3 D structure Swiss Pdb Viewer David Shiuan

3 D structure -Swiss Pdb Viewer David Shiuan Department of Life Science, Institute of Biotechnology and Interdisciplinary Program of Bioinformatics National Dong Hwa University

Swiss-Pdb. Viewer n n a tool for viewing and manipulating protein structures and models. tightly linked to Swiss-Model, an automated homology modeling server.

Swiss-Pdb. Viewer n n provides a user friendly interface allowing to analyze several proteins at the same time. The proteins can be superimposed in order to deduce structural alignments and compare their active sites or any other relevant parts. Amino acid mutations, H-bonds, angles and distances between atoms are easy to obtain thanks to the intuitive graphic and menu interface.


































Welcome to the SWISS-MODEL Protein Modelling Server n Dear David Shiuan, Title of your Request: lipase 3 D Swiss-Model '(SWISS-MODEL Version 36. 0003)' started on 16 -Dec-2005 Process identification is AAAa 03 u. OW The modelling procedure is now in progress, and its results should be sent to you shortly. In case of difficulties, please contact: Torsten Schwede <Torsten. Schwede@isb-sib. ch> Nicolas Guex

News from SWISS-MODEL: - New Server Hardware - Deep. View - Swiss. PDB-Viewer - Ex. PDB template database Good luck with your projects. The SWISS-MODEL Team Dr. Torsten Schwede Torsten. Schwede@isbsib. ch Dr. Nicolas Guex Dr. Manuel C. Peitsch

Align. Master output ======= Length of target sequence: 288 residues n Searching sequences of known 3 D structures Found 1 f 3 r. B. pdb with P(N)=6 e-76 Found 1 dzb. B. pdb with P(N)=2 e-70 Found 1 dzb. A. pdb with P(N)=2 e-70 Found 1 nqb. C. pdb with P(N)=1 e-66 Found 1 nqb. A. pdb with P(N)=1 e-66 Found 1 qok. A. pdb with P(N)=2 e-65 Found 1 lmk. G. pdb with P(N)=5 e-64 Found 1 lmk. C. pdb with P(N)=5 e-64 Found 1 lmk. E. pdb with P(N)=5 e-64 Found 1 lmk. A. pdb with P(N)=5 e-64 Found 1 dee. C. pdb with P(N)=8 e-55

Please find enclosed your results for AAAa 03 u. OW as an attachment. Title : lipase 3 D Jobid: AAAa 03 u. OW Batch. 1 n SWISS-MODEL project files are best viewed using the program - Deep. View, Swiss. Pdb. Viewer (http: //www. expasy. org/spdbv/). n The program allows you to display and manipulate several structures at the same time, e. g. the model and all the superposed templates.

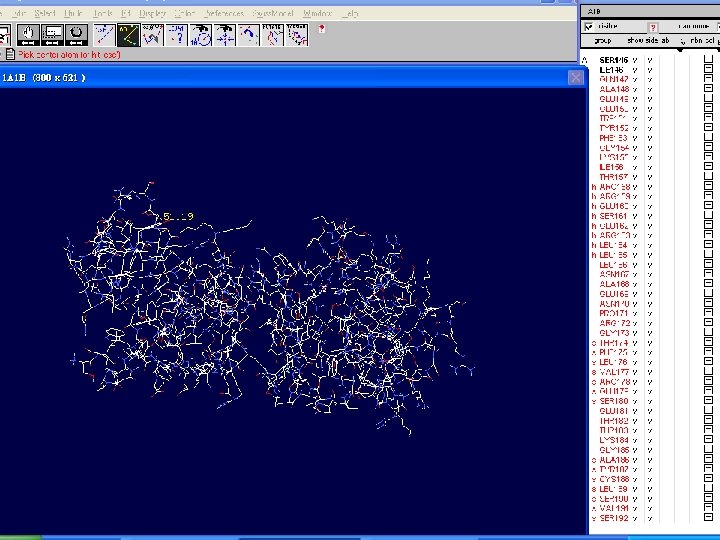
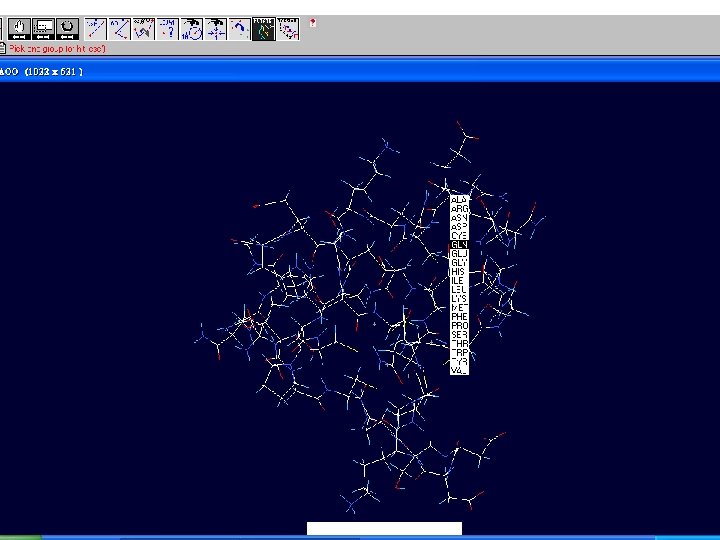
How to visualize and analyze the Swiss. Model generated structures n The result file can be visualized by Swiss-pdb Viewer or Ras. Win (Ras. Mol). n Use the Control Panel to remove all of the templates. n Select known 3 D structures of lipase (for example). n Compare the target sequence with the known lipase structure by magic fit of Swiss-pdb Viewer. n Use center the molecule on one atom tool bar to focus on the region of binding site. n Compare the above two structures.
Hermicola (Thermomyces) lanuginaosa lipase (1 DU 4)
Swiss Model – Target (Ab) with 3 templates
Target (Ab) model
Target and lipase i. DU 4
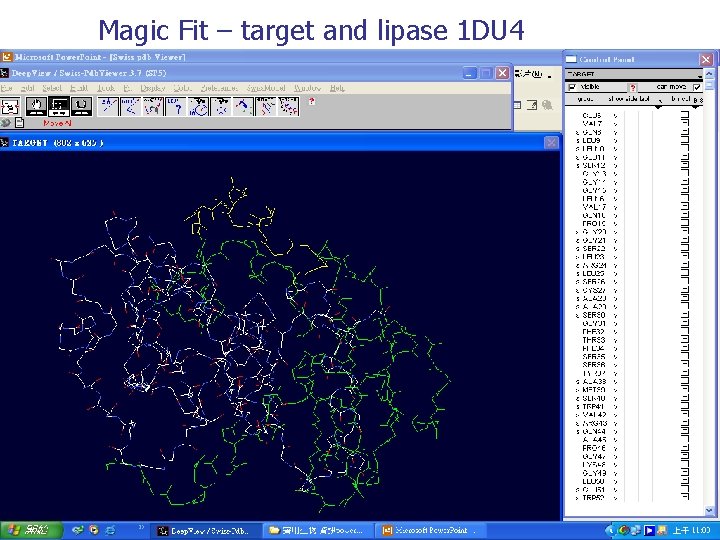
Magic Fit – target and lipase 1 DU 4
- Slides: 50