2016 12 02 JinSun Ryu Introduction se DSBs

2016. 12. 02 Jin-Sun Ryu
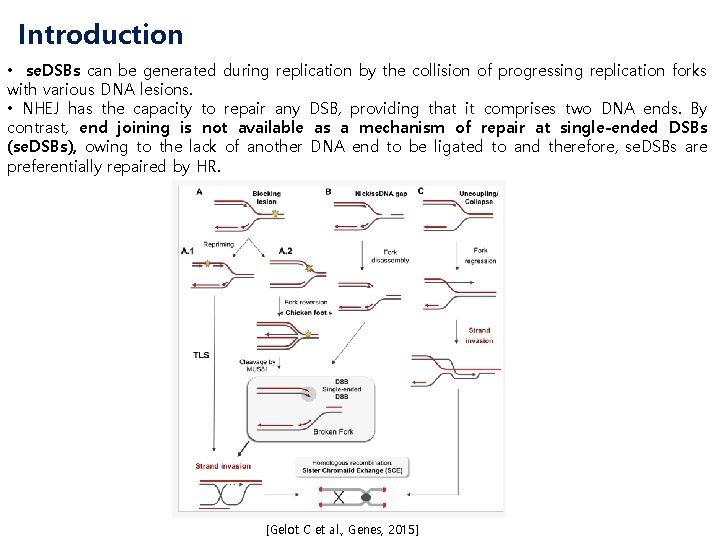
Introduction • se. DSBs can be generated during replication by the collision of progressing replication forks with various DNA lesions. • NHEJ has the capacity to repair any DSB, providing that it comprises two DNA ends. By contrast, end joining is not available as a mechanism of repair at single-ended DSBs (se. DSBs), owing to the lack of another DNA end to be ligated to and therefore, se. DSBs are preferentially repaired by HR. [Gelot C et al. , Genes, 2015]
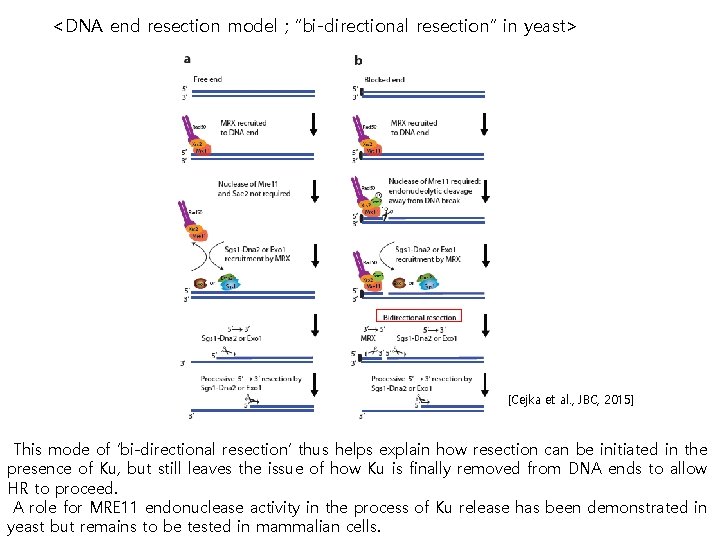
<DNA end resection model ; “bi-directional resection” in yeast> [Cejka et al. , JBC, 2015] This mode of ‘bi-directional resection’ thus helps explain how resection can be initiated in the presence of Ku, but still leaves the issue of how Ku is finally removed from DNA ends to allow HR to proceed. A role for MRE 11 endonuclease activity in the process of Ku release has been demonstrated in yeast but remains to be tested in mammalian cells.
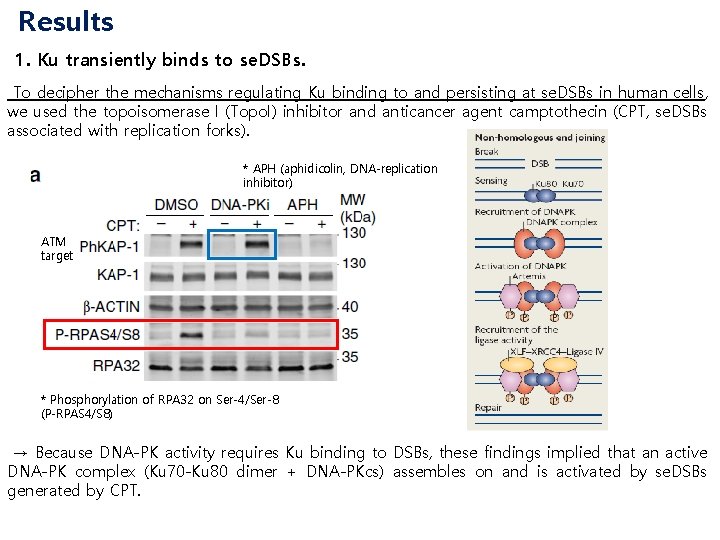
Results 1. Ku transiently binds to se. DSBs. To decipher the mechanisms regulating Ku binding to and persisting at se. DSBs in human cells, we used the topoisomerase I (Topo. I) inhibitor and anticancer agent camptothecin (CPT, se. DSBs associated with replication forks). * APH (aphidicolin, DNA-replication inhibitor) ATM target * Phosphorylation of RPA 32 on Ser-4/Ser-8 (P-RPAS 4/S 8) → Because DNA-PK activity requires Ku binding to DSBs, these findings implied that an active DNA-PK complex (Ku 70 -Ku 80 dimer + DNA-PKcs) assembles on and is activated by se. DSBs generated by CPT.
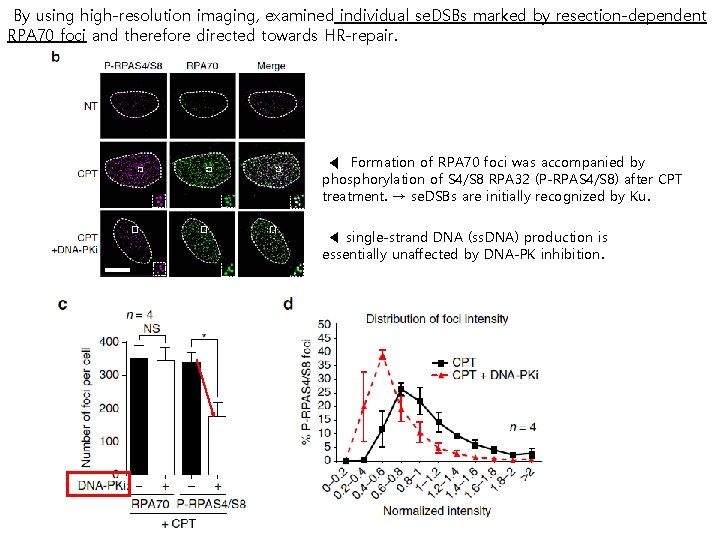
By using high-resolution imaging, examined individual se. DSBs marked by resection-dependent RPA 70 foci and therefore directed towards HR-repair. ◀ Formation of RPA 70 foci was accompanied by phosphorylation of S 4/S 8 RPA 32 (P-RPAS 4/S 8) after CPT treatment. → se. DSBs are initially recognized by Ku. ◀ single-strand DNA (ss. DNA) production is essentially unaffected by DNA-PK inhibition.
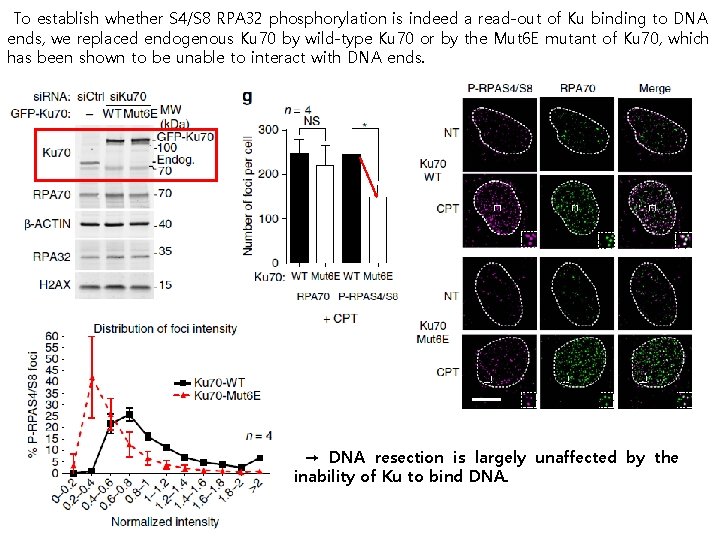
To establish whether S 4/S 8 RPA 32 phosphorylation is indeed a read-out of Ku binding to DNA ends, we replaced endogenous Ku 70 by wild-type Ku 70 or by the Mut 6 E mutant of Ku 70, which has been shown to be unable to interact with DNA ends. → DNA resection is largely unaffected by the inability of Ku to bind DNA.
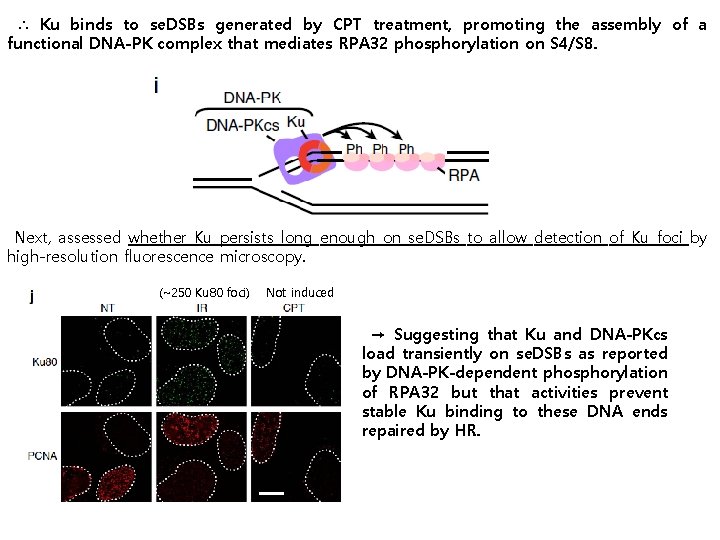
∴ Ku binds to se. DSBs generated by CPT treatment, promoting the assembly of a functional DNA-PK complex that mediates RPA 32 phosphorylation on S 4/S 8. Next, assessed whether Ku persists long enough on se. DSBs to allow detection of Ku foci by high-resolution fluorescence microscopy. (~250 Ku 80 foci) Not induced → Suggesting that Ku and DNA-PKcs load transiently on se. DSBs as reported by DNA-PK-dependent phosphorylation of RPA 32 but that activities prevent stable Ku binding to these DNA ends repaired by HR.
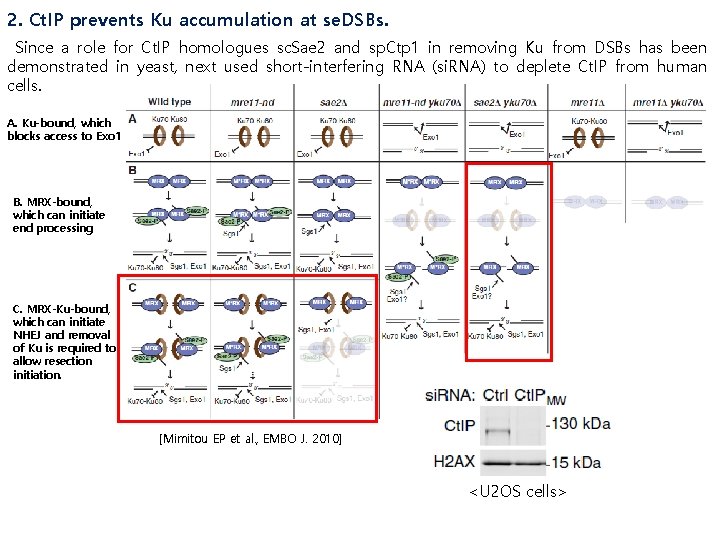
2. Ct. IP prevents Ku accumulation at se. DSBs. Since a role for Ct. IP homologues sc. Sae 2 and sp. Ctp 1 in removing Ku from DSBs has been demonstrated in yeast, next used short-interfering RNA (si. RNA) to deplete Ct. IP from human cells. A. Ku-bound, which blocks access to Exo 1 B. MRX-bound, which can initiate end processing C. MRX-Ku-bound, which can initiate NHEJ and removal of Ku is required to allow resection initiation. [Mimitou EP et al. , EMBO J. 2010] <U 2 OS cells>
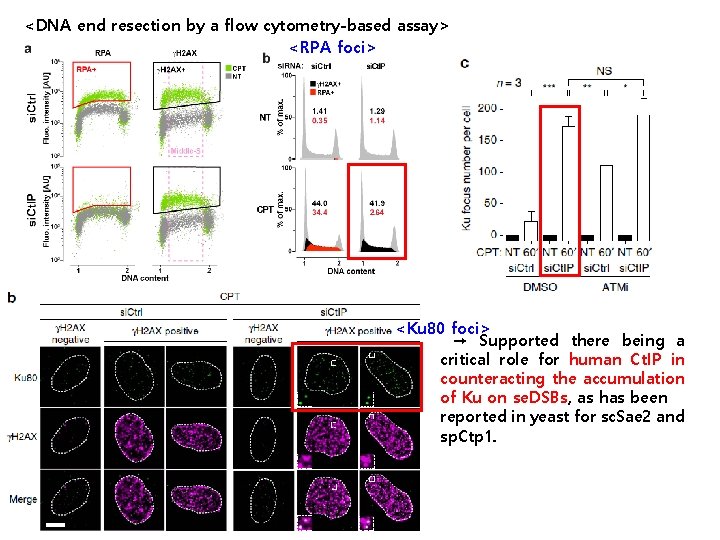
<DNA end resection by a flow cytometry-based assay> <RPA foci> <Ku 80 foci> → Supported there being a critical role for human Ct. IP in counteracting the accumulation of Ku on se. DSBs, as has been reported in yeast for sc. Sae 2 and sp. Ctp 1.
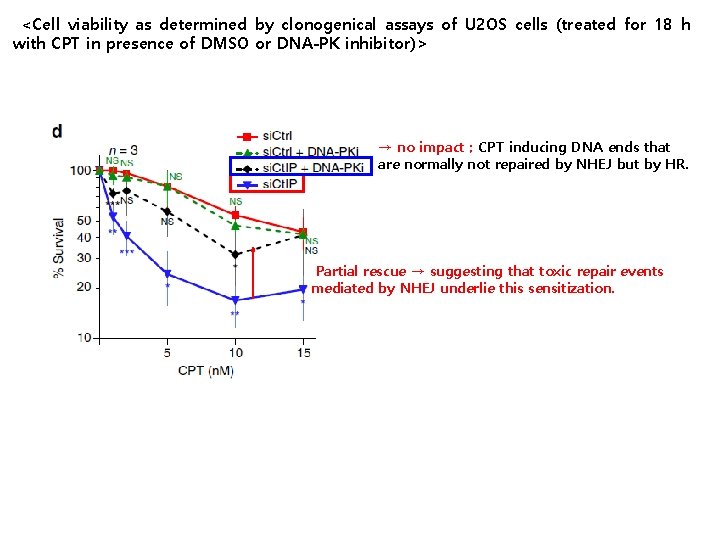
<Cell viability as determined by clonogenical assays of U 2 OS cells (treated for 18 h with CPT in presence of DMSO or DNA-PK inhibitor)> → no impact ; CPT inducing DNA ends that are normally not repaired by NHEJ but by HR. Partial rescue → suggesting that toxic repair events mediated by NHEJ underlie this sensitization.
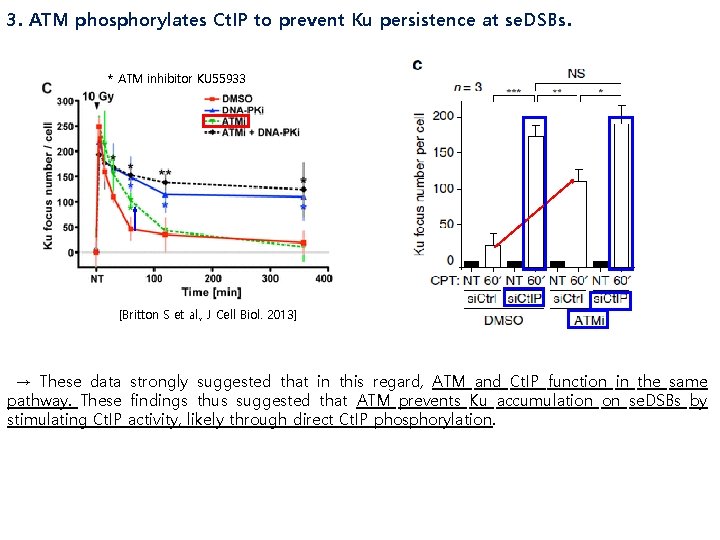
3. ATM phosphorylates Ct. IP to prevent Ku persistence at se. DSBs. * ATM inhibitor KU 55933 [Britton S et al. , J Cell Biol. 2013] → These data strongly suggested that in this regard, ATM and Ct. IP function in the same pathway. These findings thus suggested that ATM prevents Ku accumulation on se. DSBs by stimulating Ct. IP activity, likely through direct Ct. IP phosphorylation.
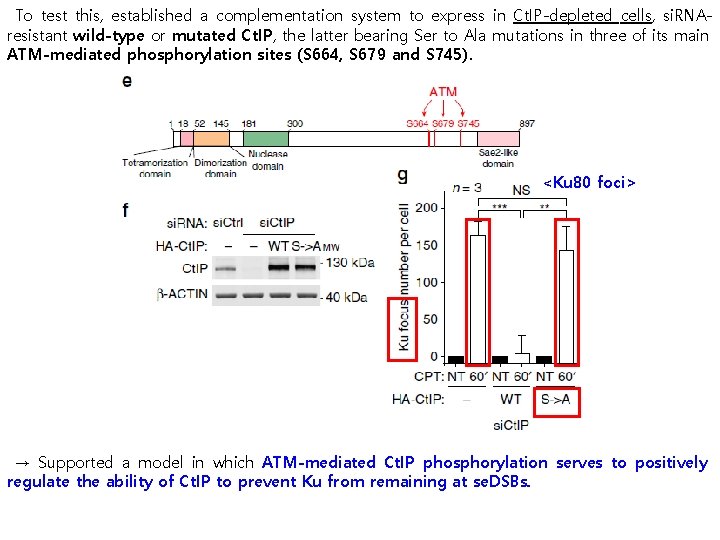
To test this, established a complementation system to express in Ct. IP-depleted cells, si. RNAresistant wild-type or mutated Ct. IP, the latter bearing Ser to Ala mutations in three of its main ATM-mediated phosphorylation sites (S 664, S 679 and S 745). <Ku 80 foci> → Supported a model in which ATM-mediated Ct. IP phosphorylation serves to positively regulate the ability of Ct. IP to prevent Ku from remaining at se. DSBs.
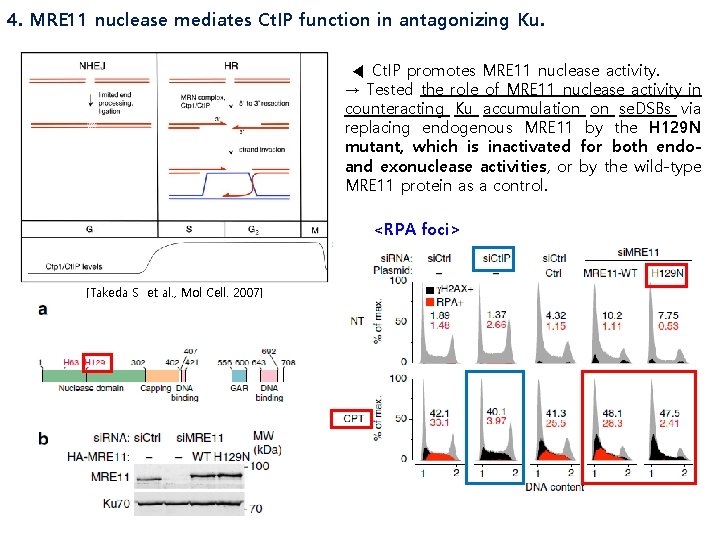
4. MRE 11 nuclease mediates Ct. IP function in antagonizing Ku. ◀ Ct. IP promotes MRE 11 nuclease activity. → Tested the role of MRE 11 nuclease activity in counteracting Ku accumulation on se. DSBs via replacing endogenous MRE 11 by the H 129 N mutant, which is inactivated for both endoand exonuclease activities, or by the wild-type MRE 11 protein as a control. <RPA foci> [Takeda S et al. , Mol Cell. 2007]
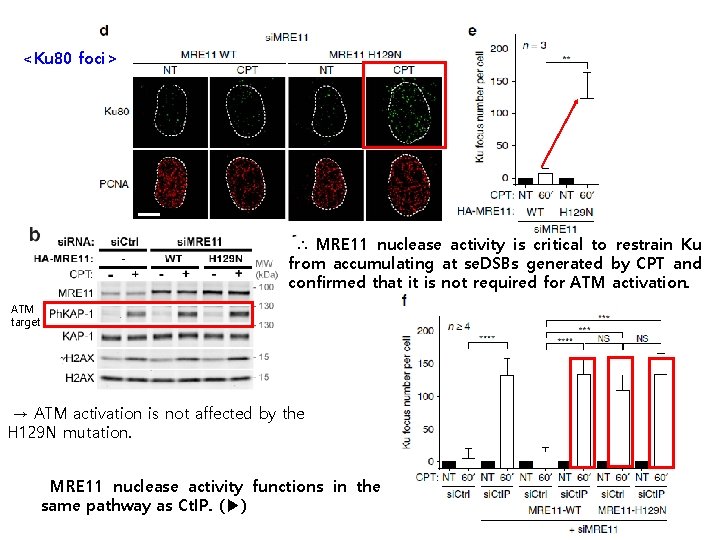
<Ku 80 foci> ∴ MRE 11 nuclease activity is critical to restrain Ku from accumulating at se. DSBs generated by CPT and confirmed that it is not required for ATM activation. ATM target → ATM activation is not affected by the H 129 N mutation. MRE 11 nuclease activity functions in the same pathway as Ct. IP. (▶)
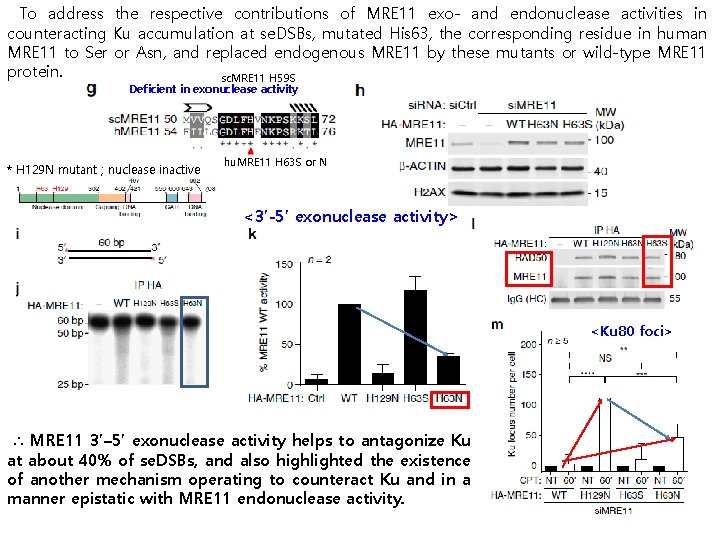
To address the respective contributions of MRE 11 exo- and endonuclease activities in counteracting Ku accumulation at se. DSBs, mutated His 63, the corresponding residue in human MRE 11 to Ser or Asn, and replaced endogenous MRE 11 by these mutants or wild-type MRE 11 protein. sc. MRE 11 H 59 S Deficient in exonuclease activity * H 129 N mutant ; nuclease inactive hu. MRE 11 H 63 S or N <3’-5’ exonuclease activity> <Ku 80 foci> ∴ MRE 11 3’– 5’ exonuclease activity helps to antagonize Ku at about 40% of se. DSBs, and also highlighted the existence of another mechanism operating to counteract Ku and in a manner epistatic with MRE 11 endonuclease activity.
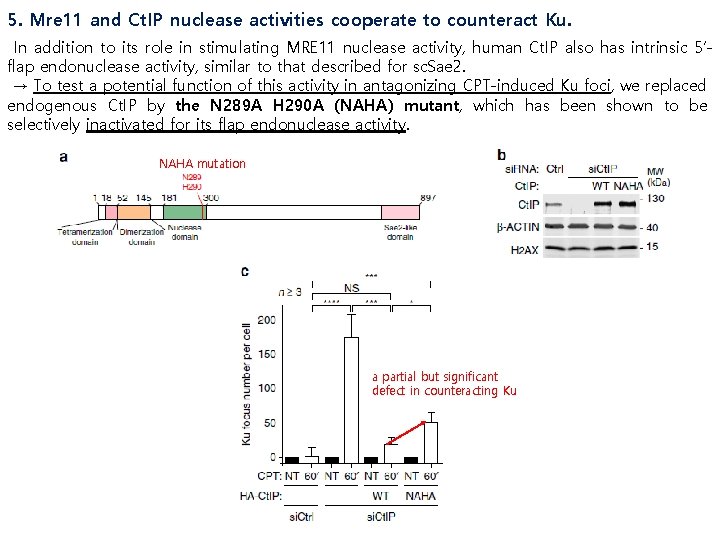
5. Mre 11 and Ct. IP nuclease activities cooperate to counteract Ku. In addition to its role in stimulating MRE 11 nuclease activity, human Ct. IP also has intrinsic 5’flap endonuclease activity, similar to that described for sc. Sae 2. → To test a potential function of this activity in antagonizing CPT-induced Ku foci, we replaced endogenous Ct. IP by the N 289 A H 290 A (NAHA) mutant, which has been shown to be selectively inactivated for its flap endonuclease activity. NAHA mutation a partial but significant defect in counteracting Ku
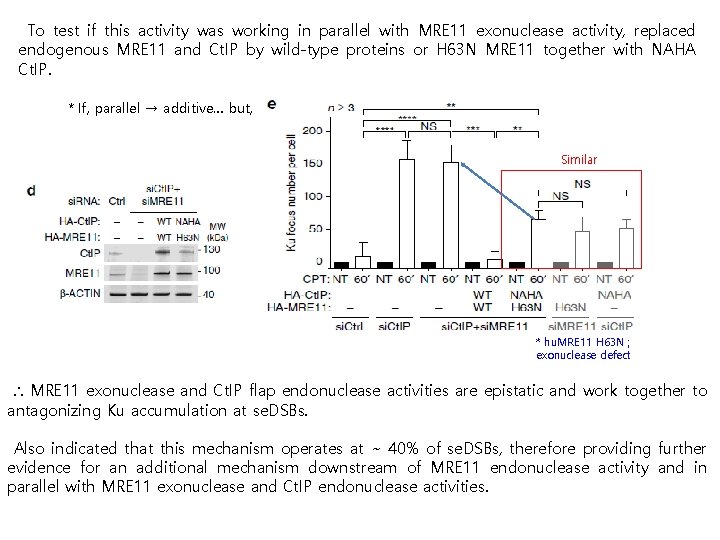
To test if this activity was working in parallel with MRE 11 exonuclease activity, replaced endogenous MRE 11 and Ct. IP by wild-type proteins or H 63 N MRE 11 together with NAHA Ct. IP. * If, parallel → additive… but, Similar * hu. MRE 11 H 63 N ; exonuclease defect ∴ MRE 11 exonuclease and Ct. IP flap endonuclease activities are epistatic and work together to antagonizing Ku accumulation at se. DSBs. Also indicated that this mechanism operates at ~ 40% of se. DSBs, therefore providing further evidence for an additional mechanism downstream of MRE 11 endonuclease activity and in parallel with MRE 11 exonuclease and Ct. IP endonuclease activities.
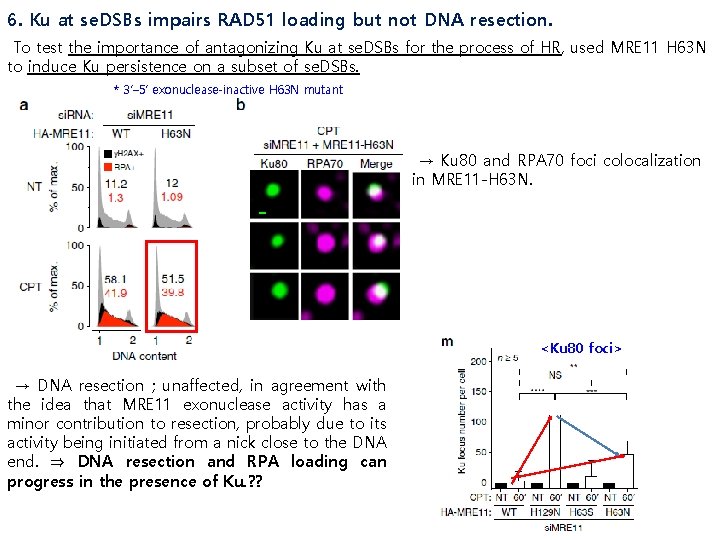
6. Ku at se. DSBs impairs RAD 51 loading but not DNA resection. To test the importance of antagonizing Ku at se. DSBs for the process of HR, used MRE 11 H 63 N to induce Ku persistence on a subset of se. DSBs. * 3’– 5’ exonuclease-inactive H 63 N mutant → Ku 80 and RPA 70 foci colocalization in MRE 11 -H 63 N. <Ku 80 foci> → DNA resection ; unaffected, in agreement with the idea that MRE 11 exonuclease activity has a minor contribution to resection, probably due to its activity being initiated from a nick close to the DNA end. ⇒ DNA resection and RPA loading can progress in the presence of Ku. ? ?
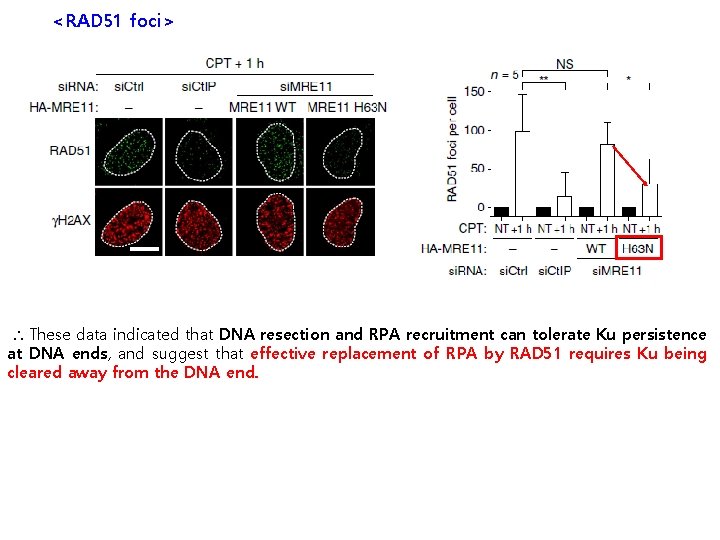
<RAD 51 foci> ∴ These data indicated that DNA resection and RPA recruitment can tolerate Ku persistence at DNA ends, and suggest that effective replacement of RPA by RAD 51 requires Ku being cleared away from the DNA end.
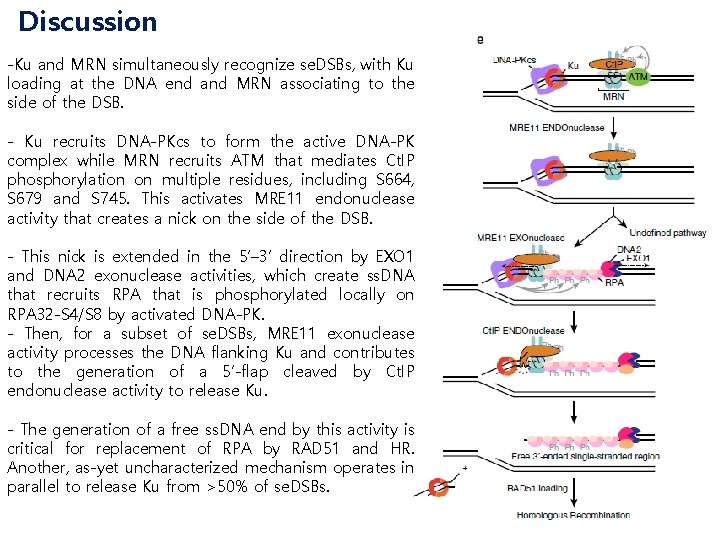
Discussion -Ku and MRN simultaneously recognize se. DSBs, with Ku loading at the DNA end and MRN associating to the side of the DSB. - Ku recruits DNA-PKcs to form the active DNA-PK complex while MRN recruits ATM that mediates Ct. IP phosphorylation on multiple residues, including S 664, S 679 and S 745. This activates MRE 11 endonuclease activity that creates a nick on the side of the DSB. - This nick is extended in the 5’– 3’ direction by EXO 1 and DNA 2 exonuclease activities, which create ss. DNA that recruits RPA that is phosphorylated locally on RPA 32 -S 4/S 8 by activated DNA-PK. - Then, for a subset of se. DSBs, MRE 11 exonuclease activity processes the DNA flanking Ku and contributes to the generation of a 5’-flap cleaved by Ct. IP endonuclease activity to release Ku. - The generation of a free ss. DNA end by this activity is critical for replacement of RPA by RAD 51 and HR. Another, as-yet uncharacterized mechanism operates in parallel to release Ku from >50% of se. DSBs.
- Slides: 20