2014 NSFCMACS Workshop on Cellular Signaling Pathways Extending












- Slides: 21

2014 NSF-CMACS Workshop on Cellular Signaling Pathways Extending The Network Analysis of Oncogenic Ras Activation In Cancer (Stites, et. al. ) Model With Positive Feedback From The Allosteric Binding Of Ras To The SOS GEF Complex. Group DAN Dylan Sun Alessandro Rosa Neno Fuller Wikipedia

Shape In Biology https: //www. gameinformer. com/blogs/members/b/subsaint_blog/archive/2013/06/14/the-push-towards-digital-distribution. aspx “Structure and Function are Correlated at All Levels of Biological Organization” – Campbell Biology, Ninth Edition In Biology, Shape Is Everything

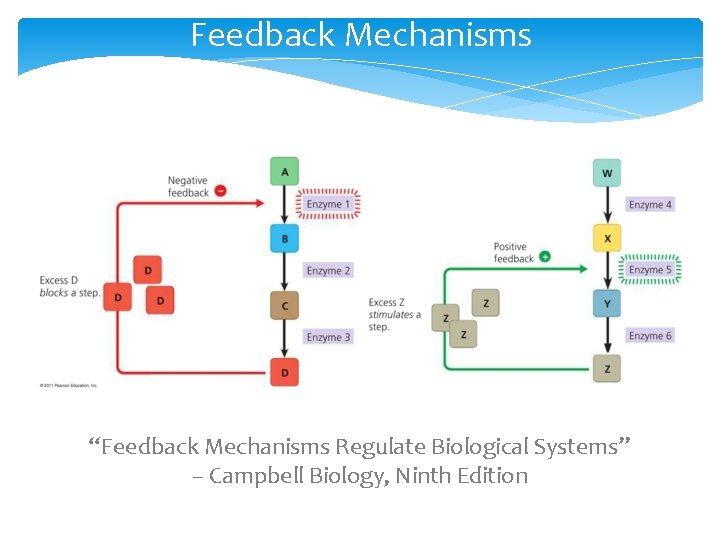
Feedback Mechanisms “Feedback Mechanisms Regulate Biological Systems” – Campbell Biology, Ninth Edition

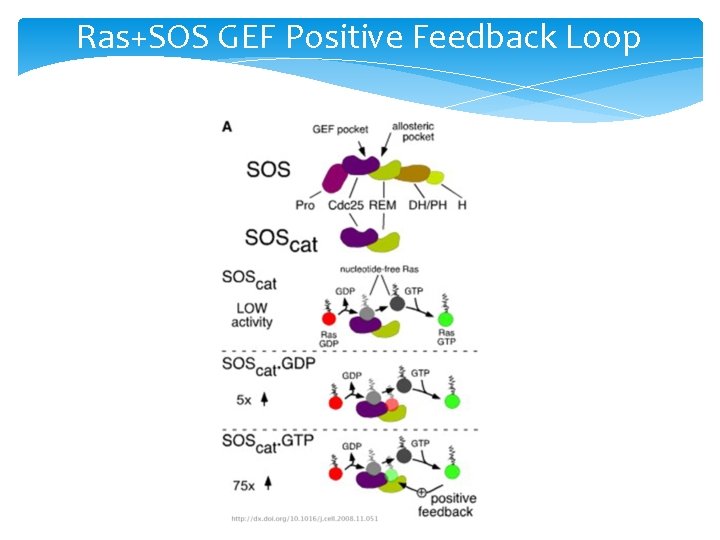
Ras+SOS GEF Positive Feedback Loop

Bio. Net. Gen Reaction Rules For Allosteric Feedback Loop Positive feedback loop:

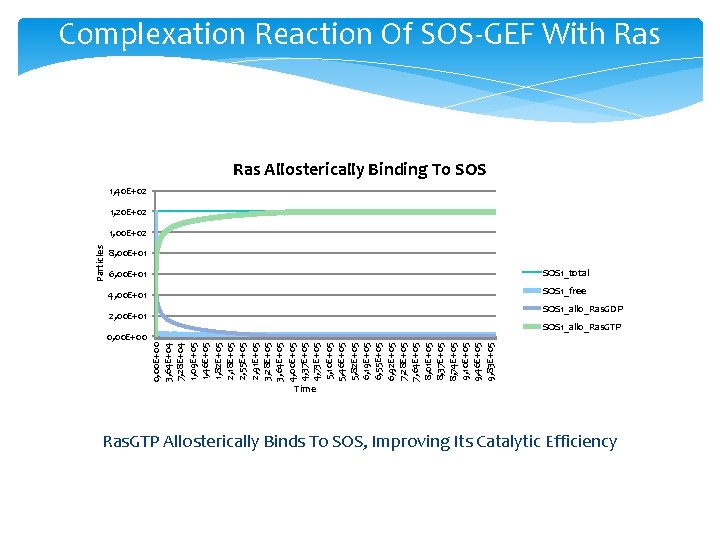
Complexation Reaction Of SOS-GEF With Ras Allosterically Binding To SOS 1, 40 E+02 1, 20 E+02 Particles 1, 00 E+02 8, 00 E+01 6, 00 E+01 SOS 1_total 4, 00 E+01 SOS 1_free SOS 1_allo_Ras. GDP 2, 00 E+01 SOS 1_allo_Ras. GTP 0, 00 E+00 3, 64 E+04 7, 28 E+04 1, 09 E+05 1, 46 E+05 1, 82 E+05 2, 18 E+05 2, 55 E+05 2, 91 E+05 3, 28 E+05 3, 64 E+05 4, 00 E+05 4, 37 E+05 4, 73 E+05 5, 10 E+05 5, 46 E+05 5, 82 E+05 6, 19 E+05 6, 55 E+05 6, 92 E+05 7, 28 E+05 7, 64 E+05 8, 01 E+05 8, 37 E+05 8, 74 E+05 9, 10 E+05 9, 46 E+05 9, 83 E+05 0, 00 E+00 Time Ras. GTP Allosterically Binds To SOS, Improving Its Catalytic Efficiency

Results Of Guanine Nucleotide Exchange Factors Transfer Reaction With Allosteric Feedback Loop Ø Original Model Based on Reaction Rates in Stites Paper Ø Produce Ras. GTP more quickly than positive Feedback Model Ø Positive Feedback Model Ø Initial reactions catalyzed at the relatively slower SOS • Ras. GDP Reaction Rate Ø As number of Ras. GTP particles increases, the reaction is catalyzed and the relatively faster SOS • Ras. GTP Reaction Rate Ø Positive Feedback Model shows that reaction rate without feedback in Original Model overestimates the production of Ras. GTP in the initial stages of signal transduction

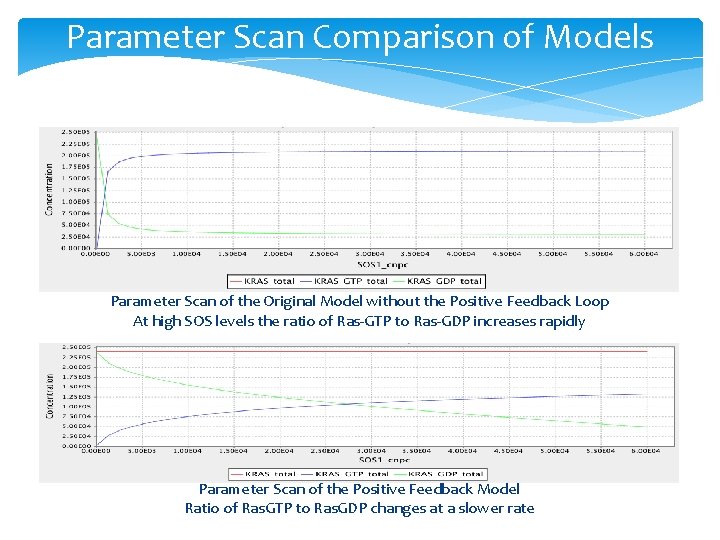
Parameter Scan Comparison of Models Parameter Scan of the Original Model without the Positive Feedback Loop At high SOS levels the ratio of Ras-GTP to Ras-GDP increases rapidly Parameter Scan of the Positive Feedback Model Ratio of Ras. GTP to Ras. GDP changes at a slower rate

Effect Of Rules For Uncatalyzed Removal Of Guanine Nucleotides From Ras On Positive Feedback Model Bio. Net. Gen Reaction Rules for the Uncatalyzed Removal of Guanine Nucleotides from Ras Positive Feedback Loop Model Results excluding uncatalyzed removal of GTP and GDP from Ras 3, 00 E+05 2, 00 E+05 1, 50 E+05 KRAS_total 1, 00 E+05 KRAS_GTP_total 5, 00 E+04 KRAS_GDP_total 0, 00 E+00 4, 84 E+04 9, 68 E+04 1, 45 E+05 1, 94 E+05 2, 42 E+05 2, 90 E+05 3, 39 E+05 3, 87 E+05 4, 36 E+05 4, 84 E+05 5, 32 E+05 5, 81 E+05 6, 29 E+05 6, 78 E+05 7, 26 E+05 7, 74 E+05 8, 23 E+05 8, 71 E+05 9, 20 E+05 9, 68 E+05 Particles 2, 50 E+05 Time

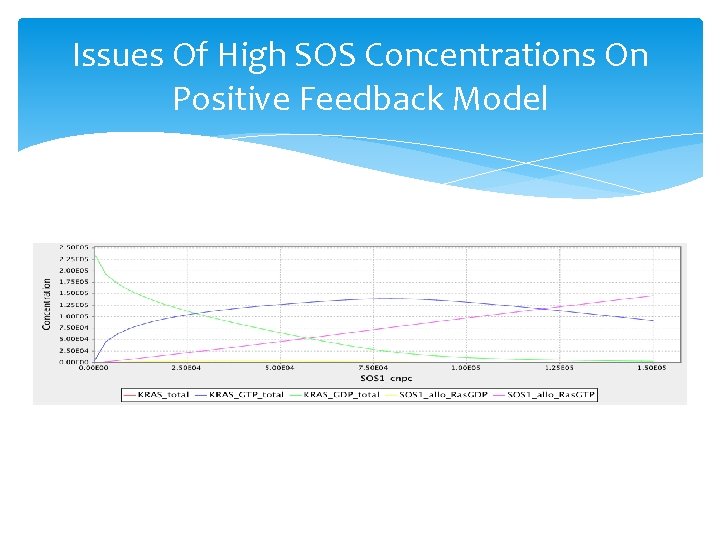
Issues Of High SOS Concentrations On Positive Feedback Model

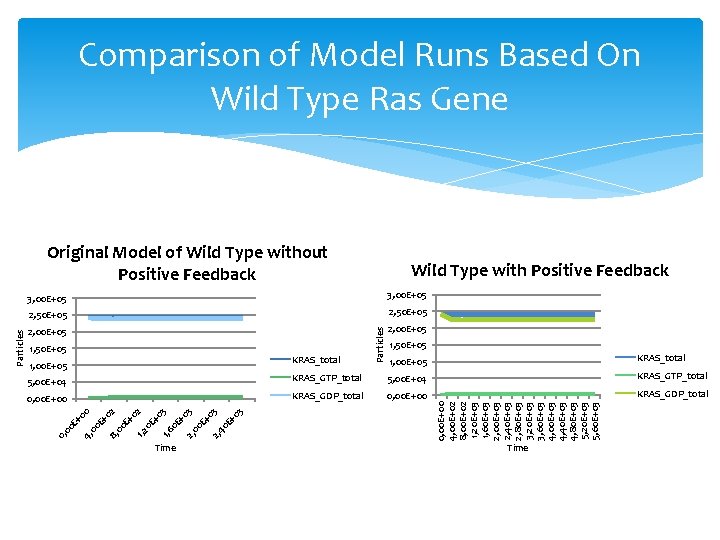
Comparison of Model Runs Based On Wild Type Ras Gene Wild Type with Positive Feedback 3, 00 E+05 2, 50 E+05 2, 00 E+05 1, 50 E+05 KRAS_total 1, 00 E+05 Particles 3, 00 E+05 1, 50 E+05 1, 00 E+05 KRAS_total 0, 00 E+00 KRAS_GDP_total 1, 2 00 E+ 0 8, 00 E+ 4, 00 E+ Time 0, 00 E+00 4, 00 E+02 8, 00 E+02 1, 20 E+03 1, 60 E+03 2, 00 E+03 2, 40 E+03 2, 80 E+03 3, 20 E+03 3, 60 E+03 4, 00 E+03 4, 40 E+03 4, 80 E+03 5, 20 E+03 5, 60 E+03 KRAS_GTP_total 02 0 E +0 1, 6 3 0 E +0 3 2, 00 E+ 03 2, 40 E+ 03 5, 00 E+04 2 KRAS_GTP_total 00 5, 00 E+04 0, Particles Original Model of Wild Type without Positive Feedback Time

Adding GAP – Insensitivity Mutant RAS to Positive Feedback Model Ø A relatively common form of mutant is G 12 V RAS Ø Ras binds to GAP but GTP is not hydrolyzed Ø This mutation cause Ras to remain in its active form when it is activated and leads to the overproduction of proteins which lead the cell into mitotic division Ø The loss of regulatory control of Ras activity is one of the steps which leads a cell towards cancer.

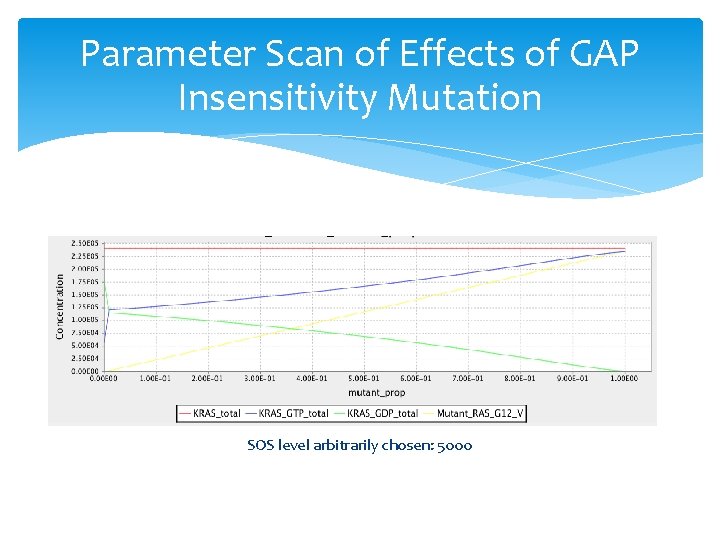
Parameter Scan of Effects of GAP Insensitivity Mutation SOS level arbitrarily chosen: 5000

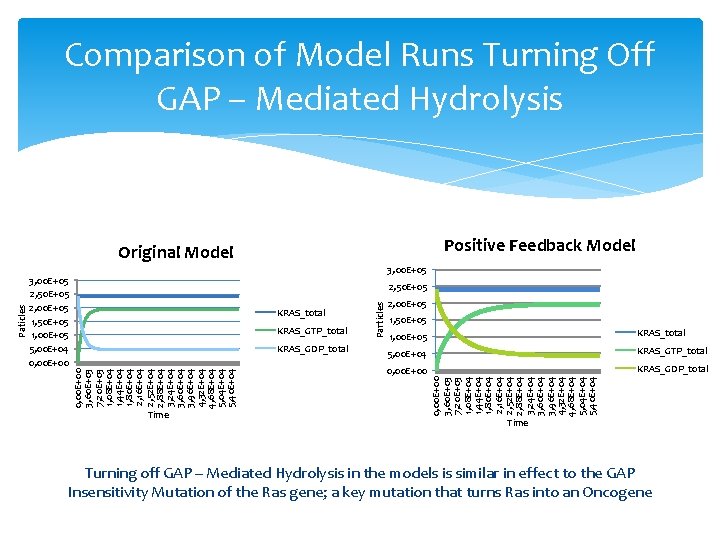
Comparison of Model Runs Turning Off GAP – Mediated Hydrolysis Positive Feedback Model 3, 00 E+05 2, 50 E+05 2, 00 E+05 1, 50 E+05 1, 00 E+05 5, 00 E+04 0, 00 E+00 KRAS_GTP_total KRAS_GDP_total Time 2, 00 E+05 1, 50 E+05 1, 00 E+05 KRAS_total 5, 00 E+04 KRAS_GTP_total 0, 00 E+00 KRAS_GDP_total 0, 00 E+00 3, 60 E+03 7, 20 E+03 1, 08 E+04 1, 44 E+04 1, 80 E+04 2, 16 E+04 2, 52 E+04 2, 88 E+04 3, 24 E+04 3, 60 E+04 3, 96 E+04 4, 32 E+04 4, 68 E+04 5, 04 E+04 5, 40 E+04 KRAS_total Particles 2, 50 E+05 0, 00 E+00 3, 60 E+03 7, 20 E+03 1, 08 E+04 1, 44 E+04 1, 80 E+04 2, 16 E+04 2, 52 E+04 2, 88 E+04 3, 24 E+04 3, 60 E+04 3, 96 E+04 4, 32 E+04 4, 68 E+04 5, 04 E+04 5, 40 E+04 Paticles Original Model Time Turning off GAP – Mediated Hydrolysis in the models is similar in effect to the GAP Insensitivity Mutation of the Ras gene; a key mutation that turns Ras into an Oncogene

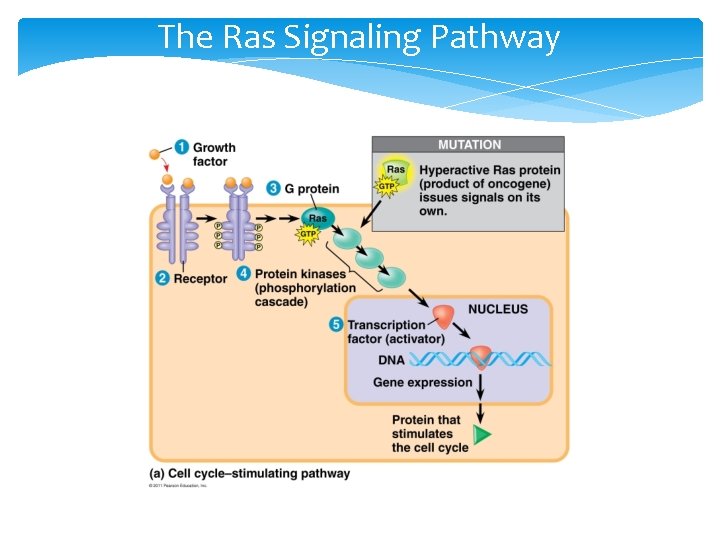
The Ras Signaling Pathway

Possible Bugs Ø Rate units and conversions Ø Model uses particles rather than concentrations Ø Rates taken from a variety of papers Ø Assumptions made Ø Concentration of SOS must be much lower than the concentration of Ras for Michaelis-Mentin equations to work Ø Unactivated SOS has no rate for converting Ras. GDP to Ras. GTP Ø SOS is equally likely to be activated by mutant Ras

Conclusions Ø Modeling specific components of a Signal Transduction Pathway provides insights into how a particular biological process works Ø A basic model of the process being investigated provides a foundation from which more detailed evaluation can be added onto, providing a more detailed view of the process Ø It is important to validate model results with experimental data to ensure that the model accurately reflects the modeled system Ø Different molecular interactions within the same pathway affect the efficiency of a molecules ability to catalyze downstream reactions

Conclusions Ø By not simulating the activation of the signal transduction pathway the model does not seem to accurately representing the signaling of a healthy cell Ø When Epithelial Growth Factor concentration decrease in the biological system, it is possible that the ligand would release from Receptor Tyrosine Kinase, deactivating it and shutting down the pathway Ø The cell can also endocytose the Receptor Tyrosine Kinase, turning off the signal transduction pathway Ø SOS inactivation/degradation will effect the concentrations of active Ras and thus the production of cyclins

Conclusions Ø While modeling Ras activation and degradation allows us to see the change in Ras concentration over time, the model does not take into account the activity of Tumor Suppressor molecules and their role in mediating the cell cycle Ø As The Hallmarks of Cancer paper by Hanahan and Weinberg shows, multiple errors need to occur within a cell in order for it to proceed towards cancer Ø While valuable in understanding one aspect of a cells journey down the path to cancer, modeling one aspect does not give the complete picture of the disease process within the cell Ø However, a reductionist approach can lead to an understanding of vulnerabilities within cancer that can be exploited by treatment options that target those particular areas

Suggested Extensions To Positive Feedback Model Ø Modeling the activation/deactivation of the Tyrosine Kinase Receptor and SOS recruitment to the plasma membrane would improve the overall accuracy of the model Ø Modeling SOS inactivation and degradation Ø Adding inhibitory molecules to the pathway to reflect the cell’s natural regulatory environment

Acknowledgements Group DAN would like to thank: Ø The National Science Foundation for funding the Workshop Ø Dr. Griffeth for providing direction, support, and leadership Ø Naralys Batista for great guidance with Rule Bender programming and keeping us well caffeinated and functioning Ø Dr. James Faeder Ø Dr. Stephen Redenti Ø CUNY – Lehman College for hosting the 2014 NSF-CMACS Workshop