1 Professor Mark Ragan Institute for Molecular Bioscience

1. Professor Mark Ragan (Institute for Molecular Bioscience) 2. Dr Thomas Huber (Department of Mathematics) Computational Biology and Bioinformatics Environment Com. Bin. E Queensland Parallel Supercomputing Foundation National Facility Projects
The scientific problem: Handcrafted analyses suggest that gene transfer in nature may be not only from parents to offspring (“vertical”), but also from one lineage to another (“lateral” or “horizontal”) From microbial genomics we have complete inventories of genes & proteins in ~ 80 genomes Comparative analysis should identify all cases of vertical and lateral gene transfer Queensland Parallel Supercomputing Foundation Comparison of protein families among completely sequenced microbial genomes

Computational requirement for 80 genomes: Find all interestingly large protein families in all microbial genomes 1012 BLAST comparisons Generate structure-sensitive multiple alignments 5000 T-Coffee alignments Infer phylogenetic trees with appropriate statistics 5000 Bayesian inference trees Compare trees, look for topological incongruence 107 topological comparisons Queensland Parallel Supercomputing Foundation The approach

Usage of NF: Motif-based multiple alignment 30 -50 sequences = 2 -5 hours per run Will need ~5000 runs @ 4 - 60 seqs Code not yet parallelised Bayesian inference Parameterisation of (MC)3 search NF used for trials of up to 106 Markov chain generations (~200 hours / run) 1. 5 -2. 0 Gb RAM per run With each run costing a few 10 s of hours and need for 1000 s analyses, it’s more efficient to use many processors simultaneously Queensland Parallel Supercomputing Foundation Computations on APAC National Facility
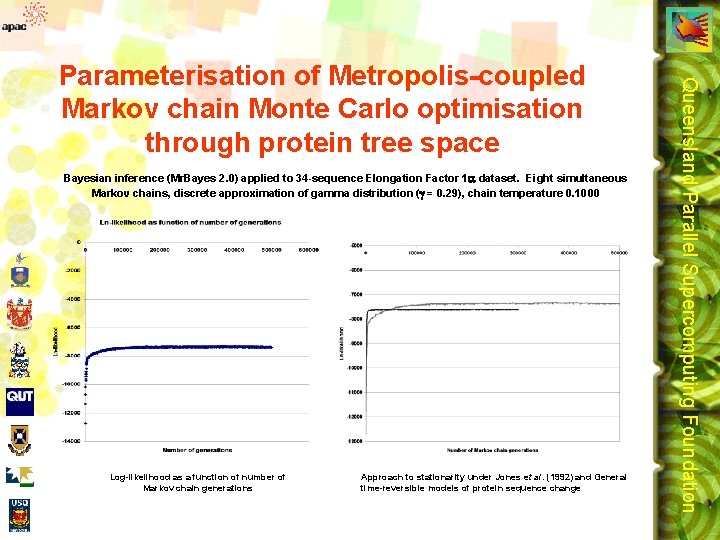
Bayesian inference (Mr. Bayes 2. 0) applied to 34 -sequence Elongation Factor 1 dataset. Eight simultaneous Markov chains, discrete approximation of gamma distribution ( = 0. 29), chain temperature 0. 1000 Log-likelihood as a function of number of Markov chain generations Approach to stationarity under Jones et al. (1992) and General time-reversible models of protein sequence change Queensland Parallel Supercomputing Foundation Parameterisation of Metropolis-coupled Markov chain Monte Carlo optimisation through protein tree space

Mark Borodovsky, Georgia Tech Robert Charlebois, NGI Inc. (Ottawa) Tim Harlow, University of Queensland Jeffrey Lawrence, University of Pittsburgh Thomas Rand, St Mary’s University Queensland Parallel Supercomputing Foundation With thanks to collaborators

1. Professor Mark Ragan (Institute for Molecular Bioscience) 2. Dr Thomas Huber (Department of Mathematics) Computational Biology and Bioinformatics Environment Com. Bin. E Queensland Parallel Supercomputing Foundation National Facility Projects

Protein Structure Prediction • The bioinformatics approach – Compare sequence to other sequence – huge datasets (0. 5*106 sequences) – Match sequence with known structure – (Low resolution force field development) • The biophysics approach – Simulations that mimic natural behaviour Queensland Parallel Supercomputing Foundation Two Lineages
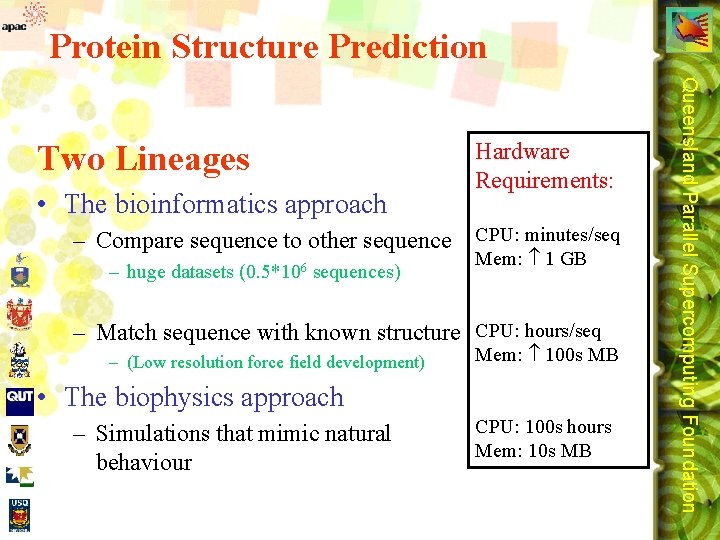
Protein Structure Prediction • The bioinformatics approach Hardware Requirements: – Compare sequence to other sequence CPU: minutes/seq – huge datasets (0. 5*106 sequences) Mem: 1 GB – Match sequence with known structure CPU: hours/seq – (Low resolution force field development) Mem: 100 s MB • The biophysics approach – Simulations that mimic natural behaviour CPU: 100 s hours Mem: 10 s MB Queensland Parallel Supercomputing Foundation Two Lineages
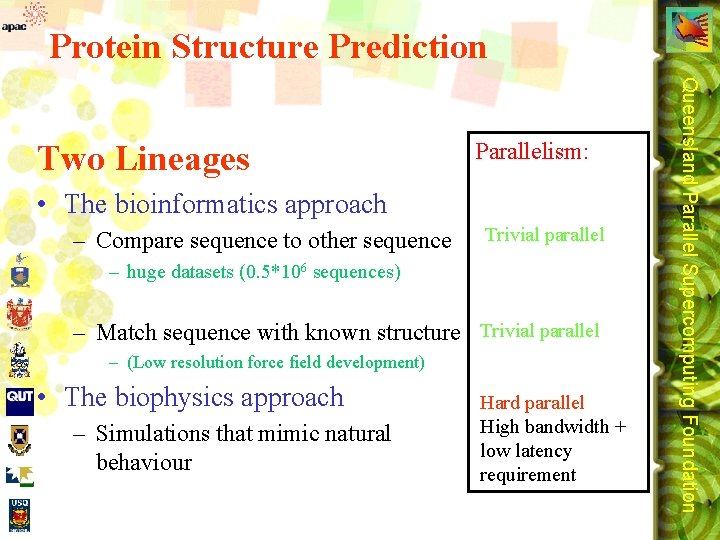
Protein Structure Prediction Parallelism: • The bioinformatics approach – Compare sequence to other sequence Trivial parallel – huge datasets (0. 5*106 sequences) – Match sequence with known structure Trivial parallel – (Low resolution force field development) • The biophysics approach – Simulations that mimic natural behaviour Hard parallel High bandwidth + low latency requirement Queensland Parallel Supercomputing Foundation Two Lineages
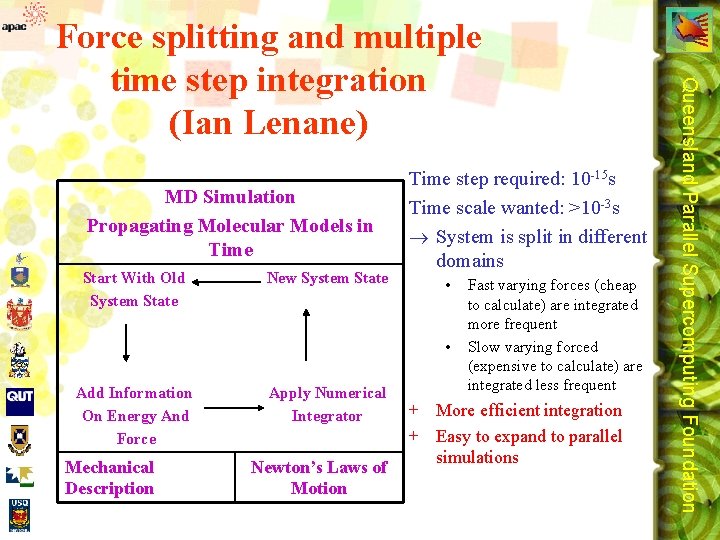
MD Simulation Propagating Molecular Models in Time Start With Old System State New System State Time step required: 10 -15 s Time scale wanted: >10 -3 s ® System is split in different domains • • Add Information On Energy And Force Mechanical Description Apply Numerical Integrator Newton’s Laws of Motion Fast varying forces (cheap to calculate) are integrated more frequent Slow varying forced (expensive to calculate) are integrated less frequent + More efficient integration + Easy to expand to parallel simulations Queensland Parallel Supercomputing Foundation Force splitting and multiple time step integration (Ian Lenane)

What if start and end points are given? • proteins: unfolded • Molecular machines: 1 cycle • Shortest path calculations – Floyd, Dijkstra • Hamilton’s principle of least action + Computationally very attractive • Extremely long time steps • Very well suited for parallel architectures (Floyd algorithm parallelized, but performance problems >4 PE on -GS NUMA architecture) Queensland Parallel Supercomputing Foundation Path simulations (Ben Gladwin)

• 2001 CPU quota: 2*5250 + 8000 service units – Total use 12000 units ( 3000 units in parallel) • 2002 CPU quota: 4 * 6000 service units – First quarter: 2000 units – Second quarter: 85 units • Collaborators • Dr A. Torda (ANU) Low resolution force fields / protein structure prediction • Prof. D. Hume, A/Prof. B. Kobe and Dr. J. Martin (UQ) Structural genomics project • Prof. K. Burrage, I. Lenane and B. Galdwin (UQ) Numerical integration and path simulations • Special Thanks • Mrs J. Jenkinson and Dr D. Singleton (NF/ANUSF) Queensland Parallel Supercomputing Foundation National Facility supercomputer use
- Slides: 13