1 SELFDIFFERENTIATED BACTERIAL ASSEMBLY LINE PEKING 2007 IGEM

1 SELF-DIFFERENTIATED BACTERIAL ASSEMBLY LINE PEKING 2007 IGEM All ideas and images from Peking 2007 IGEM team unless otherwise noted Allen Lin

Issue 2 Proteins whose production interferes with each other Need for seperation Seperation from homogenous conditions in division of labor Solutions: Temporal Differentiation Spatial Differentiation

Push-on-Push-off Switch 3 Temporal Seperation Same input at different times -> different output Simple finite state machine (has a current state and cyclic) – binary
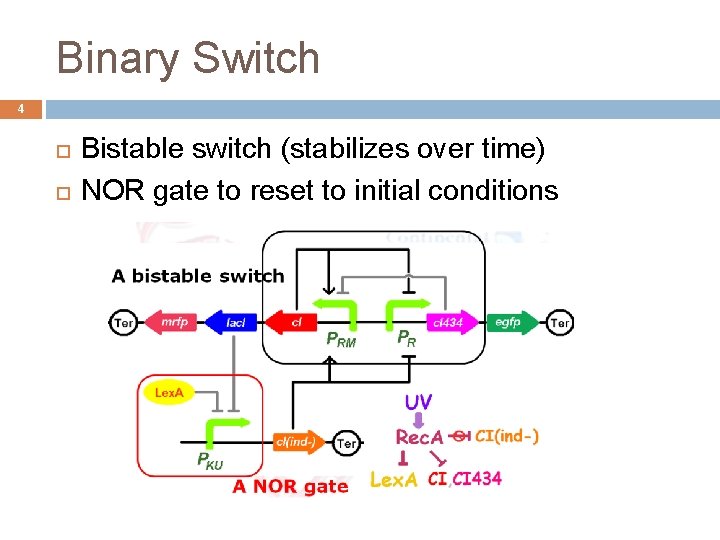
Binary Switch 4 Bistable switch (stabilizes over time) NOR gate to reset to initial conditions

Binary Switch 5

Details of Binary Switch 6 At first, Prm on -> CI and lac. I produced, red color C 1 represses Pr, lac. I represses Pku After UV light, CI degraded; Pr stronger promoter -> CI 434 produced, green color Pr represses Prm, lac. I production falls (m. RNA degraded), so only Lex. A represss Pku After 2 nd UV light, Lex. A degraded, Pku on -> Prm on, red color, back to start c. I(ind-) repressions Pr
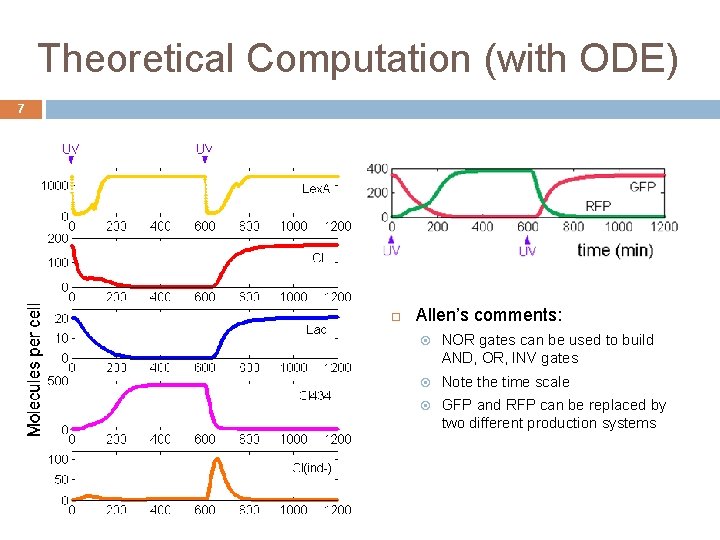
Theoretical Computation (with ODE) 7 Allen’s comments: NOR gates can be used to build AND, OR, INV gates Note the time scale GFP and RFP can be replaced by two different production systems
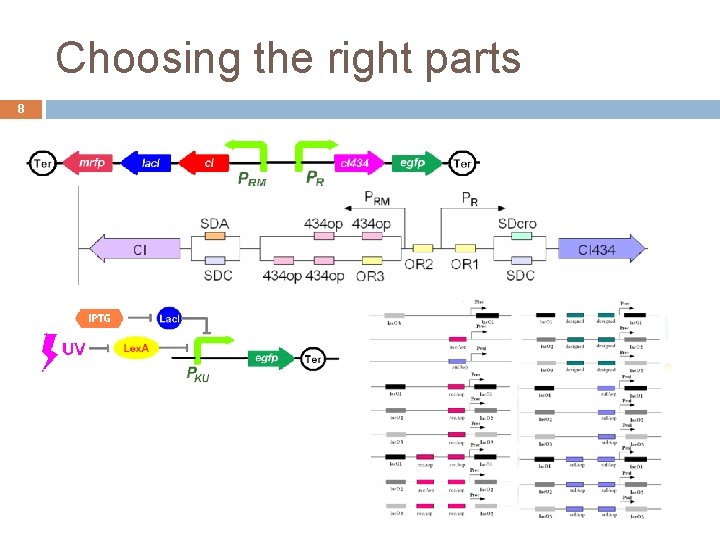
Choosing the right parts 8
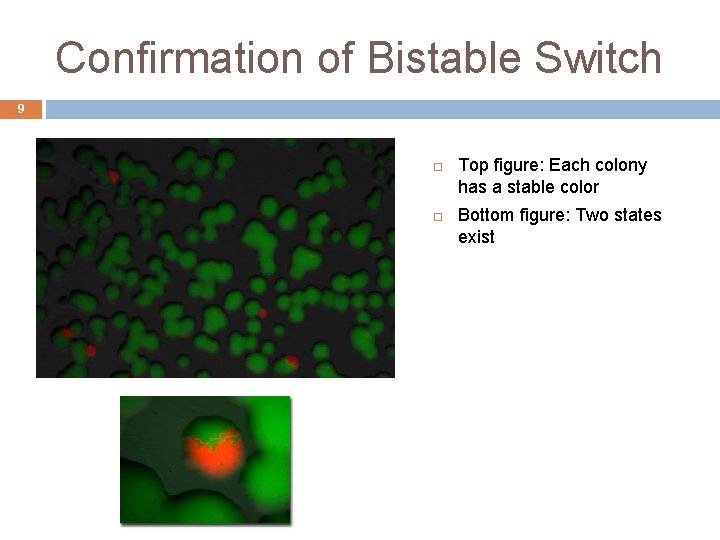
Confirmation of Bistable Switch 9 Top figure: Each colony has a stable color Bottom figure: Two states exist
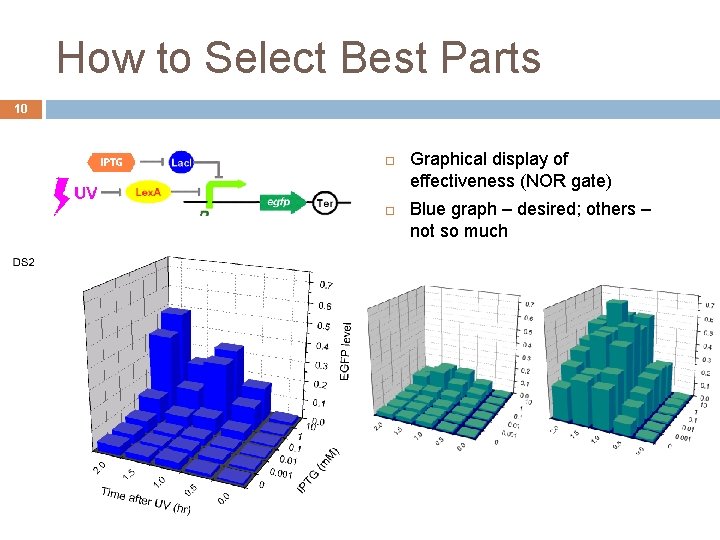
How to Select Best Parts 10 Graphical display of effectiveness (NOR gate) Blue graph – desired; others – not so much

Allen’s Thoughts 11 How was the circuit designed? The circuit is a 2 -state, cyclic system – can it be expanded? What components are specific to this implementation? Needed a NOR gate to reset from 2 nd state to 1 st state Use of computation beforehand Identical input to produce varied output based on previous history -> counter?

Hop Count 12 Spatial Seperation – need for different production systems that interfere with each other to happen in fixed proportions
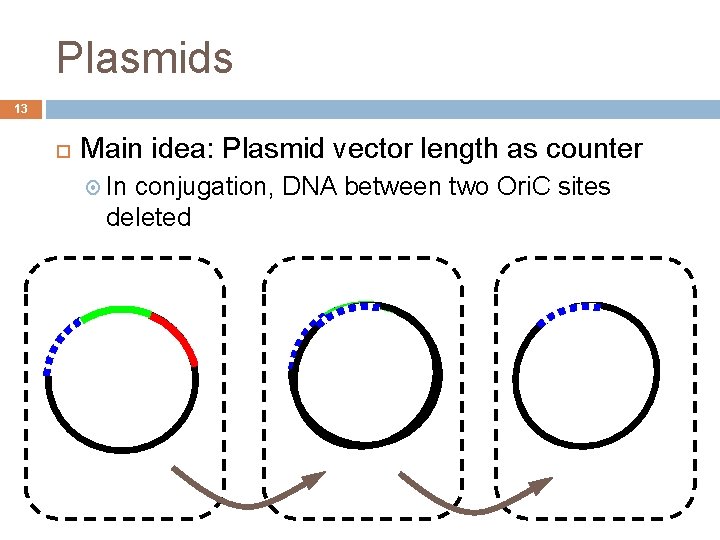
Plasmids 13 Main idea: Plasmid vector length as counter In conjugation, DNA between two Ori. C sites deleted
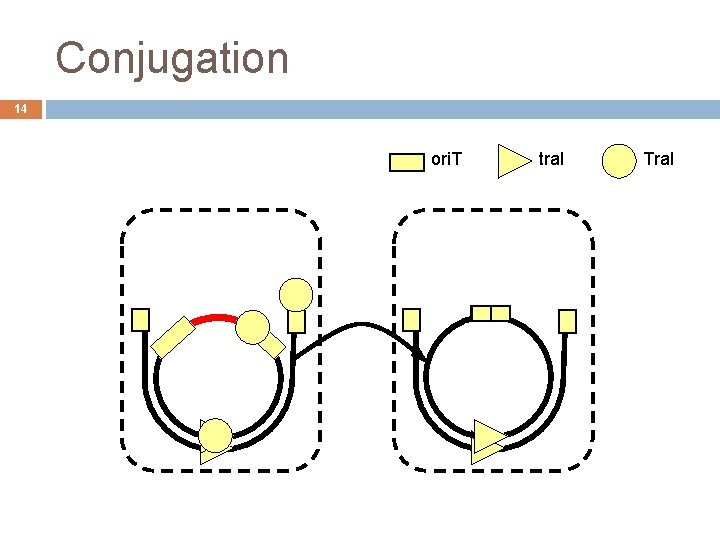
Conjugation 14 ori. T tra. I Tra. I
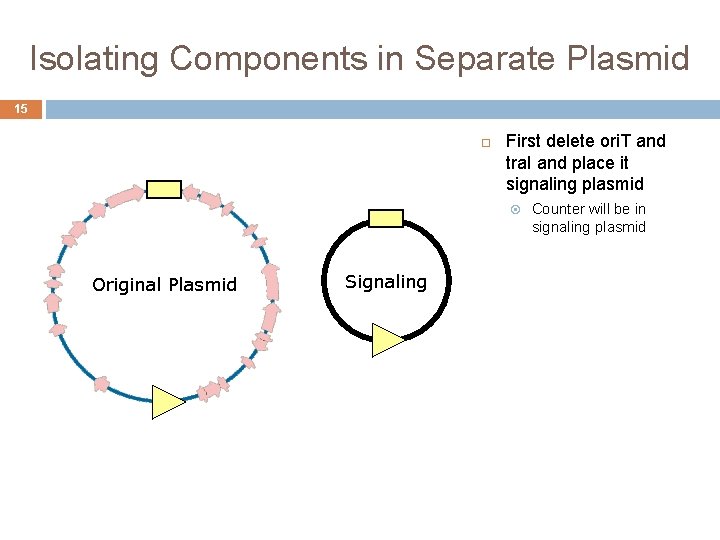
Isolating Components in Separate Plasmid 15 First delete ori. T and tra. I and place it signaling plasmid Original Plasmid Signaling Counter will be in signaling plasmid
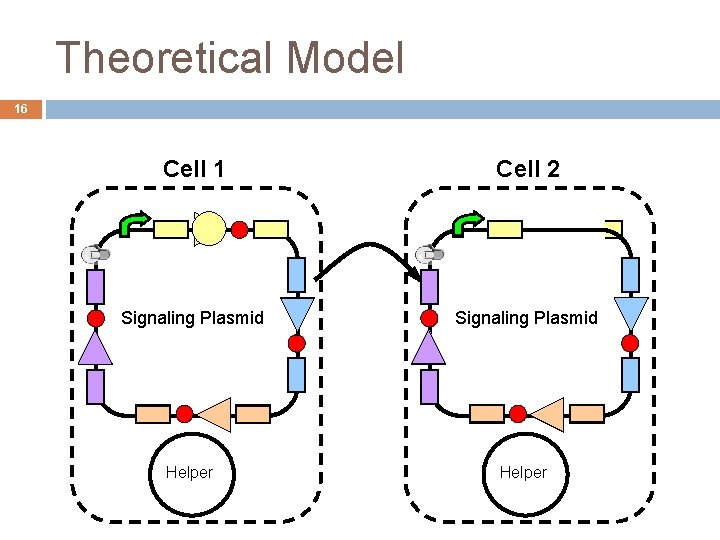
Theoretical Model 16 Cell 1 Cell 2 Signaling Plasmid Helper

Details 17 Important that the different ori. T/tra. I systems do not interfere (crosstalk) with each other Things to note: One promoter – only tra. I nearest promoter is transcribed, terminator ends that transcription DNA replication occurs in opposite direction of promoter Thus lose tra. I + terminator + functional ori. T every step
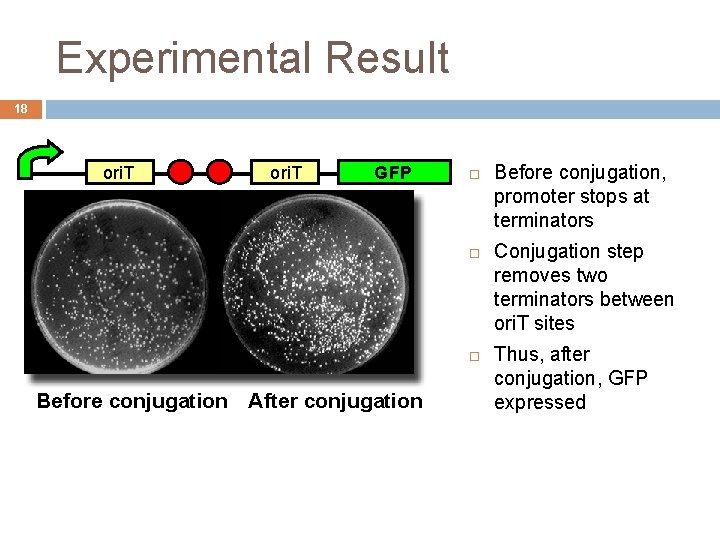
Experimental Result 18 ori. T GFP Before conjugation After conjugation Before conjugation, promoter stops at terminators Conjugation step removes two terminators between ori. T sites Thus, after conjugation, GFP expressed
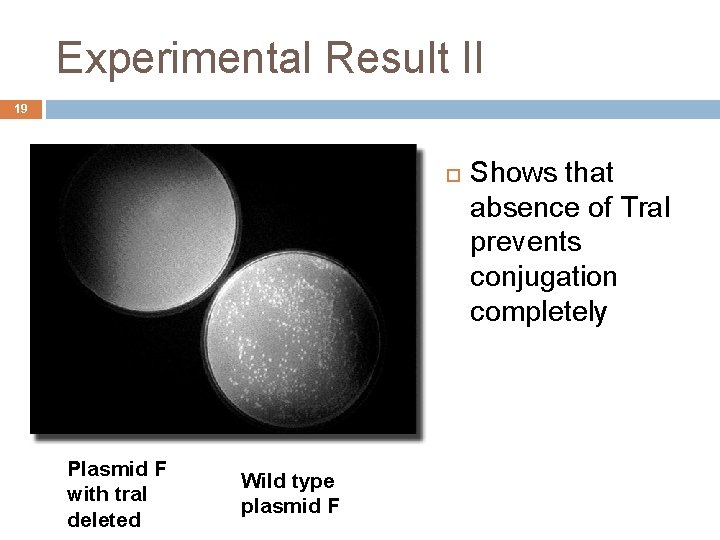
Experimental Result II 19 Plasmid F with tra. I deleted Wild type plasmid F Shows that absence of Tra. I prevents conjugation completely

Allen’s Thoughts 20 Shows two steps needed for hop count system to work once Place assembly line specific genes in between ori. T sites before terminator Ensures that one cell does each step; other cells in assembly line not far away What would happen if we combined temporal and seperation? Get bistable neighboring cells? Checkerboard configuration?

Sources 21 Peking 2007 i. GEM presentation and powerpoint http: //parts. mit. edu/igem 07/index. php/Presentatio ns Peking 2007 Wiki Project Notes http: //parts. mit. edu/igem 07/index. php/Peking_The _Projects
- Slides: 21