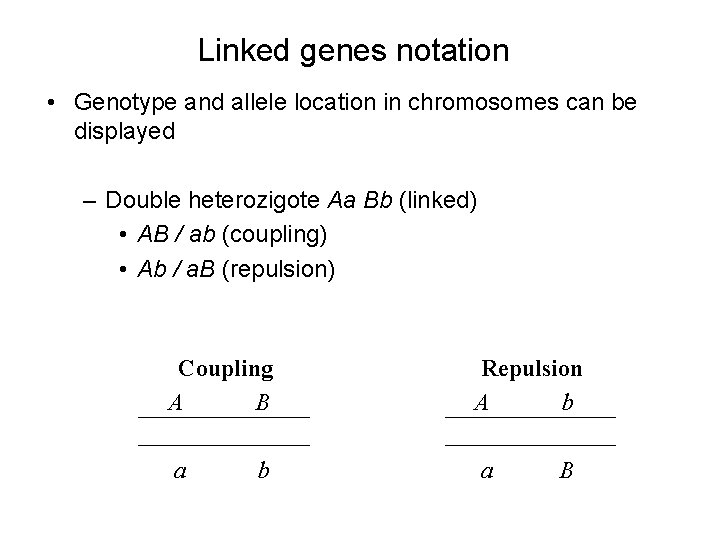
Linked genes notation • Genotype and allele location in chromosomes can be displayed – Double heterozigote Aa Bb (linked) • AB / ab (coupling) • Ab / a. B (repulsion) Coupling A B Repulsion A b a a b B

Recombination frequency • Morgan y Sturtevant (1911): recombination frequencies can be used for making maps, since they reflect the genes separation distances. • pr pr ct ct X pr+ ct+ • Test cross pr ct/ pr ct X pr+ ct+ / ct pr pr + ct/ ct pr pr ct + / ct pr pr ct / ct pr pr + ct + / ct pr TOTAL 21 Recombinants 23 165 Parentals 191 RF=recombinants/total 400
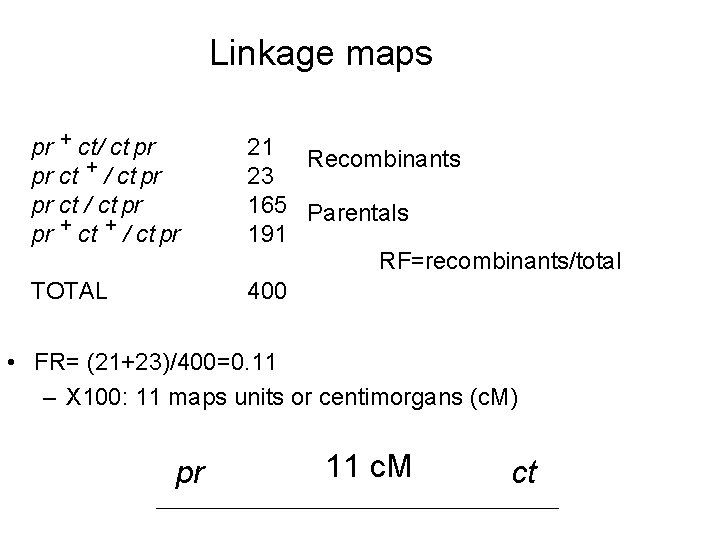
Linkage maps pr + ct/ ct pr pr ct + / ct pr pr ct / ct pr pr + ct + / ct pr TOTAL 21 Recombinants 23 165 Parentals 191 RF=recombinants/total 400 • FR= (21+23)/400=0. 11 – X 100: 11 maps units or centimorgans (c. M) pr 11 c. M ct
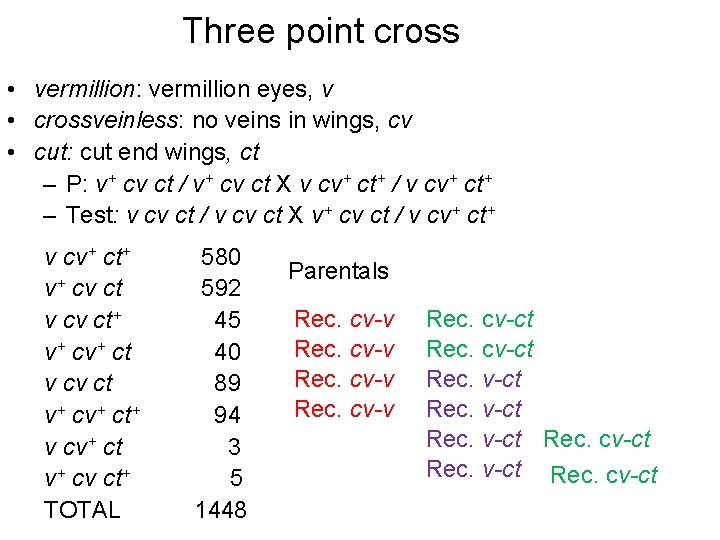
Three point cross • vermillion: vermillion eyes, v • crossveinless: no veins in wings, cv • cut: cut end wings, ct – P: v+ cv ct / v+ cv ct X v cv+ ct+ / v cv+ ct+ – Test: v cv ct / v cv ct X v+ cv ct / v cv+ ct+ v+ cv ct v cv ct+ v+ ct v cv ct v+ ct+ v cv+ ct v+ cv ct+ TOTAL 580 592 45 40 89 94 3 5 1448 Parentals Rec. cv-v Rec. cv-ct Rec. cv-ct
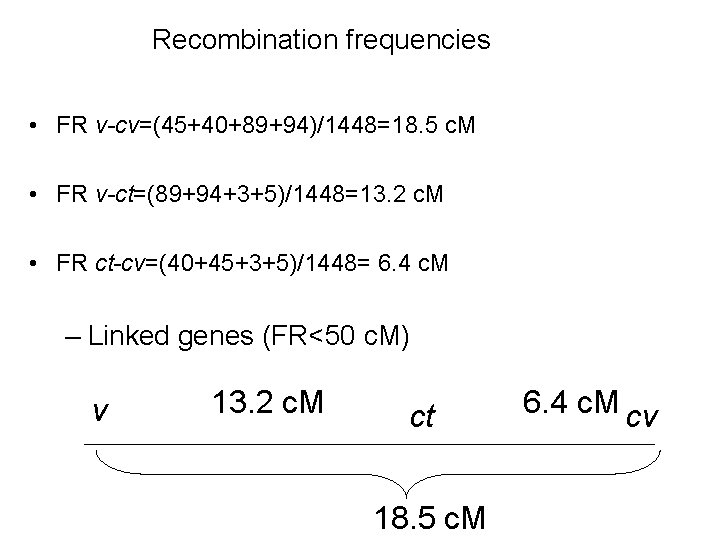
Recombination frequencies • FR v-cv=(45+40+89+94)/1448=18. 5 c. M • FR v-ct=(89+94+3+5)/1448=13. 2 c. M • FR ct-cv=(40+45+3+5)/1448= 6. 4 c. M – Linked genes (FR<50 c. M) v 13. 2 c. M ct 18. 5 c. M 6. 4 c. M cv

Coefficient of coincidence and interference ncv-ct nct-v = 0. 064 v 13. 2 c. M ct 6. 4 c. M cv = 0. 132 §Positive interference: when the occurrence of one crossover reduces the probability that a second one will occur in the same region.

Steps to perform a three point maps 1. Determine phenotypes and numbers of progeny 2. Determine parental genotypes 3. Determine the no recombinant phenotypic classes no recombinants and double ones (more and less frequent) 4. Determine the central locus 5. Determine the order 6. Determine the location of crossovers that led to phenotypic classes 7. Determine the recombination frequencies 8. Construct the map 9. Determine the coefficient of coincidence and the interference
- Slides: 15