11205 RNA Structure Prediction 1212020 D Dobbs ISU

11/2/05 RNA Structure Prediction 12/1/2020 D Dobbs ISU - BCB 444/544 X: RNA Structure Prediction 1

Announcements Seminar 12: 10 PM Fri BCB Faculty Seminar in E 164 Lago How to do sequence alignments on parallel computers Srinivas Aluru, ECpr. E & Chair, BCB Program http: //www. bcb. iastate. edu/courses/BCB 691 F 2005. html 12/1/2020 D Dobbs ISU - BCB 444/544 X: RNA Structure Prediction 2

Announcements BCB 544 Projects - Important Dates: Nov 2 Wed noon - Project proposals due to David/Drena Nov 4 Fri 10 A - Approvals/responses to students Dec 2 Fri noon - Written project reports due Dec 5, 7, 8, 9 class/lab - Oral Presentations (20') (Dec 15 Thurs = Final Exam) 12/1/2020 D Dobbs ISU - BCB 444/544 X: RNA Structure Prediction 3
RNA Structure & Function Prediction Mon Review - promoter prediction RNA structure & function Wed RNA structure prediction 2' & 3' structure prediction mi. RNA & target prediction - perhaps. . RNA function prediction? Won't have time to cover this… 12/1/2020 D Dobbs ISU - BCB 444/544 X: RNA Structure Prediction 4

Reading Assignment (for Mon/Wed) Mount Bioinformatics • Chp 8 Prediction of RNA Secondary Structure • pp. 327 -355 • Ck Errata: http: //www. bioinformaticsonline. org/help/errata 2. html Cates (Online) RNA Secondary Structure Prediction Module • http: //cnx. rice. edu/content/m 11065/latest/ 12/1/2020 D Dobbs ISU - BCB 444/544 X: RNA Structure Prediction 5

Review last lecture: RNA Structure & Function 12/1/2020 D Dobbs ISU - BCB 444/544 X: RNA Structure Prediction 6

RNA Structure & Function • RNA structure • Levels of organization • Energetics (more about this on Wed) • RNA types & functions • • Genomic information storage/transfer Structural Catalytic Regulatory 12/1/2020 D Dobbs ISU - BCB 444/544 X: RNA Structure Prediction 7
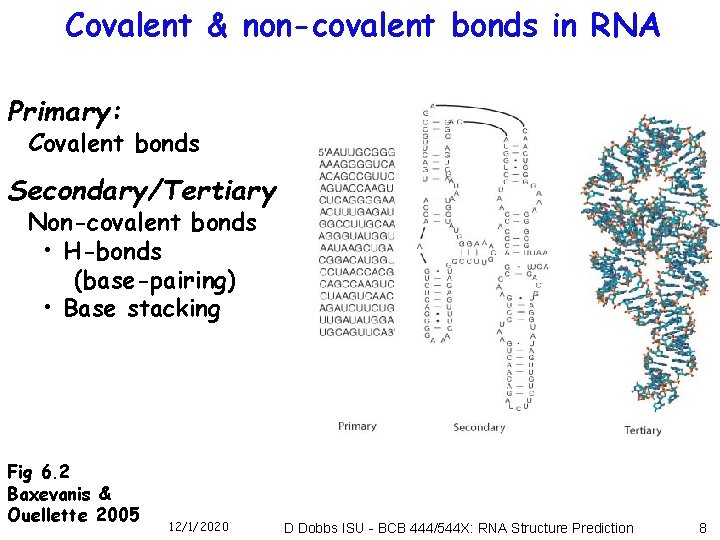
Covalent & non-covalent bonds in RNA Primary: Covalent bonds Secondary/Tertiary Non-covalent bonds • H-bonds (base-pairing) • Base stacking Fig 6. 2 Baxevanis & Ouellette 2005 12/1/2020 D Dobbs ISU - BCB 444/544 X: RNA Structure Prediction 8

Base-pairing in RNA 1) G-C, A-U, G-U ("wobble") & variants U can form base-pairs with both A & G 2) Nucleotides in RNA are frequently modified this is not very common in DNA These features & flexible "single-stranded" RNA backbone allow for many potential base-pairs Modified bases are especially important) in t. RNA: e. g. , pseudo-Uridine, r. D, 5 -CH 3 -C 6 -isopentenyl-A 7 -CH 3 -G, many others… See: IMB Image Library of Biological Molecules 12/1/2020 D Dobbs ISU - BCB 444/544 X: RNA Structure Prediction 9
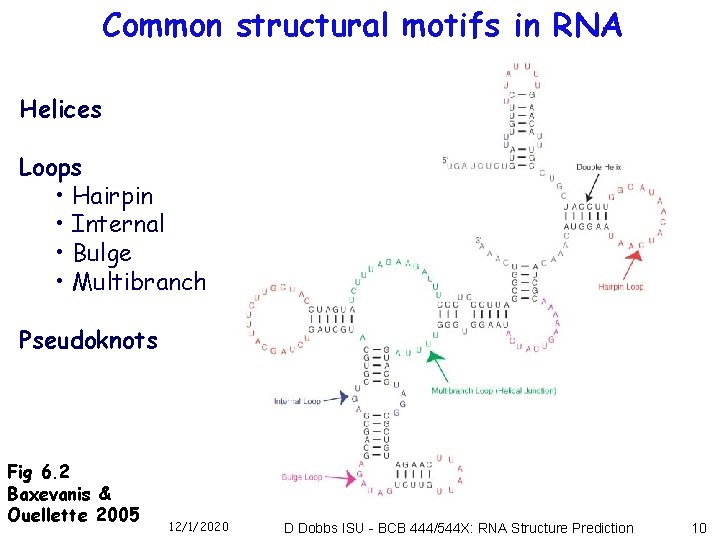
Common structural motifs in RNA Helices Loops • Hairpin • Internal • Bulge • Multibranch Pseudoknots Fig 6. 2 Baxevanis & Ouellette 2005 12/1/2020 D Dobbs ISU - BCB 444/544 X: RNA Structure Prediction 10

RNA functions Storage/transfer of genetic information • Genomes • many viruses have RNA genomes single-stranded (ss. RNA) e. g. , retroviruses (HIV) double-stranded (ds. RNA) • Transfer of genetic information • m. RNA = "coding RNA" - encodes proteins 12/1/2020 D Dobbs ISU - BCB 444/544 X: RNA Structure Prediction 11

RNA functions Structural • e. g. , r. RNA, which is major structural component of ribosomes (Gloria Culver, ISU) BUT - its role is not just structural, also: Catalytic RNA in ribosome has peptidyltransferase activity • Enzymatic activity responsible for peptide bond formation between amino acids in growing peptide chain • Also, many small RNAs are enzymes "ribozymes" (W Allen Miller, ISU) 12/1/2020 D Dobbs ISU - BCB 444/544 X: RNA Structure Prediction 12

RNA functions Regulatory Recently discovered important new roles for RNAs In normal cells: • in "defense" - esp. in plants • in normal development e. g. , si. RNAs, mi. RNA As tools: • for gene therapy or to modify gene expression • RNAi (used by many at ISU: Diane Bassham, • Thomas Baum, Jeff Essner, Kristen Johansen, Jo Anne Powell-Coffman, Roger Wise, etc. ) RNA aptamers (Marit Nilsen-Hamilton, ISU) 12/1/2020 D Dobbs ISU - BCB 444/544 X: RNA Structure Prediction 13
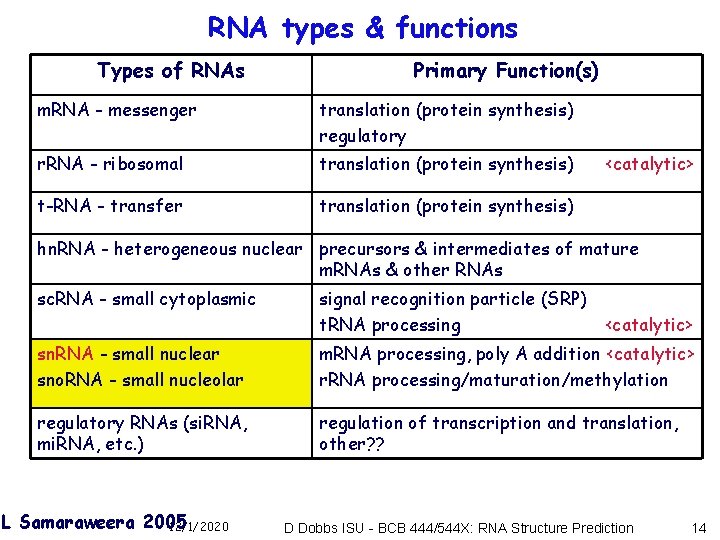
RNA types & functions Types of RNAs Primary Function(s) m. RNA - messenger translation (protein synthesis) regulatory r. RNA - ribosomal translation (protein synthesis) t-RNA - transfer translation (protein synthesis) <catalytic> hn. RNA - heterogeneous nuclear precursors & intermediates of mature m. RNAs & other RNAs sc. RNA - small cytoplasmic signal recognition particle (SRP) t. RNA processing <catalytic> sn. RNA - small nuclear sno. RNA - small nucleolar m. RNA processing, poly A addition <catalytic> r. RNA processing/maturation/methylation regulatory RNAs (si. RNA, mi. RNA, etc. ) regulation of transcription and translation, other? ? L Samaraweera 2005 12/1/2020 D Dobbs ISU - BCB 444/544 X: RNA Structure Prediction 14

Thanks to Chris Burge, MIT for following slides Slightly modified from: Gene Regulation and Micro. RNAs Session introduction presented at ISMB 2005, Detroit, MI Chris Burge cburge@MIT. EDU C Burge 2005 12/1/2020 D Dobbs ISU - BCB 444/544 X: RNA Structure Prediction 15
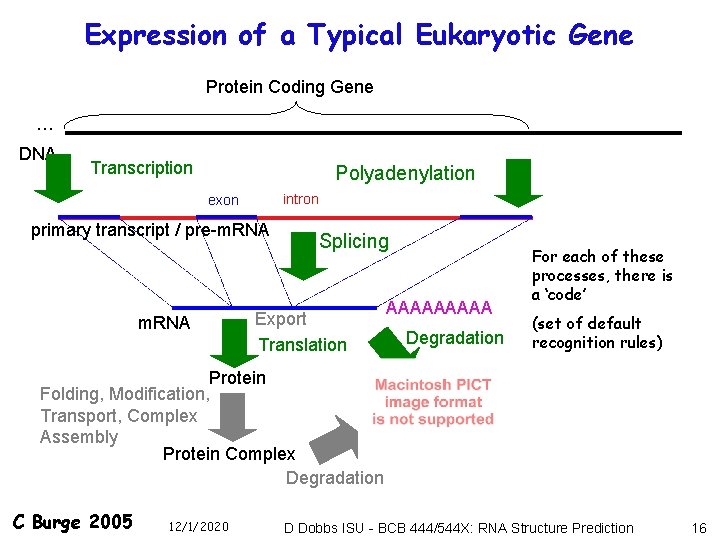
Expression of a Typical Eukaryotic Gene Protein Coding Gene … DNA Transcription Polyadenylation intron exon primary transcript / pre-m. RNA Splicing Export Translation AAAAA Degradation For each of these processes, there is a ‘code’ (set of default recognition rules) Protein Folding, Modification, Transport, Complex Assembly Protein Complex Degradation C Burge 2005 12/1/2020 D Dobbs ISU - BCB 444/544 X: RNA Structure Prediction 16

Gene Expression Challenges for Computational Biology • Understand the ‘code’ for each step in gene expression (set of default recognition rules), e. g. , the ‘splicing code’ • Understand the rules for sequence-specific recognition of nucleic acids by protein and ribonucleoprotein (RNP) factors • Understand the regulatory events that occur at each step and the biological consequences of regulation Lots of data Genomes, structures, transcripts, microarrays, Ch. IP-Chip, etc. C Burge 2005 12/1/2020 D Dobbs ISU - BCB 444/544 X: RNA Structure Prediction 17

Sequence-specific Transcription Factors • have modular organization » Understand DNA-binding specificity Yan (ISU) A computational method to identify amino acid residues involved in protein-DNA interactions ATF-2/c-Jun/IRF-3 DNA complex Panne et al. EMBO J. 2004 C Burge 2005 12/1/2020 D Dobbs ISU - BCB 444/544 X: RNA Structure Prediction 18
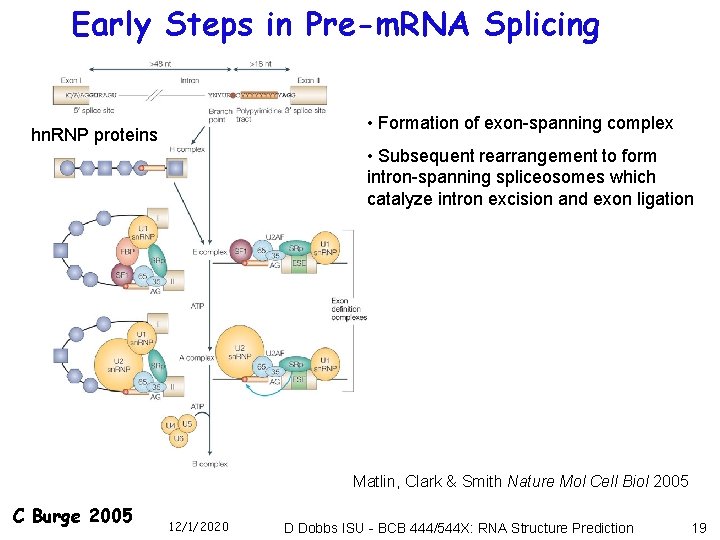
Early Steps in Pre-m. RNA Splicing • Formation of exon-spanning complex hn. RNP proteins • Subsequent rearrangement to form intron-spanning spliceosomes which catalyze intron excision and exon ligation Matlin, Clark & Smith Nature Mol Cell Biol 2005 C Burge 2005 12/1/2020 D Dobbs ISU - BCB 444/544 X: RNA Structure Prediction 19
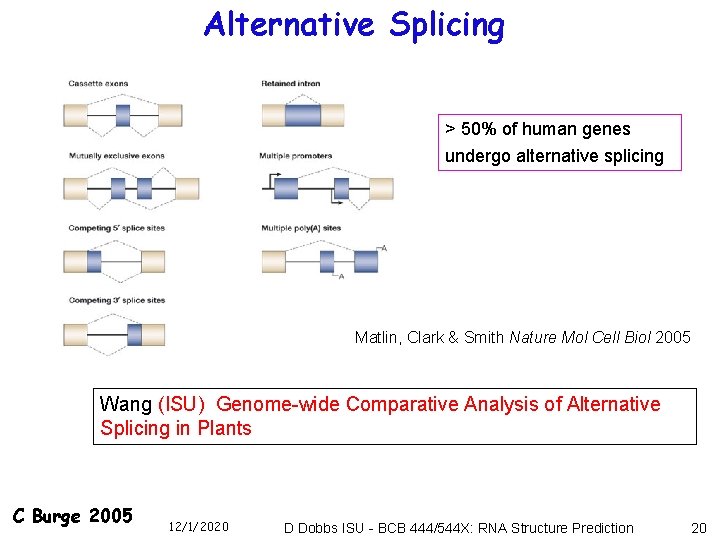
Alternative Splicing > 50% of human genes undergo alternative splicing Matlin, Clark & Smith Nature Mol Cell Biol 2005 Wang (ISU) Genome-wide Comparative Analysis of Alternative Splicing in Plants C Burge 2005 12/1/2020 D Dobbs ISU - BCB 444/544 X: RNA Structure Prediction 20

Splicing Regulation ESE/ESS = Exonic Splicing Enhancers/Silencers ISE/ISS = Intronic Splicing Enhancers/Silencers Matlin, Clark & Smith Nature Mol Cell Biol 2005 C Burge 2005 12/1/2020 D Dobbs ISU - BCB 444/544 X: RNA Structure Prediction 21
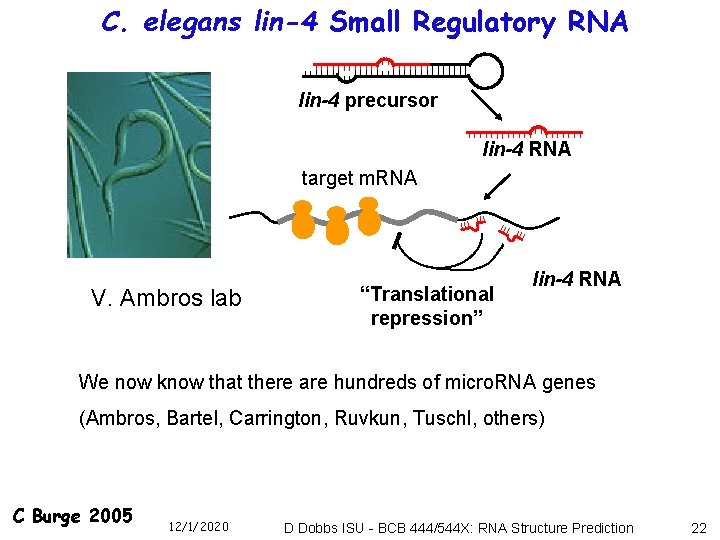
C. elegans lin-4 Small Regulatory RNA lin-4 precursor lin-4 RNA target m. RNA V. Ambros lab “Translational repression” lin-4 RNA We now know that there are hundreds of micro. RNA genes (Ambros, Bartel, Carrington, Ruvkun, Tuschl, others) C Burge 2005 12/1/2020 D Dobbs ISU - BCB 444/544 X: RNA Structure Prediction 22
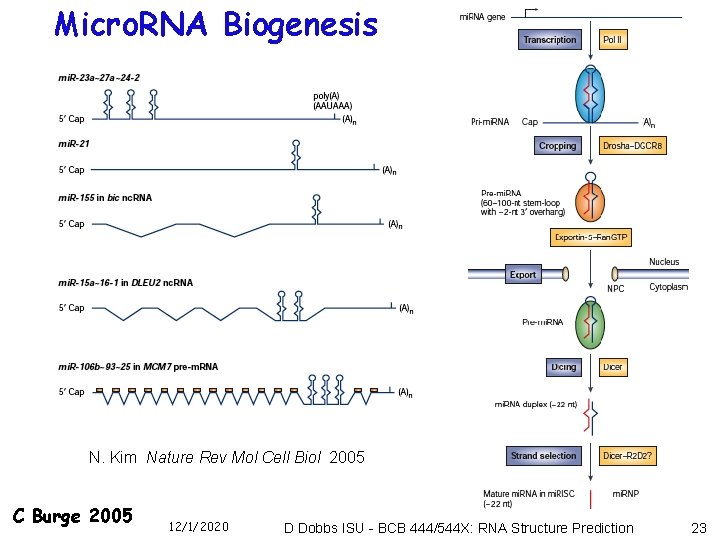
Micro. RNA Biogenesis N. Kim Nature Rev Mol Cell Biol 2005 C Burge 2005 12/1/2020 D Dobbs ISU - BCB 444/544 X: RNA Structure Prediction 23
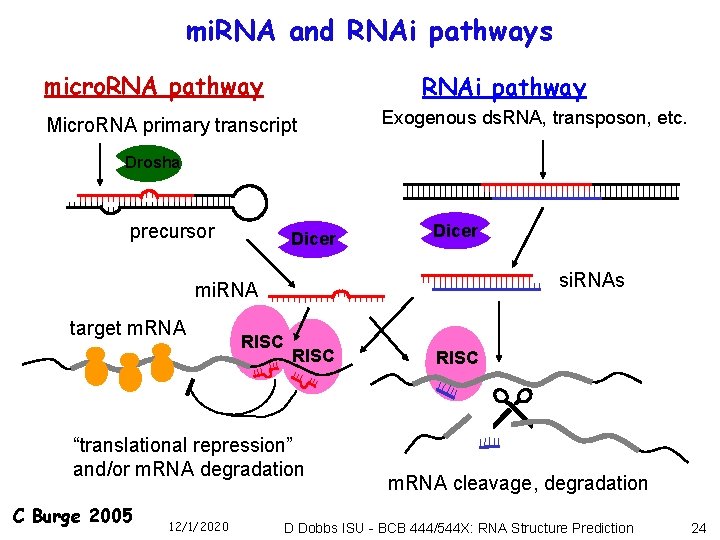
mi. RNA and RNAi pathways micro. RNA pathway RNAi pathway Micro. RNA primary transcript Exogenous ds. RNA, transposon, etc. Drosha precursor Dicer si. RNAs mi. RNA target m. RNA RISC “translational repression” and/or m. RNA degradation C Burge 2005 12/1/2020 RISC m. RNA cleavage, degradation D Dobbs ISU - BCB 444/544 X: RNA Structure Prediction 24

mi. RNA Challenges for Computational Biology • Find the genes encoding micro. RNAs • Predict their regulatory targets Computational Prediction of Micro. RNA Genes & Targets • Integrate mi. RNAs into gene regulatory pathways & networks Need to modify traditional paradigm of "transcriptional control" primarily by protein-DNA interactions to include mi. RNA regulatory mechanisms! C Burge 2005 12/1/2020 D Dobbs ISU - BCB 444/544 X: RNA Structure Prediction 25

New Today: RNA Structure Prediction 12/1/2020 D Dobbs ISU - BCB 444/544 X: RNA Structure Prediction 26
RNA structure prediction strategies Secondary structure prediction 1) Energy minimization (thermodynamics) 2) Comparative sequence analysis (co-variation) 3) Combined experimental & computational 12/1/2020 D Dobbs ISU - BCB 444/544 X: RNA Structure Prediction 27

Secondary structure prediction strategies 1) Energy minimization (thermodynamics) • Algorithm: Dynamic programming to find high probability pairs (also, some Genetic algorithms) • Software: Mfold - Zuker Vienna RNA Package - Hofacker RNAstructure - Mathews Sfold - Ding & Lawrence R Knight 2005 12/1/2020 D Dobbs ISU - BCB 444/544 X: RNA Structure Prediction 28

Secondary structure prediction strategies 2) Comparative sequence analysis (co-variation) • Algorithm: Mutual information Context-free grammars • Software: Con. Struct Alifold Pfold FOLDALIGN Dynalign R Knight 2005 12/1/2020 D Dobbs ISU - BCB 444/544 X: RNA Structure Prediction 29

Secondary structure prediction strategies 3) Combined experimental & computational • Experiment: Map single-stranded vs double-stranded regions in folded RNA • How? Enzymes: S 1 nuclease, T 1 RNase Chemicals: kethoxal, DMS R Knight 2005 12/1/2020 D Dobbs ISU - BCB 444/544 X: RNA Structure Prediction 30

Experimental RNA structure determination? • X-ray crystallography • NMR spectroscopy • Enzymatic/chemical mapping 12/1/2020 D Dobbs ISU - BCB 444/544 X: RNA Structure Prediction 31

1) Energy minimization method What are the assumptions? Native tertiary structure or "fold" of an RNA molecule is (one of) its "lowest" free energy configuration(s) Gibbs free energy = G in kcal/mol at 37 C = equilibrium stability of structure lower values (negative) are more favorable Is this assumption valid? in vivo? - this may not hold, but we don't really know 12/1/2020 D Dobbs ISU - BCB 444/544 X: RNA Structure Prediction 32

Free energy minimization What are the rules? A A A U C Staben 2005 U U Basepair A=U What gives here? G = -1. 2 kcal/mole U A Basepair A=U U=A G = -1. 6 kcal/mole 12/1/2020 D Dobbs ISU - BCB 444/544 X: RNA Structure Prediction 33

Energy minimization calculations: Base-stacking is critical - Tinocco et al. C Staben 2005 12/1/2020 D Dobbs ISU - BCB 444/544 X: RNA Structure Prediction 34

Nearest-neighbor parameters Most methods for free energy minimization use nearest-neighbor parameters (derived from experiment) for predicting stability of an RNA secondary structure (in terms of G at 37 C) & most available software packages use the same set of parameters: Mathews, Sabina, Zuker & Turner, 1999 12/1/2020 D Dobbs ISU - BCB 444/544 X: RNA Structure Prediction 35

Energy minimization - calculations: Total free energy of a specific conformation for a specific RNA molecule = sum of incremental energy terms for: • helical stacking (sequence dependent) • loop initiation • unpaired stacking (favorable "increments" are < 0) Fig 6. 3 Baxevanis & Ouellette 2005 12/1/2020 D Dobbs ISU - BCB 444/544 X: RNA Structure Prediction 36

But how many possible conformations for a single RNA molecule? Huge number: Zuker estimates (1. 8)N possible secondary structures for a sequence of N nucleotides for 100 nts (small RNA…) = 3 X 1025 structures! Solution? Not exhaustive enumeration… Ø Dynamic programming O(N 3) in time O(N 2) in space/storage iff pseudoknots excluded, otherwise: O(N 6 ), time O(N 4 ), space 12/1/2020 D Dobbs ISU - BCB 444/544 X: RNA Structure Prediction 37

2) Comparative sequence analysis (co-variation) Two basic approaches: • Algorithms constrained by initial alignment Much faster, but not as robust as unconstrained Base-pairing probabilities determined by a partition function • Algorithms not constrained by initial alignment Genetic algorithms often used for finding an alignment & set of structures 12/1/2020 D Dobbs ISU - BCB 444/544 X: RNA Structure Prediction 38

RNA Secondary structure prediction: Performance? How evaluate? • Not many experimentally determined structures currently, ~ 50% are r. RNA structures so "Gold Standard" (in absence of tertiary structure): compare with predicted RNA secondary structure with that determined by comparative sequence analysis (!!? ? ) using Benchmark Datasets NOTE: Base-pairs predicted by comparative sequence analysis for large & small subunit r. RNAs are 97% accurate when compared with high resolution crystal structures! - Gutell, Pace 12/1/2020 D Dobbs ISU - BCB 444/544 X: RNA Structure Prediction 39

RNA Secondary structure prediction: Performance? 1) Energy minimization (via dynamic programming) 73% avg. prediction accuracy - single sequence 2) Comparative sequence analysis 97% avg. prediction accuracy - multiple sequences (e. g. , highly conserved r. RNAs) much lower if sequence conservation is lower &/or fewer sequences are available for alignment 1) 3) Combined - recent developments: 2) combine thermodynamics & co-variation 3) & experimental constraints? IMPROVED RESULTS 12/1/2020 D Dobbs ISU - BCB 444/544 X: RNA Structure Prediction 40
RNA structure prediction strategies Tertiary structure prediction Requires "craft" & significant user input & insight 1) Extensive comparative sequence analysis to predict tertiary contacts (co-variation) e. g. , MANIP - Westhof 2) Use experimental data to constrain model building e. g. , MC-CYM - Major • Homology modeling using sequence alignment & reference tertiary structure (not many of these!) 1) Low resolution molecular mechanics e. g. , yammp - Harvey 12/1/2020 D Dobbs ISU - BCB 444/544 X: RNA Structure Prediction 41
- Slides: 41