2012 The Illumina Individual Genome Sequencing IGS service



























- Slides: 41




2012 Цена генома Геномное секвенирование The Illumina Individual Genome Sequencing (IGS) service Rapid. Track WGS - $9, 500 http: //www. everygenome. com/test_process/process. ilmn Экзомное секвенирование To date, BGI has sequenced more than 30, 000 human exomes. http: //bgiamericas. com/deep-exome-sequencing-promotion/ http: //dna. macrogen. com/eng/support/event_view. jsp? board_number=13509

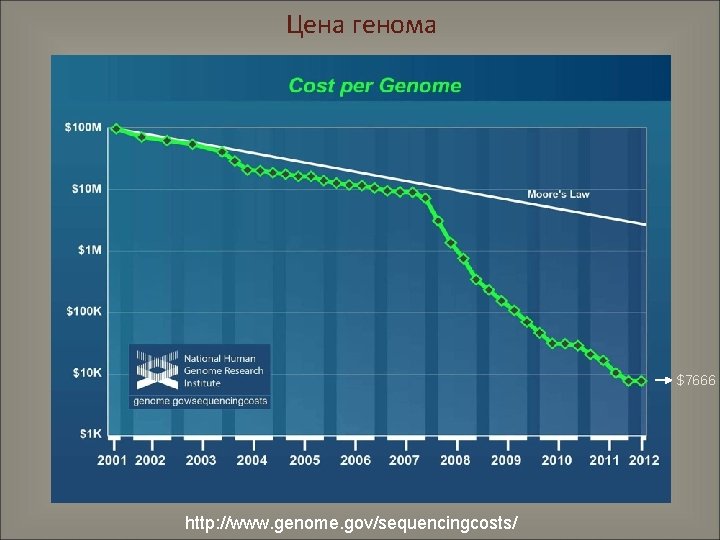
Цена генома $7666 http: //www. genome. gov/sequencingcosts/

Основные модели секвенаторов 2005 GS 20 2006 2007 2008 GS FLX GA 1 G 2009 2010 Titanium GS Junior GA II……x………e Hi. Seq 2000 SOLi. D 2. 0 SOLi. D 3 2011 FLX+ Hi. Seq 1000 Mi. Seq SOLi. D 4 5500 xl 5500 PGM Pac. Bio RS Max. Seq


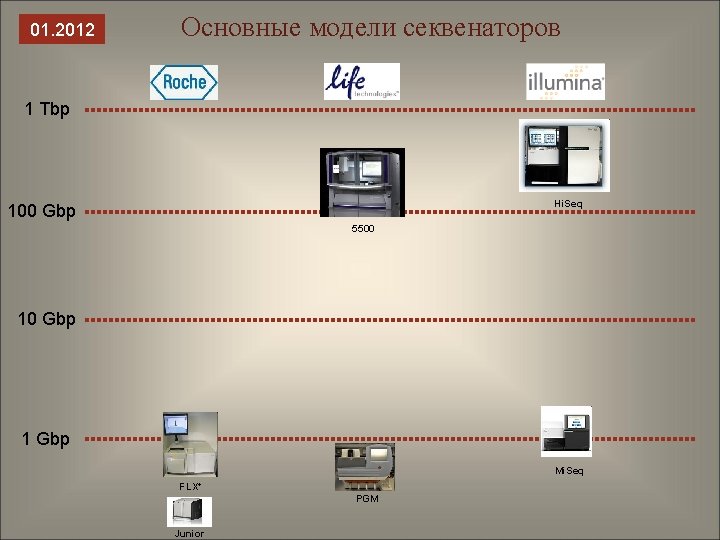
01. 2012 Основные модели секвенаторов 1 Tbp Hi. Seq 100 Gbp 5500 10 Gbp 1 Gbp Mi. Seq FLX+ PGM Junior

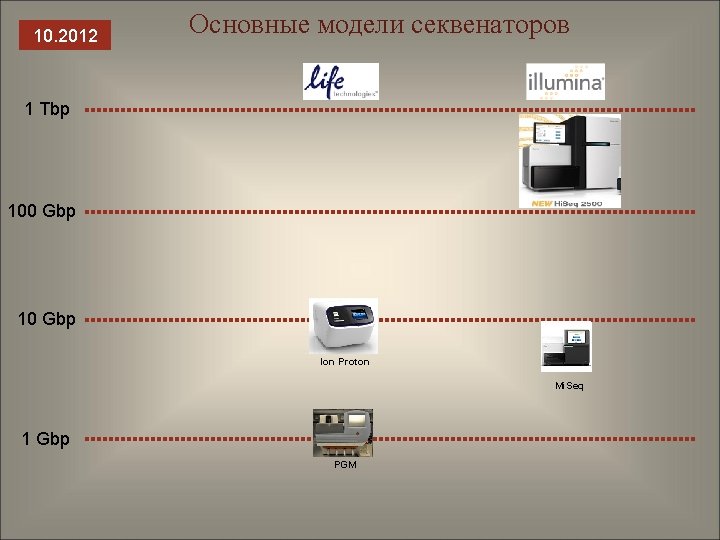
10. 2012 Основные модели секвенаторов 1 Tbp 100 Gbp 10 Gbp Ion Proton Mi. Seq 1 Gbp PGM



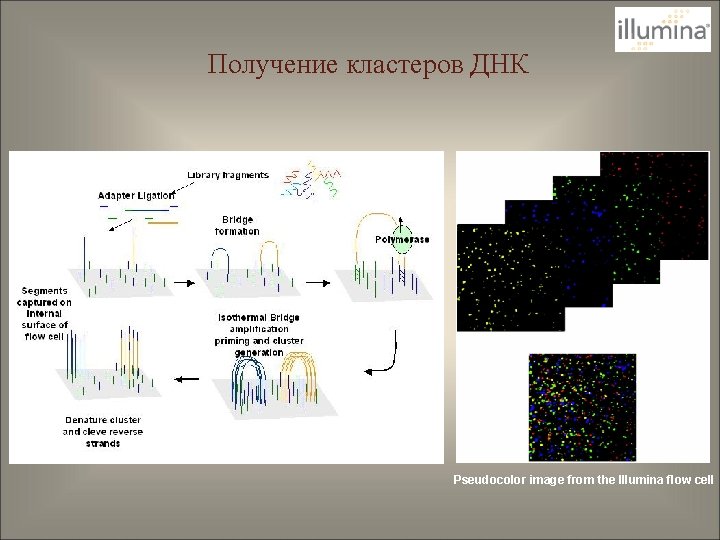
Получение кластеров ДНК Pseudocolor image from the Illumina flow cell

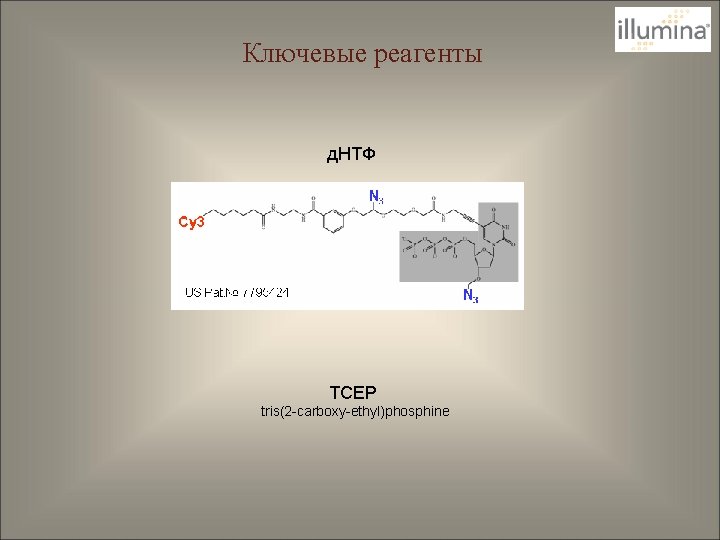
Ключевые реагенты д. НТФ TCEP tris(2 -carboxy-ethyl)phosphine




PGM (Personal Genome Machine)


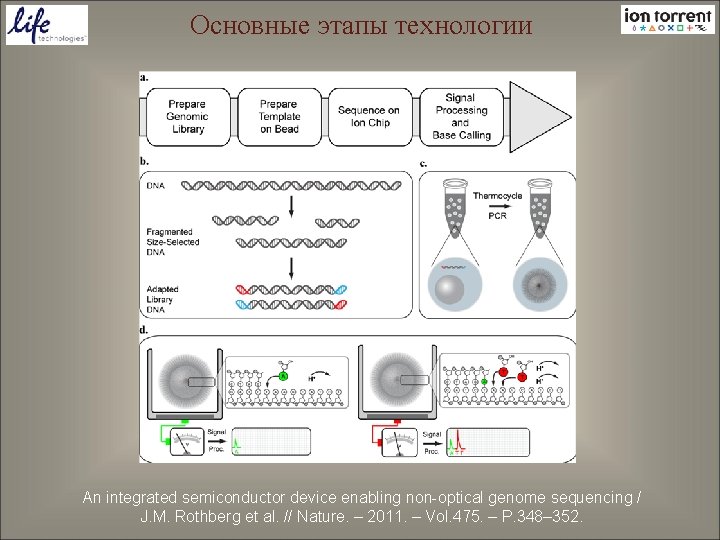
Основные этапы технологии An integrated semiconductor device enabling non-optical genome sequencing / J. M. Rothberg et al. // Nature. – 2011. – Vol. 475. – P. 348– 352.


Секвенаторы компании Ion Torrent $150 000 $49 500 PGM Throughput: Read Length: Run Time: Reads⁄Run: Weight: Up to 1 Gb per run Up to 300 bp 2 -4 hours 0, 6… 8 million reads 30 kg Ion Proton Up to 10 Gb per run Up to 200 bp 2 -4 hours 60 -80 million reads 59 kg

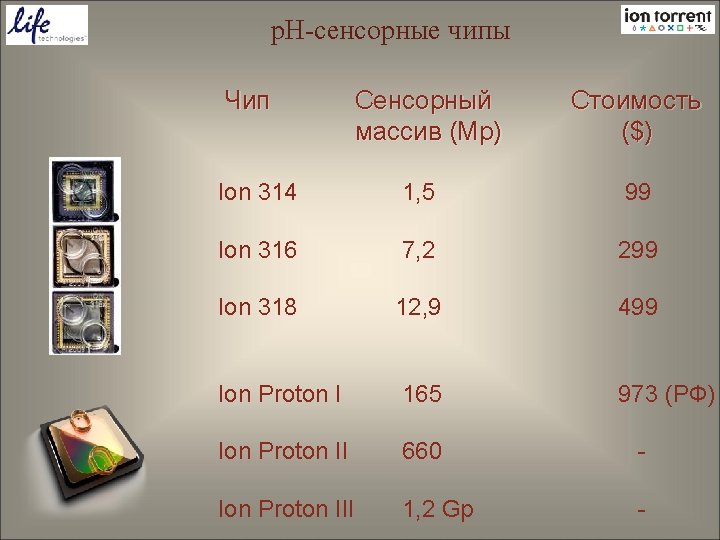
p. H-сенсорные чипы Чип Сенсорный массив (Mp) Стоимость ($) Ion 314 1, 5 99 Ion 316 7, 2 299 Ion 318 12, 9 499 Ion Proton I 165 973 (РФ) Ion Proton II 660 - Ion Proton III 1, 2 Gp -



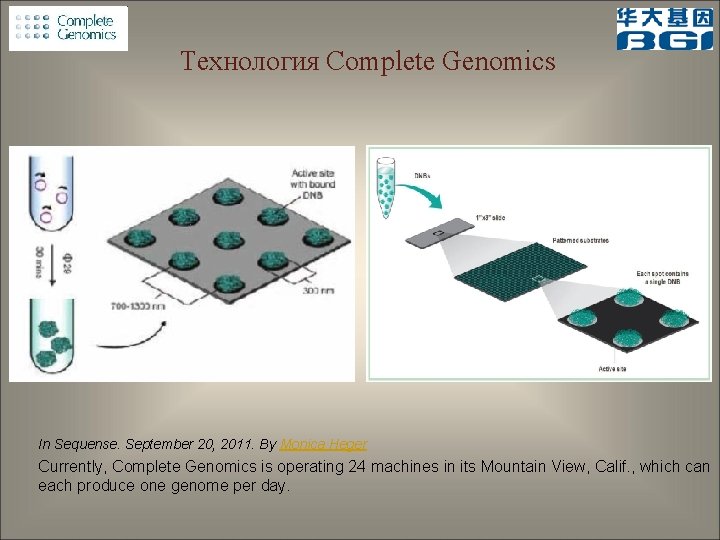
Технология Complete Genomics In Sequense. September 20, 2011. By Monica Heger Currently, Complete Genomics is operating 24 machines in its Mountain View, Calif. , which can each produce one genome per day.




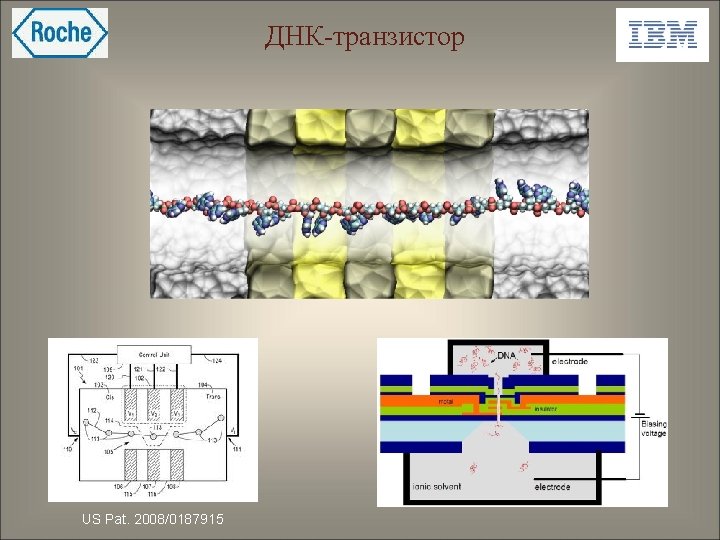
ДНК-транзистор US Pat. 2008/0187915

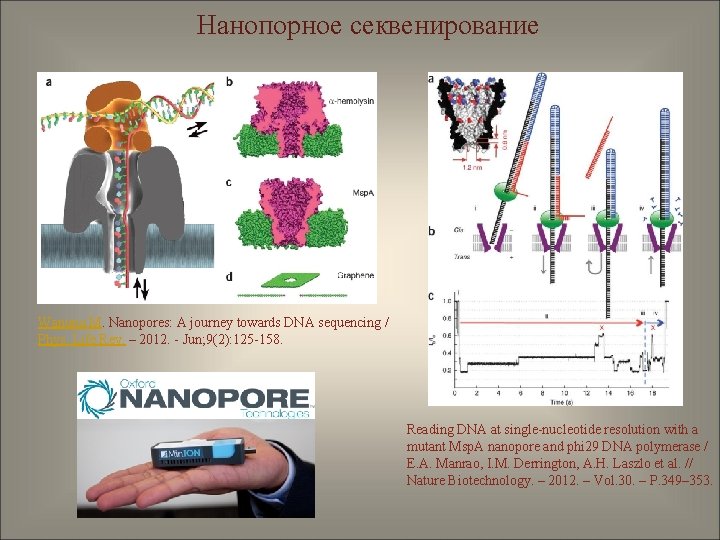
Нанопорное секвенирование Wanunu M. Nanopores: A journey towards DNA sequencing / Phys. Life Rev. – 2012. - Jun; 9(2): 125 -158. Reading DNA at single-nucleotide resolution with a mutant Msp. A nanopore and phi 29 DNA polymerase / E. A. Manrao, I. M. Derrington, A. H. Laszlo et al. // Nature Biotechnology. – 2012. – Vol. 30. – P. 349– 353.

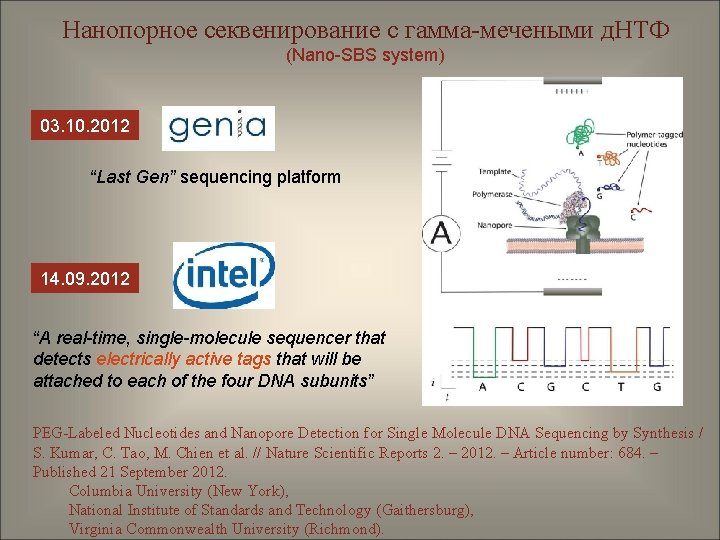
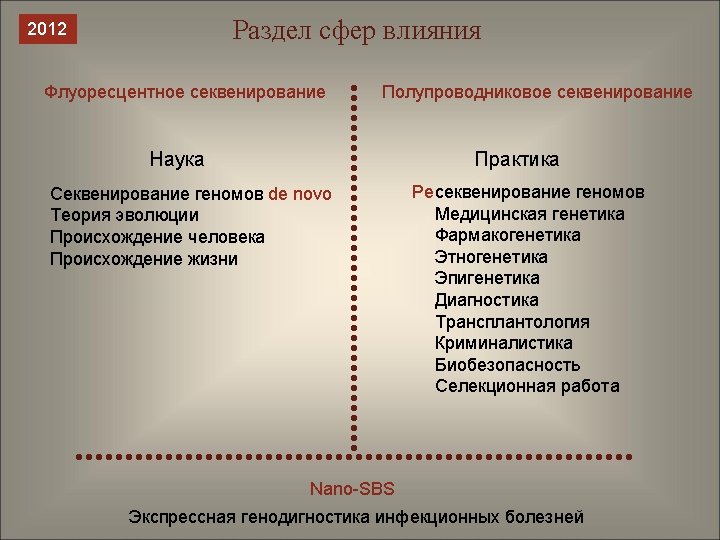
Нанопорное секвенирование с гамма-мечеными д. НТФ (Nano-SBS system) 03. 10. 2012 “Last Gen” sequencing platform 14. 09. 2012 “A real-time, single-molecule sequencer that detects electrically active tags that will be attached to each of the four DNA subunits” PEG-Labeled Nucleotides and Nanopore Detection for Single Molecule DNA Sequencing by Synthesis / S. Kumar, C. Tao, M. Chien et al. // Nature Scientific Reports 2. – 2012. – Article number: 684. – Published 21 September 2012. Columbia University (New York), National Institute of Standards and Technology (Gaithersburg), Virginia Commonwealth University (Richmond).



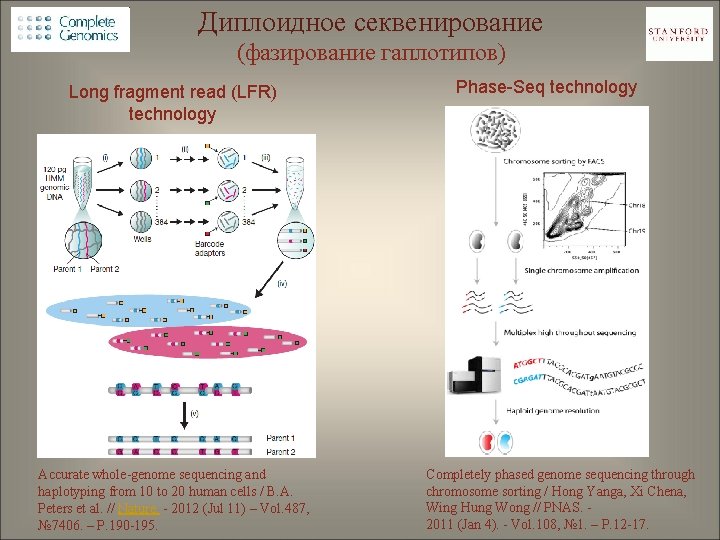
Диплоидное секвенирование (фазирование гаплотипов) Long fragment read (LFR) technology Accurate whole-genome sequencing and haplotyping from 10 to 20 human cells / B. A. Peters et al. // Nature. - 2012 (Jul 11) – Vol. 487, № 7406. – P. 190 -195. Phase-Seq technology Completely phased genome sequencing through chromosome sorting / Hong Yanga, Xi Chena, Wing Hung Wong // PNAS. 2011 (Jan 4). - Vol. 108, № 1. – P. 12 -17.

Cеквенирование гаплоидных клеток Stephen Quake, Ph. D, a professor of bioengineering and of applied physics at Stanford University Genome-wide Single-Cell Analysis of Recombination Activity and De Novo Mutation Rates in Human Sperm / J. Wang, H. Ch. Fan, B. Behr, S. R. Quake // Cell. – 2012 (July 20). – Vol. 150. – P. 402– 412.

Предсказательное секвенирование Extremely low-coverage sequencing and imputation increases power for genome-wide association studies / B. Pasaniuc, N. Rohland, P. L. Mc. Laren et al. // Nat. Genet. – 2012 (May 20). – Vol. 44, № 6. – P. 631 -635. “Here, we show that extremely low-coverage sequencing (0. 1– 0. 5×) captures almost as much of the common (>5%) and low-frequency (1– 5%) variation across the genome as SNP arrays. ” Предсказательные геномные секвенаторы? PGM Throughput: Run Time: Up to 1 Gb per run (0, 3 х) 2 -4 hours Ion Proton Up to 10 Gb per run (3 х) 2 -4 hours




