3 DSIG 2012 Jul 14 Long Beach California

3 DSIG 2012 Jul 14, Long Beach, California Protein structure alignment beyond spatial proximity Sheng WANG Toyota Technological Institute at Chicago

Related works on Pairwise Structure Alignment 1 Almost all the structure alignment tools 2 TMalign, fr-TMalign 3 DALI, MUSTANG 4 MAMMOTH, Vorolign, YAKUSA 5 FATCAT, CE, MATT, Flex. Prot Note: all proteins we align only consider their C-alpha atom

Our contribution Design a scoring function • local sub-structure similarity • evolutionary and functional information • angular similarity for hydrogen bonding Employ a fast and efficient search algorithm • from highly similar local sub-structures pair (SFP) • recruit new SFPs that satisfies spatial constrains • final refine the alignment within a bound

Scoring Function local similarity global similarity Score(i, j)=( max(0, BLOSUM(i, j) )+CLESUM(i, j) )*v(i, j)*d(i, j) CLESUM is the local structure substitution matrix; BLOSUM is the amino acid substitution matrix; v(i, j) measures the angular similarity using three vectors; d(i, j) measures the spatial proximity of two aligned residues. Note: both v(i, j) and d(i, j) are calculated after rigid-body superposition.
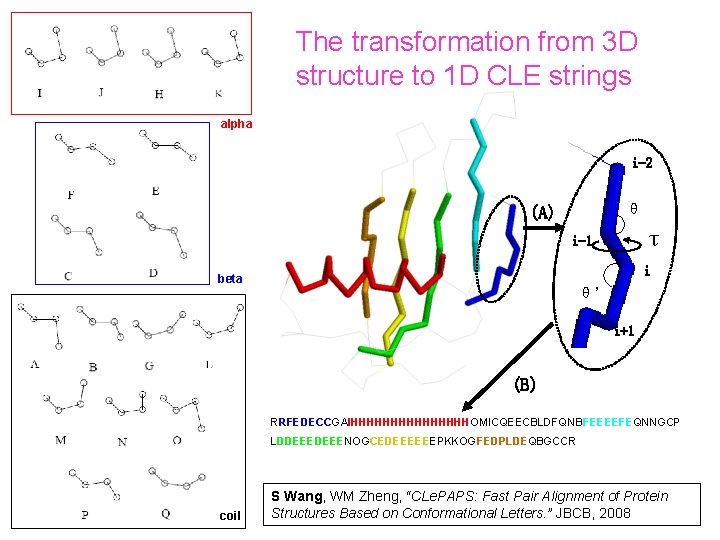
The transformation from 3 D structure to 1 D CLE strings alpha i-2 θ (A) τ i-1 i beta θ’ i+1 (B) RRFEDECCGAIHHHHHHHHOMICQEECBLDFQNBFEEEEFEQNNGCP LDDEEENOGCEDEEEEEEPKKOGFEDPLDEQBGCCR coil S Wang, WM Zheng, “CLe. PAPS: Fast Pair Alignment of Protein Structures Based on Conformational Letters. ” JBCB, 2008

CLESUM : Conformational LEtter SUbstitution Matrix typical helix evolutionary + geometric typical sheet Mij = 20* log 2 (Pij/Pi. Pj) Note: CLESUM is constructed using FSSP representatives.
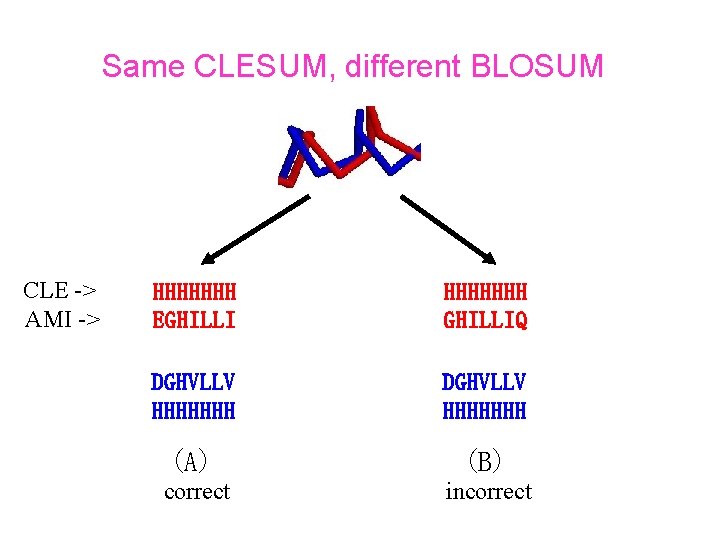
Same CLESUM, different BLOSUM CLE -> AMI -> HHHHHHH EGHILLI HHHHHHH GHILLIQ DGHVLLV HHHHHHH (A) (B) correct incorrect
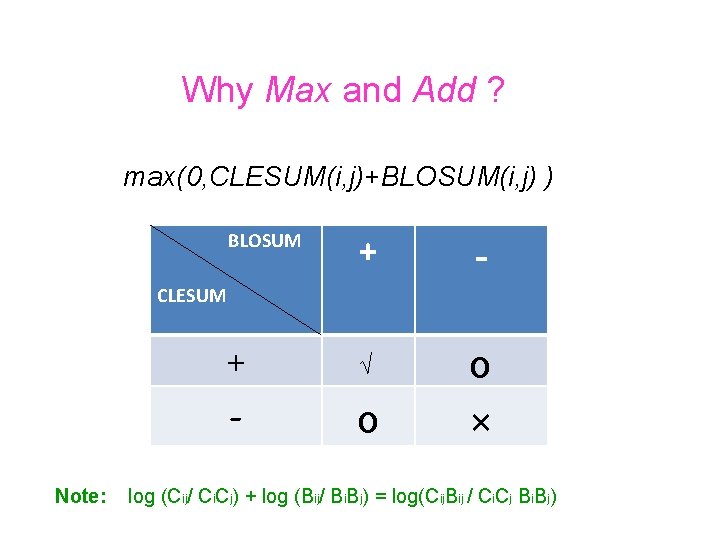
Why Max and Add ? max(0, CLESUM(i, j)+BLOSUM(i, j) ) BLOSUM + - + √ - o o × CLESUM Note: log (Cij/ Ci. Cj) + log (Bij/ Bi. Bj) = log(Cij. Bij / Ci. Cj Bi. Bj)
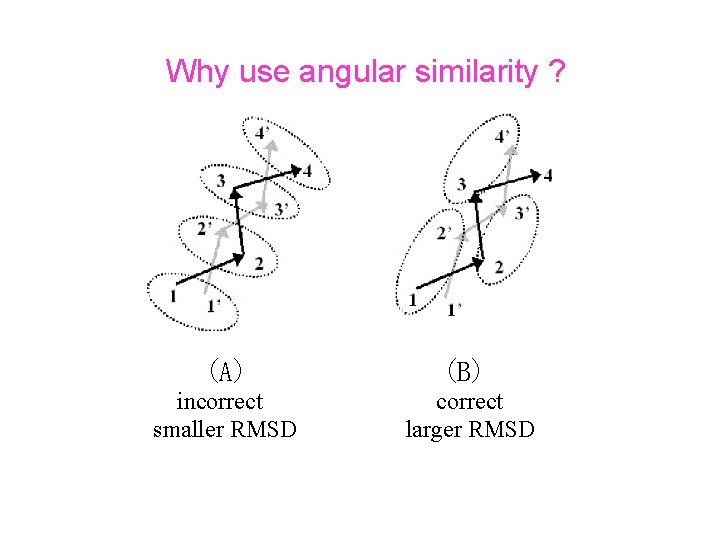
Why use angular similarity ? (A) (B) incorrect smaller RMSD correct larger RMSD
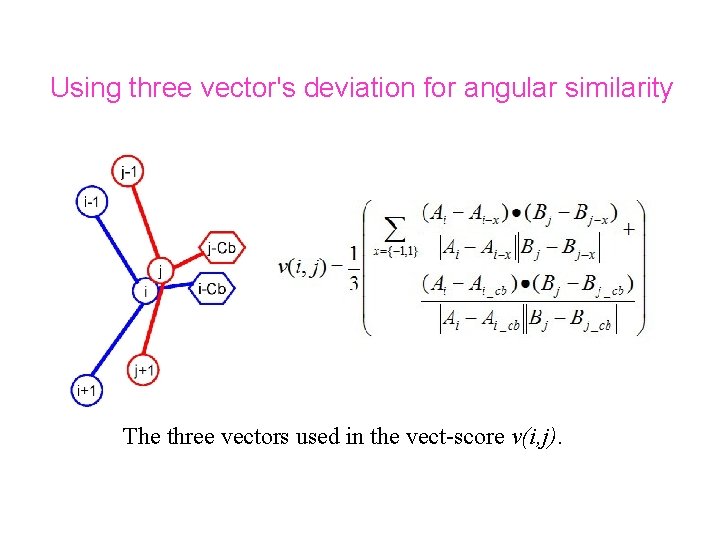
Using three vector's deviation for angular similarity The three vectors used in the vect-score v(i, j).
![Search Algorithm SFP_long SFP_short Sort both SFP lists Deep. Align-score Note: [1] Top. K Search Algorithm SFP_long SFP_short Sort both SFP lists Deep. Align-score Note: [1] Top. K](http://slidetodoc.com/presentation_image_h/ef2ee15fe440291b1bc4afd5f02044a9/image-11.jpg)
Search Algorithm SFP_long SFP_short Sort both SFP lists Deep. Align-score Note: [1] Top. K > Top. J > M [2] SFP stands for Similar Fragment Pair, using ∑max(0, CLESUM(i, j)+BLOSUM(i, j) )
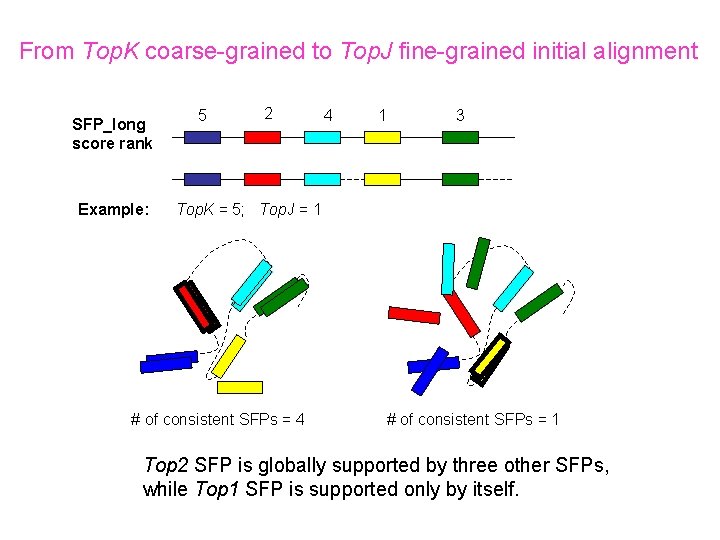
From Top. K coarse-grained to Top. J fine-grained initial alignment SFP_long score rank Example: 5 2 4 1 3 Top. K = 5; Top. J = 1 # of consistent SFPs = 4 # of consistent SFPs = 1 Top 2 SFP is globally supported by three other SFPs, while Top 1 SFP is supported only by itself.
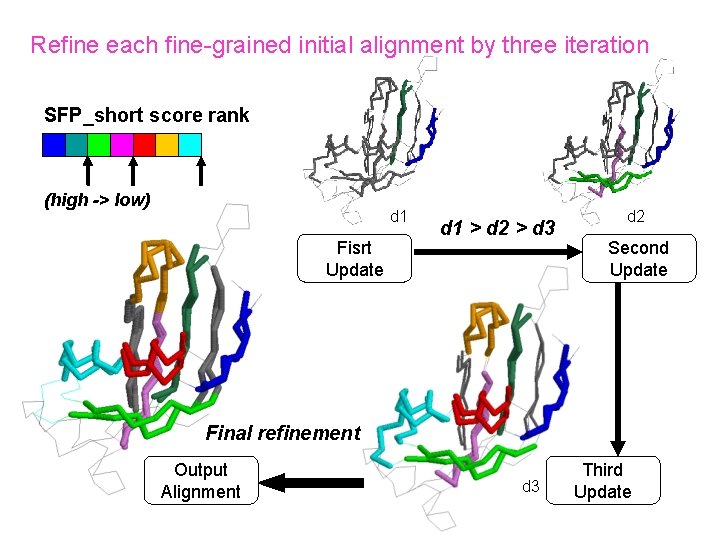
Refine each fine-grained initial alignment by three iteration SFP_short score rank (high -> low) d 1 Fisrt Update d 1 > d 2 > d 3 d 2 Second Update Final refinement Output Alignment d 3 Third Update
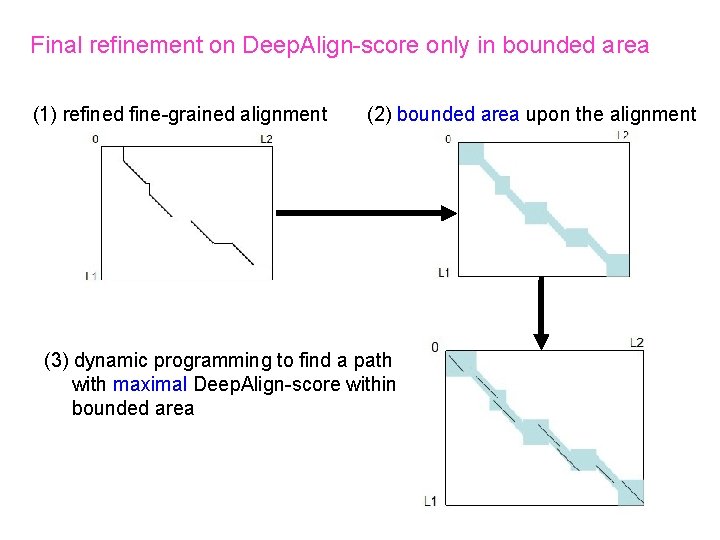
Final refinement on Deep. Align-score only in bounded area (1) refined fine-grained alignment (2) bounded area upon the alignment (3) dynamic programming to find a path with maximal Deep. Align-score within bounded area
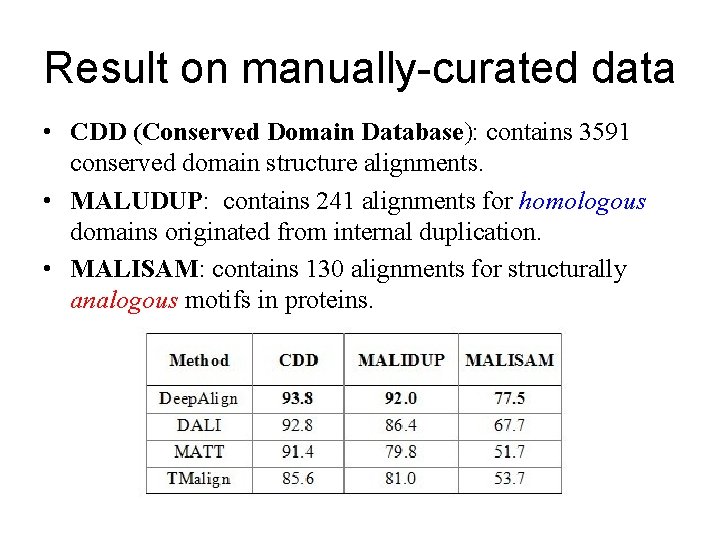
Result on manually-curated data • CDD (Conserved Domain Database): contains 3591 conserved domain structure alignments. • MALUDUP: contains 241 alignments for homologous domains originated from internal duplication. • MALISAM: contains 130 alignments for structurally analogous motifs in proteins.
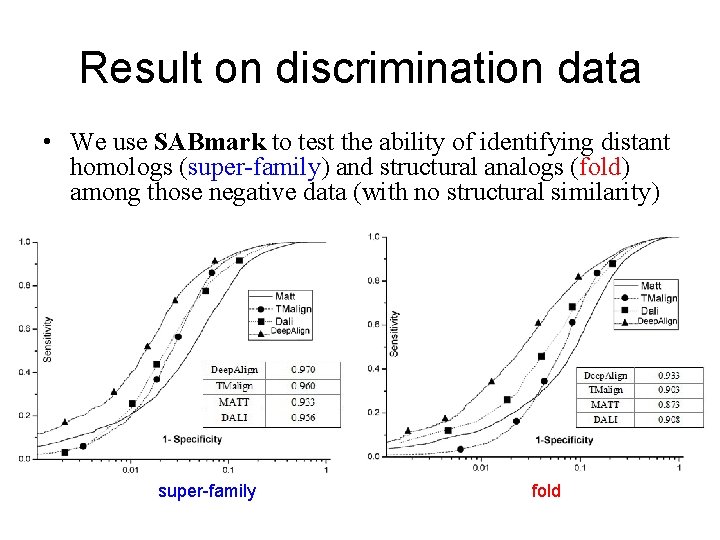
Result on discrimination data • We use SABmark to test the ability of identifying distant homologs (super-family) and structural analogs (fold) among those negative data (with no structural similarity) super-family fold
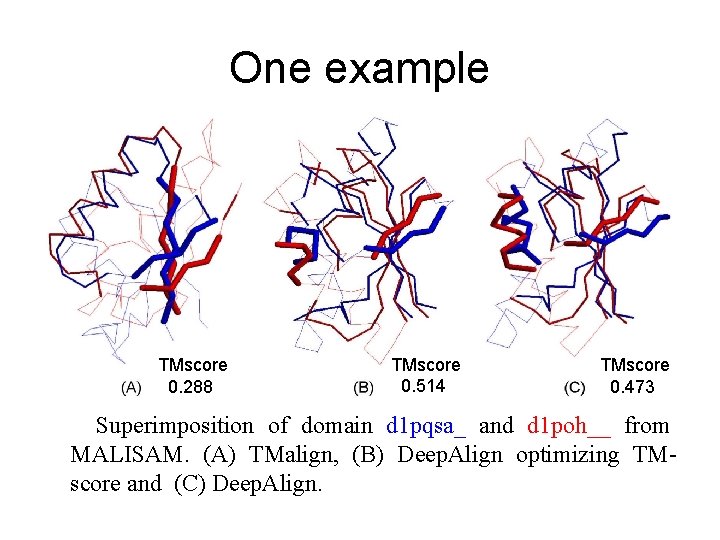
One example TMscore 0. 288 TMscore 0. 514 TMscore 0. 473 Superimposition of domain d 1 pqsa_ and d 1 poh__ from MALISAM. (A) TMalign, (B) Deep. Align optimizing TMscore and (C) Deep. Align.
Thank you !! Please find the executable program of Deep. Align at: http: //ttic. uchicago. edu/~jinbo/Deep. Align_exe_V 1. 00. tar. gz
- Slides: 18