127 Molecular Flexibility Esther Kellenberger Facult de Pharmacie

1/27 Molecular Flexibility Esther Kellenberger Faculté de Pharmacie UMR 7200, Illkirch Tel: 03 68 85 42 21 e-mail: ekellen@unistra. fr
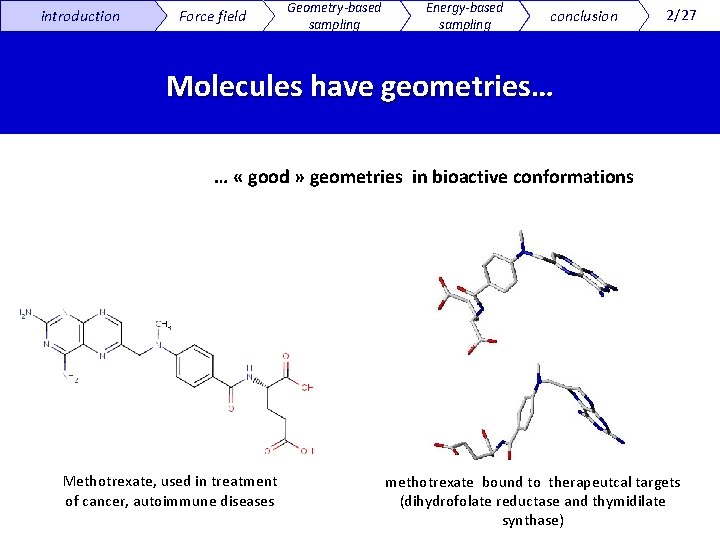
introduction Force field Geometry-based sampling Energy-based sampling conclusion 2/27 Molecules have geometries… … « good » geometries in bioactive conformations Methotrexate, used in treatment of cancer, autoimmune diseases methotrexate bound to therapeutcal targets (dihydrofolate reductase and thymidilate synthase)
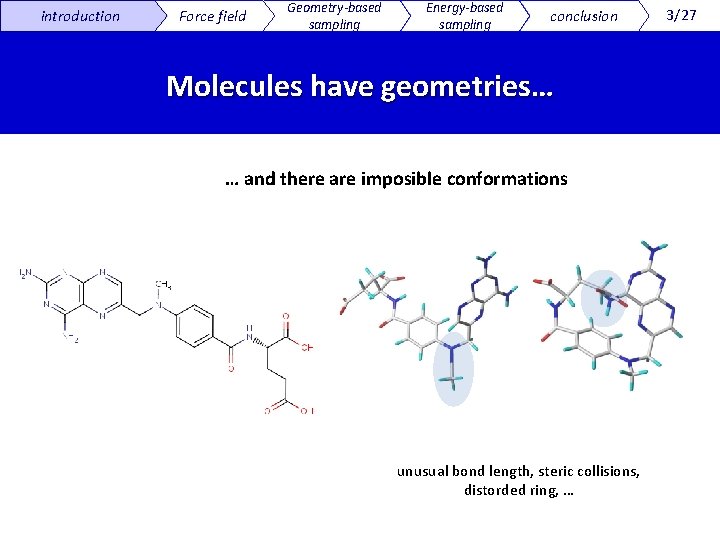
introduction Force field Geometry-based sampling Energy-based sampling conclusion Molecules have geometries… … and there are imposible conformations unusual bond length, steric collisions, distorded ring, … 3/27
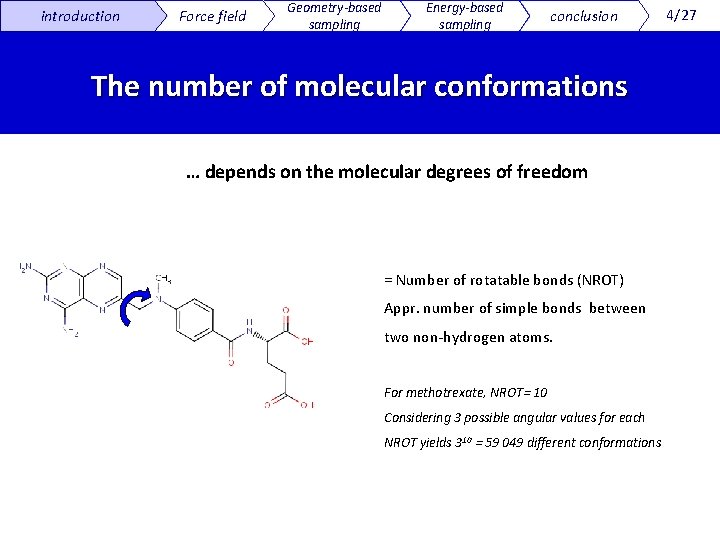
introduction Force field Geometry-based sampling Energy-based sampling conclusion The number of molecular conformations … depends on the molecular degrees of freedom = Number of rotatable bonds (NROT) Appr. number of simple bonds between two non-hydrogen atoms. For methotrexate, NROT= 10 Considering 3 possible angular values for each NROT yields 310 = 59 049 different conformations 4/27

introduction Force field Geometry-based sampling Energy-based sampling conclusion 5/27 How to evaluate the conformations? potential energy In physics, potential energy exists when a force acts stable (good) conformation low energy upon an object that tends to restore it to a lower energy configuration. Potential energy is the energy stored in a body or in a system due to its position in a force field or due to its configuration (SI unit= Joules, common unit = kcal/mol, 1 cal = 4. 1868 J) unstable (bad) conformation high energy A force field is a vector field that describes a noncontact force acting on a particle at various positions in space.
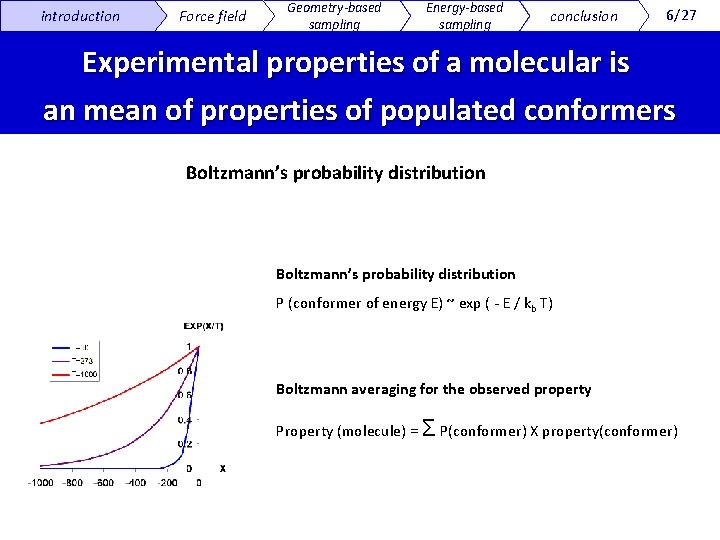
introduction Force field Geometry-based sampling Energy-based sampling conclusion 6/27 Experimental properties of a molecular is an mean of properties of populated conformers Boltzmann’s probability distribution P (conformer of energy E) ~ exp ( - E / kb T) Boltzmann averaging for the observed property Property (molecule) = Σ P(conformer) X property(conformer)

7/27 Chapter 1: Evaluation of the potential energy of conformers

introduction Force field Geometry-based sampling Energy-based sampling conclusion 8/27 Molecular mechanics Molecular systems are modeled using Newton’s laws: • each atom is simulated as a single particle • each particle is assigned a radius (van der Waals), polarizability, and a constant net charge • bonded interactions are treated as "springs" with an equilibrium distance equal to the bond length Molecular system's potential energy (E) in a given conformation as a sum of individual energy terms: E = E covalent + E non covalent
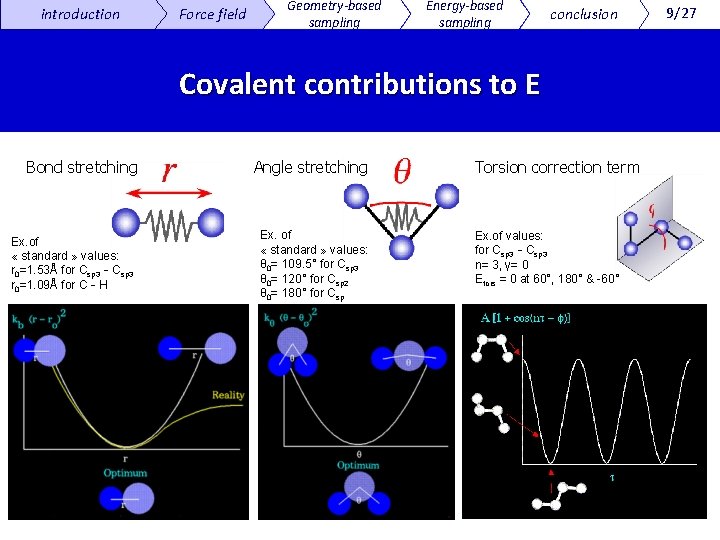
introduction Force field Geometry-based sampling Energy-based sampling conclusion Covalent contributions to E Bond stretching Ex. of « standard » values: r 0=1. 53Å for Csp 3‐Csp 3 r 0=1. 09Å for C‐H Angle stretching Ex. of « standard » values: θ 0= 109. 5° for Csp 3 θ 0= 120° for Csp 2 θ 0= 180° for Csp Torsion correction term Ex. of values: for Csp 3‐Csp 3 n= 3, γ= 0 Etors = 0 at 60°, 180° & -60° 9/27

introduction Force field Geometry-based sampling Energy-based sampling conclusion 10/27 Non covalent contributions to E Van der Waals term Electrostatic term Desolvation and Lennard Jones potential (6 -12) Coulomb’s law hydrophobic term EVd. W = A / rij 12 – B/rij 6 Ecoulomb = δ + δ - / 4πε 0 rij where A = 4 εσ12 B = 4 εσ6 where δ = charge ε = depth of the well ε 0 = solvent dielectric constant σ ~ distance with minimum EVd. W
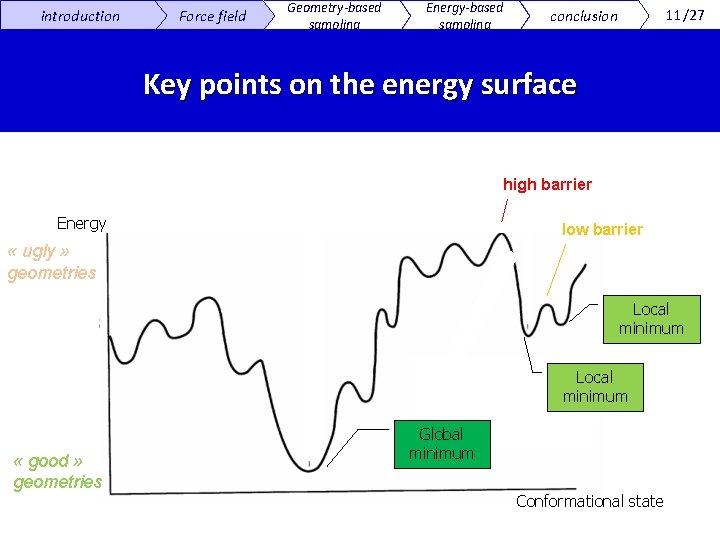
introduction Force field Geometry-based sampling Energy-based sampling 11/27 conclusion Key points on the energy surface high barrier Energy low barrier « ugly » geometries Local minimum « good » geometries Global minimum Conformational state
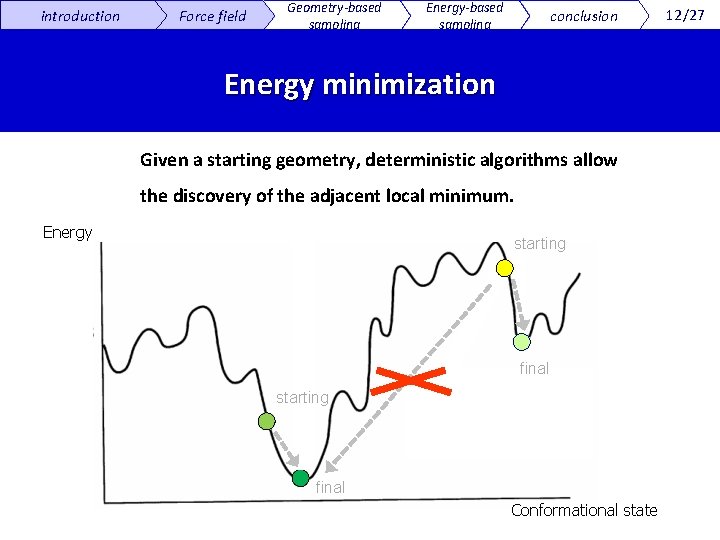
introduction Force field Geometry-based sampling Energy-based sampling conclusion Energy minimization Given a starting geometry, deterministic algorithms allow the discovery of the adjacent local minimum. Energy starting final Conformational state 12/27

introduction Force field Geometry-based sampling Energy-based sampling conclusion 13/27 The limits of conformational exploration by molecular dynamics Molecular dynamics trajectory may be seen as an exchange of potential and kinetic energy, with total energy being conserved. The dynamic system consists of moving particles (i. e. molecular atoms with coordinates and velocities). Particle position as a function of time is obtained by solving equation from the Newton’s laws. Energy sampling depends on the number of frames (time) starting Amplitude of motion controled by heating minimisation Conformational state

14/27 Chapter 2: exploration of the molecular energy landscape
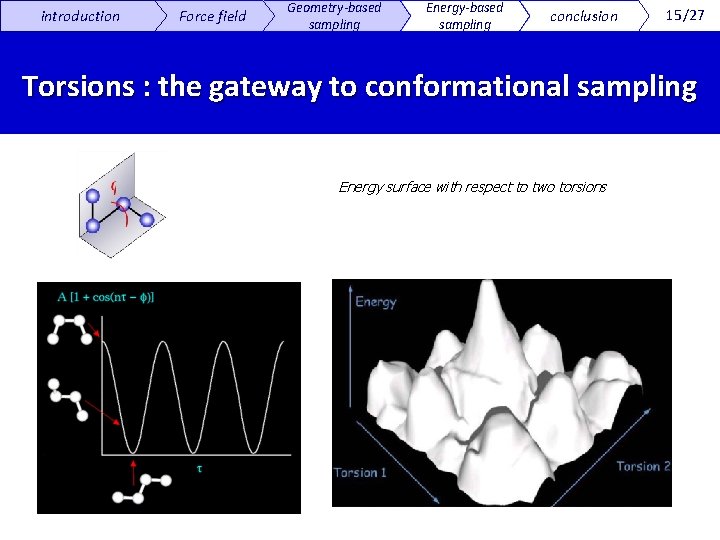
introduction Force field Geometry-based sampling Energy-based sampling conclusion 15/27 Torsions : the gateway to conformational sampling Energy surface with respect to two torsions
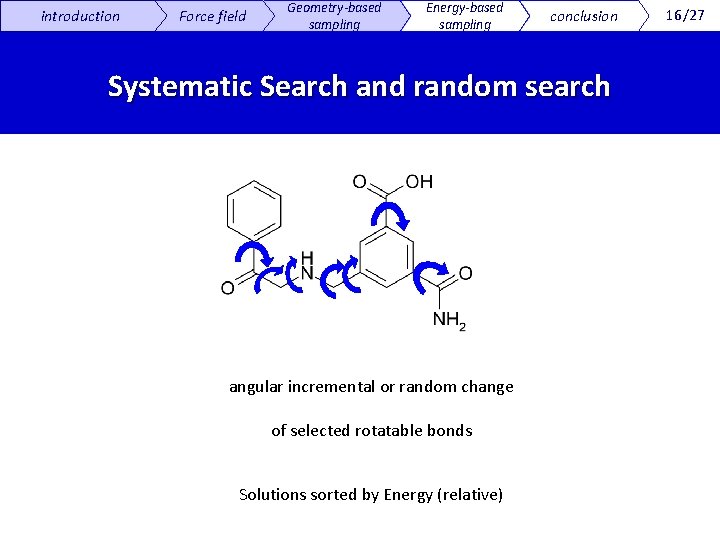
introduction Force field Geometry-based sampling Energy-based sampling conclusion Systematic Search and random search angular incremental or random change of selected rotatable bonds Solutions sorted by Energy (relative) 16/27
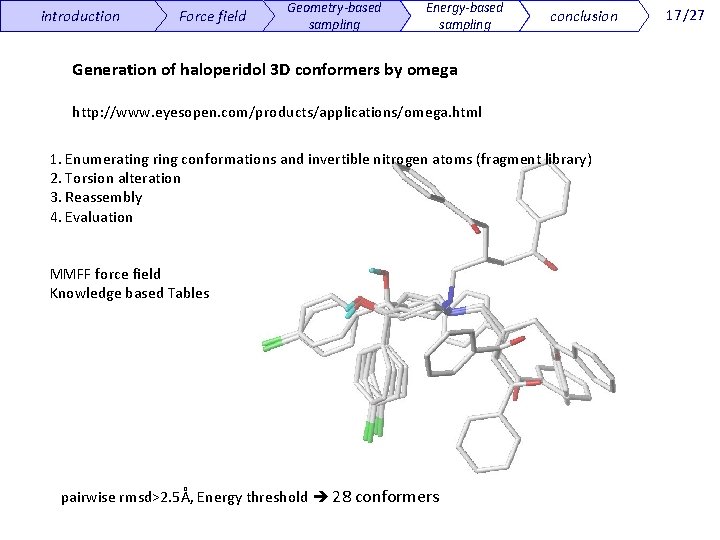
introduction Force field Geometry-based sampling Energy-based sampling conclusion Generation of haloperidol 3 D conformers by omega http: //www. eyesopen. com/products/applications/omega. html 1. Enumerating ring conformations and invertible nitrogen atoms (fragment library) 2. Torsion alteration 3. Reassembly 4. Evaluation MMFF force field Knowledge based Tables pairwise rmsd>2. 5Å, Energy threshold 28 conformers 17/27

introduction Force field Geometry-based sampling Energy-based sampling conclusion Increasing complexity of energy hypersurface … Geometry-based sampling methods: • a systematic search is possible if NROT < 4 -5 • Enumeration restricted to a fixed number of conformers for flexible compounds (Ex: 200 in omega) Energy-based sampling methods: • (molecular dynamics ) • stochastic sampling: Monte-Carlo and Genetic algorithm 18/27
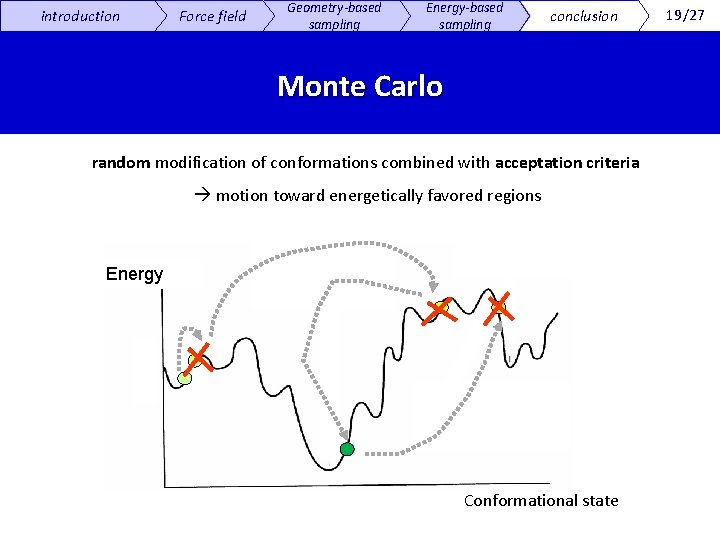
introduction Force field Geometry-based sampling Energy-based sampling conclusion Monte Carlo random modification of conformations combined with acceptation criteria motion toward energetically favored regions Energi Energy Conformational state 19/27
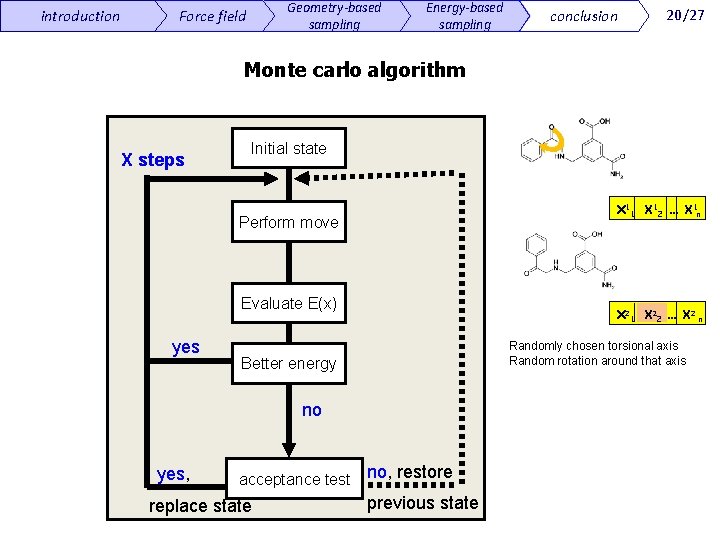
introduction Geometry-based sampling Force field Energy-based sampling conclusion 20/27 Monte carlo algorithm X steps Initial state Χ 11 Χ 12 … Χ 1 n Perform move Evaluate E(x) yes Χ 21 Χ 22 … Χ 2 n Randomly chosen torsional axis Random rotation around that axis Better energy no yes, acceptance test replace state no, restore previous state
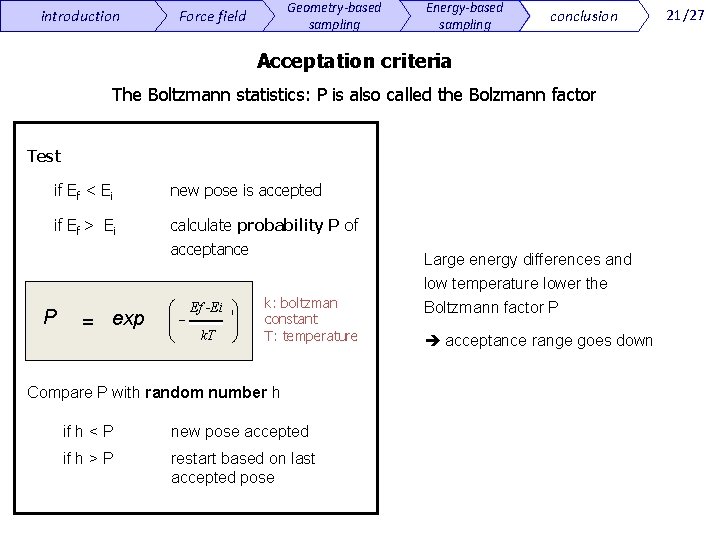
introduction Geometry-based sampling Force field Energy-based sampling conclusion Acceptation criteria The Boltzmann statistics: P is also called the Bolzmann factor Test if Ef < Ei new pose is accepted if Ef > Ei calculate probability P of acceptance exp = e æ ç ç è P æ Ef -Ei ç÷ ç ÷ k. T è k: boltzman constant T: temperature Compare P with random number h if h < P new pose accepted if h > P restart based on last accepted pose Large energy differences and low temperature lower the Boltzmann factor P acceptance range goes down 21/27
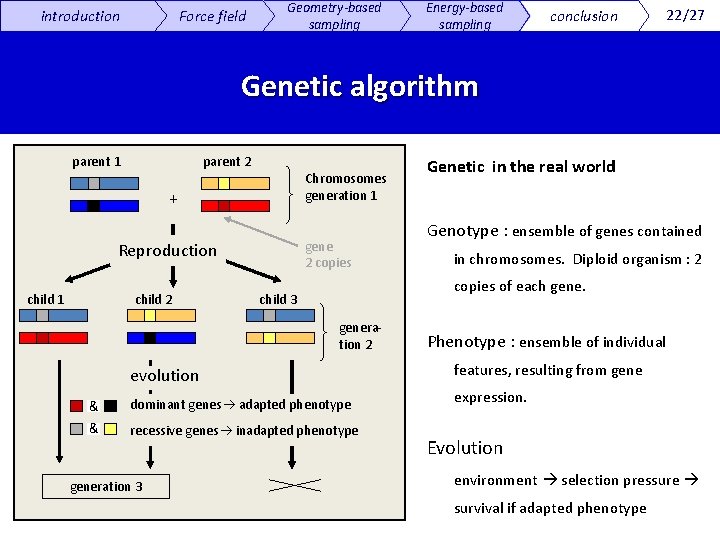
introduction Force field Geometry-based sampling Energy-based sampling conclusion 22/27 Genetic algorithm parent 1 parent 2 Chromosomes generation 1 + gene 2 copies Reproduction child 1 child 2 Genetic in the real world Genotype : ensemble of genes contained in chromosomes. Diploid organism : 2 copies of each gene. child 3 generation 2 Phenotype : ensemble of individual evolution features, resulting from gene & dominant genes adapted phenotype expression. & recessive genes inadapted phenotype generation 3 Evolution environment selection pressure survival if adapted phenotype
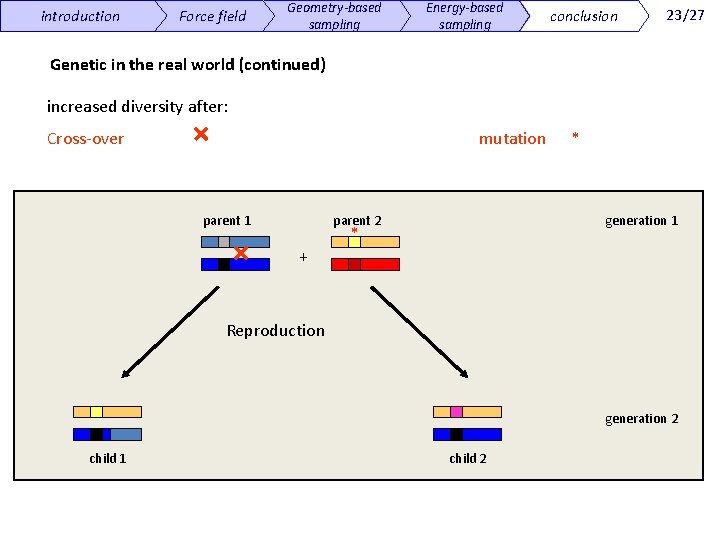
introduction Force field Geometry-based sampling Energy-based sampling conclusion 23/27 Genetic in the real world (continued) increased diversity after: Cross-over mutation * parent 1 parent 2 generation 1 * + Reproduction generation 2 child 1 child 2
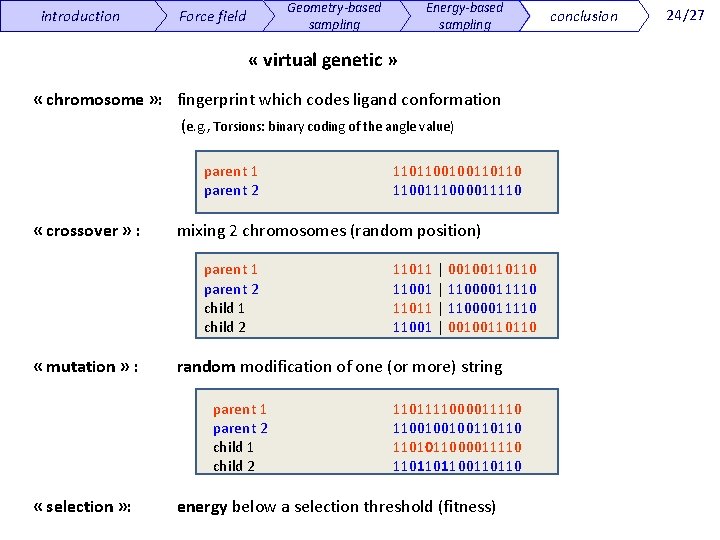
introduction Geometry-based sampling Force field Energy-based sampling « virtual genetic » « chromosome » : fingerprint which codes ligand conformation (e. g. , Torsions: binary coding of the angle value) parent 1 parent 2 « crossover » : mixing 2 chromosomes (random position) parent 1 parent 2 child 1 child 2 « mutation » : 11011 | 00100110110 11001 | 11000011110 11011 | 11000011110 11001 | 00100110110 random modification of one (or more) string parent 1 parent 2 child 1 child 2 « selection » : 1101100100110110 1100111000011110 1101111000011110 1100100100110110 1101011000011110 110110110 energy below a selection threshold (fitness) conclusion 24/27
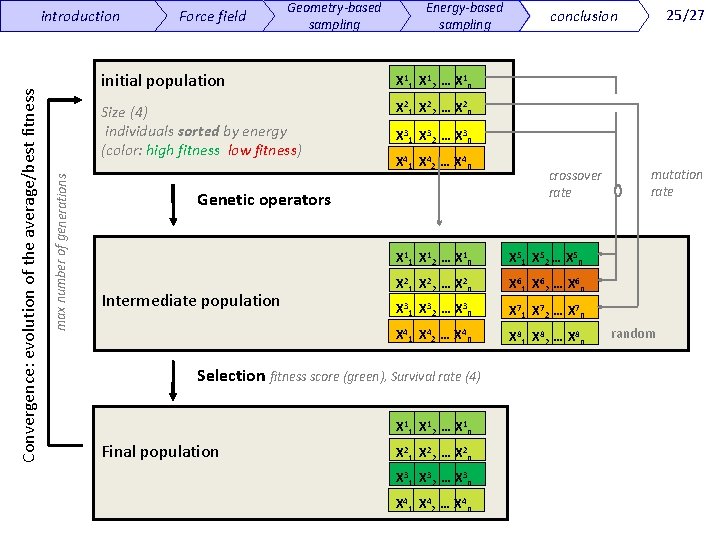
max number of generations Convergence: evolution of the average/best fitness introduction Force field Geometry-based sampling Energy-based sampling initial population Χ 11 Χ 12 … Χ 1 n Size (4) individuals sorted by energy (color: high fitness low fitness) Χ 21 Χ 22 … Χ 2 n Χ 31 Χ 32 … Χ 3 n Χ 41 Χ 42 … Χ 4 n Genetic operators Intermediate population crossover rate Χ 11 Χ 12 … Χ 1 n Χ 51 Χ 52 … Χ 5 n Χ 21 Χ 22 … Χ 2 n Χ 61 Χ 62 … Χ 6 n Χ 31 Χ 32 … Χ 3 n Χ 71 Χ 72 … Χ 7 n Χ 41 Χ 42 … Χ 4 n Χ 81 Χ 82 … Χ 8 n Selection fitness score (green), Survival rate (4) Χ 11 Χ 12 … Χ 1 n Final population 25/27 conclusion Χ 21 Χ 22 … Χ 2 n Χ 31 Χ 32 … Χ 3 n Χ 41 Χ 42 … Χ 4 n mutation rate random
introduction Force field Geometry-based sampling Energy-based sampling conclusion 26/27 Genetic algorithm is an optimization method: How to preserve the diversity? • Selection pressure: child chromosome replace the worst members of the population / bias in the selection of parent chromosomes (towards high fitness or favoring torsion values seen in in previous populations) • Multiple islands model: population split into sub-populations, with parallel simulations and occasionally swapping solutions (migration) • Discard of redundant chromosomes (requires a metric to evaluate the similarity of individuals) the niche model: a niche is a ensemble of similar individuals in a population (as estimated by RMSD). If there a more than niche size individuals in the niche, then the new individual is replaces the worst individual of the niche rather than the worse individual of the population, in order to preserve diversity within the population.

introduction Force field Geometry-based sampling Energy-based sampling conclusion 27/27 CONCLUSION • Conformational Sampling is the key element for understanding of molecular behavior • It may range from very simple to extremely difficult, to impossible • If you don’t do it well, better don’t do it at all: empirical methods based on molecular topology only may be more accurate than 3 D models based on wrong – or too few – conformations • Two main sources of errors: A. ) wrong calculated energy- geometry landscape (poor Force Field parameterization) and B. ) – insufficient sampling! Thanks to Dragos Howarth!
- Slides: 27