1 Pacific Biosciences Use a circular template to

1 Pacific Biosciences Use a circular template to get redundant reads and so more accuracy.
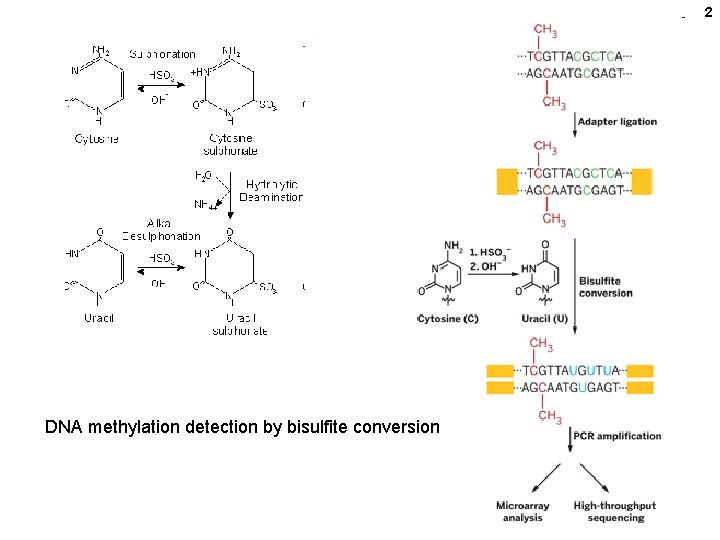
2 DNA methylation detection by bisulfite conversion
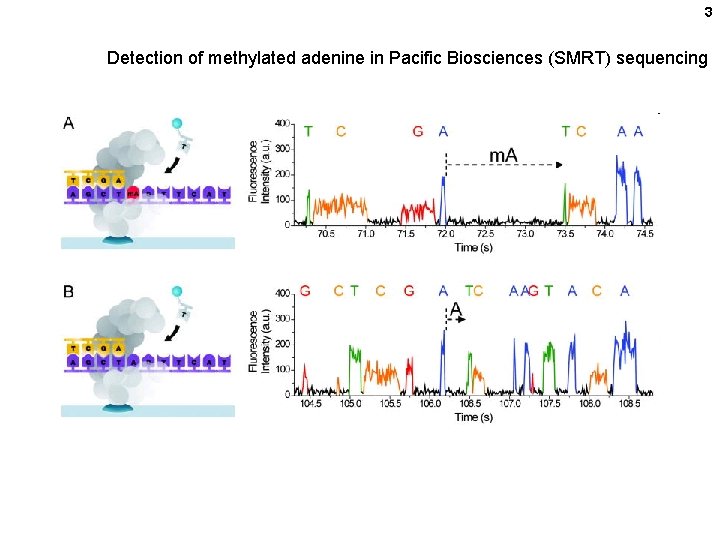
3 Detection of methylated adenine in Pacific Biosciences (SMRT) sequencing
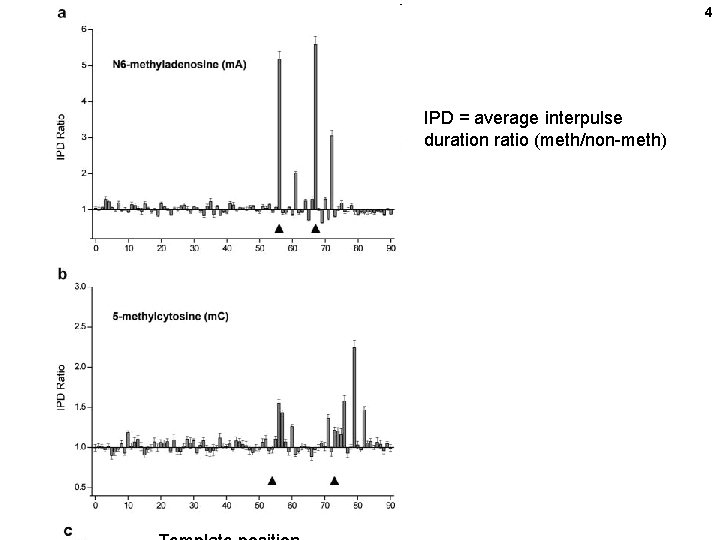
4 IPD = average interpulse duration ratio (meth/non-meth)

5 Pacific Biosciences • 50, 000 ZMWs (Aug. , 2011), and density may climb • Long reads (e. g. , full molecules to determine full length splicing isoforms) • Direct RNA sequencing possible. • DNA methylation detectable
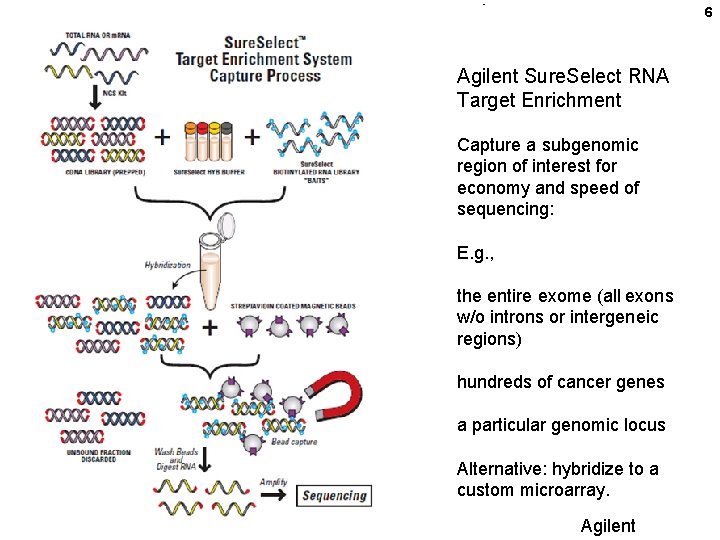
6 Agilent Sure. Select RNA Target Enrichment Capture a subgenomic region of interest for economy and speed of sequencing: E. g. , the entire exome (all exons w/o introns or intergeneic regions) hundreds of cancer genes a particular genomic locus Alternative: hybridize to a custom microarray. Agilent
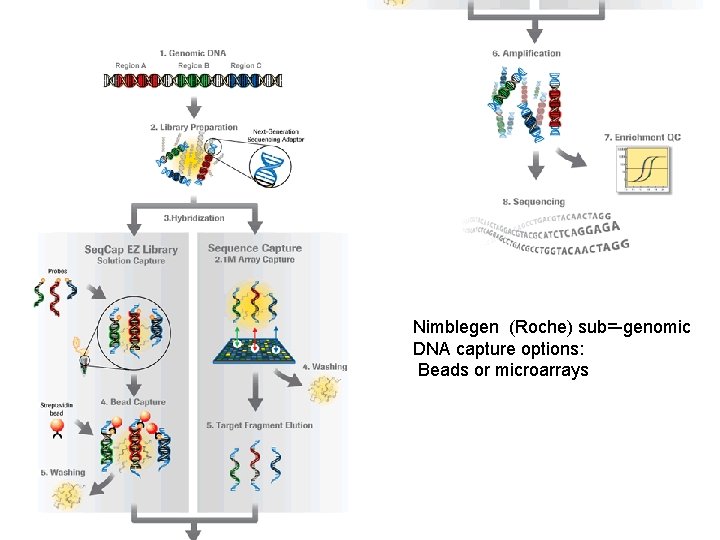
7 Nimblegen (Roche) sub=-genomic DNA capture options: Beads or microarrays
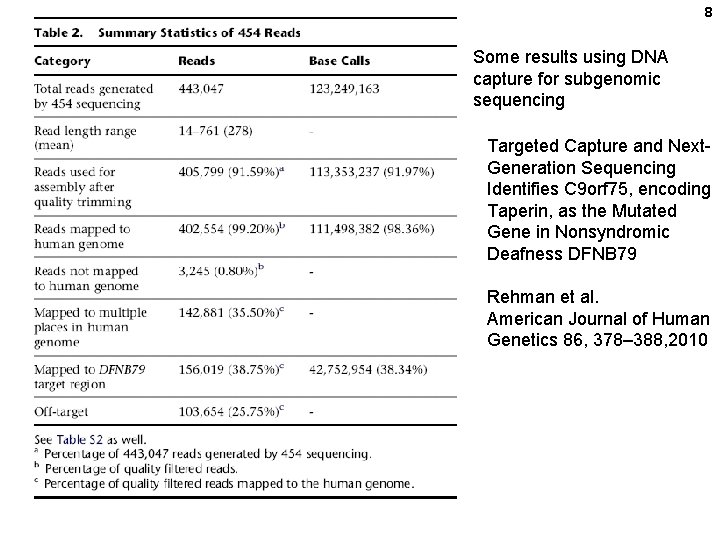
8 Some results using DNA capture for subgenomic sequencing Targeted Capture and Next. Generation Sequencing Identifies C 9 orf 75, encoding Taperin, as the Mutated Gene in Nonsyndromic Deafness DFNB 79 Rehman et al. American Journal of Human Genetics 86, 378– 388, 2010
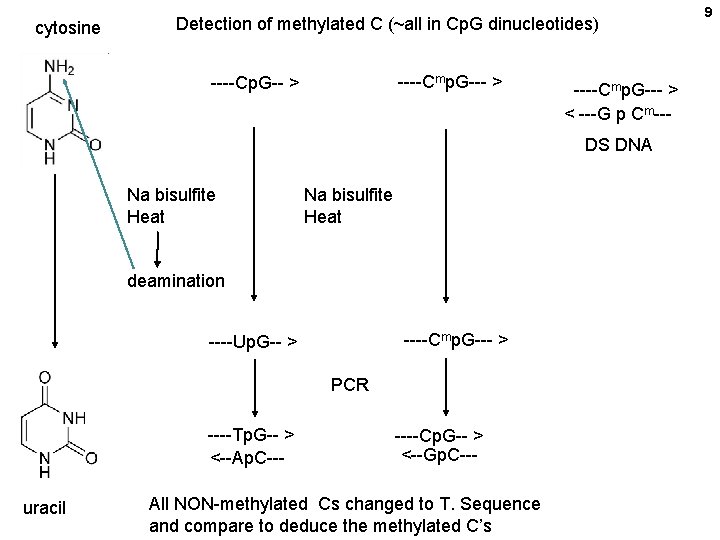
cytosine Detection of methylated C (~all in Cp. G dinucleotides) ----Cmp. G--- > ----Cp. G-- > ----Cmp. G--- > < ---G p Cm--DS DNA Na bisulfite Heat deamination ----Cmp. G--- > ----Up. G-- > PCR ----Tp. G-- > <--Ap. C--uracil ----Cp. G-- > <--Gp. C--- All NON-methylated Cs changed to T. Sequence and compare to deduce the methylated C’s 9

10 DEEP SEQUENCING (Next generation sequencing, High throughput sequencing, Massively parallel sequencing) applications: Human genome re-sequencing (mutations, SNPs, haplotypes, disease associations, personalized medicine) Tumor genome sequencing Microbial flora sequencing (microbiome, viruses) Metagenomic sequencing (without cell culturing) RNA sequencing (RNAseq; gene expression levels, mi. RNAs, lnc. RNAs, splicing isoforms) Chromatin structure (Ch. IP-seq; histone modifications, nucleosome positioning) Epigenetic modifications (DNA Cp. G methylation and hydroxymethylation) Transcription kinetics (GROseq; nascent RNA, Brd. U pulse labeled RNA) High throughput genetics (QUEPASA; cis-acting regulatory motif discovery) Drug discovery (bar-coded organic molecule libraries) [Manocci PNAS paper]

11 Ke et al, and Chasin, Quantitative evaluation of all hexamers as exonic splicing elements. Genome Res. 2011. 21: 1360 -1374 ). Order an equal mixture of all 4 bases at these 6 positions
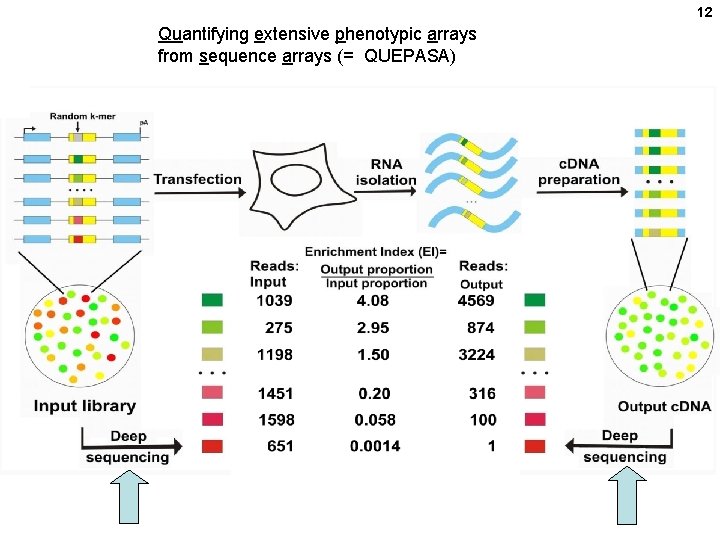
12 Quantifying extensive phenotypic arrays from sequence arrays (= QUEPASA)
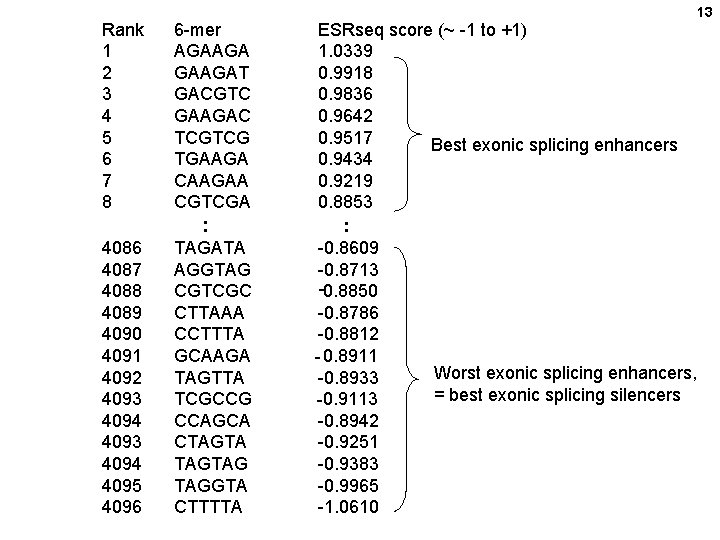
Rank 1 2 3 4 5 6 7 8 6 -mer AGAAGAT GACGTC GAAGAC TCGTCG TGAAGA CAAGAA CGTCGA : 4086 4087 4088 4089 4090 4091 4092 4093 4094 4095 4096 TAGATA AGGTAG CGTCGC CTTAAA CCTTTA GCAAGA TAGTTA TCGCCG CCAGCA CTAGTAG TAGGTA CTTTTA ESRseq score (~ -1 to +1) 1. 0339 0. 9918 0. 9836 0. 9642 0. 9517 Best exonic splicing enhancers 0. 9434 0. 9219 0. 8853 : -0. 8609 -0. 8713 -0. 8850 -0. 8786 -0. 8812 - 0. 8911 Worst exonic splicing enhancers, -0. 8933 = best exonic splicing silencers -0. 9113 -0. 8942 -0. 9251 -0. 9383 -0. 9965 -1. 0610 13
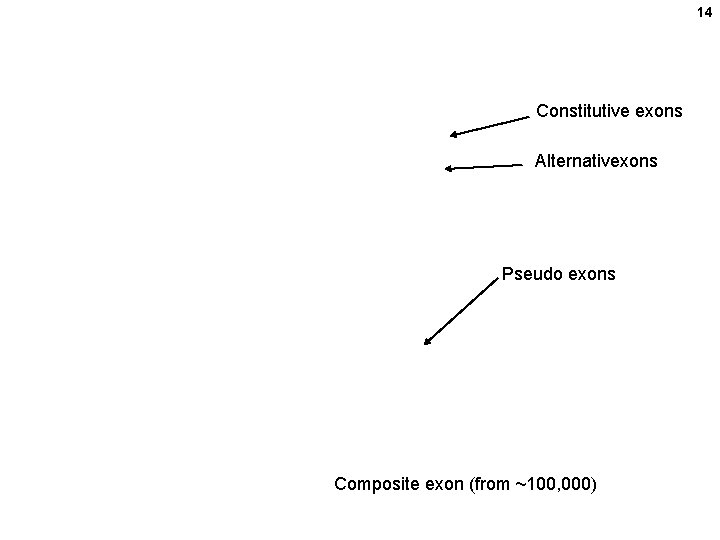
14 Constitutive exons Alternativexons Pseudo exons Composite exon (from ~100, 000)
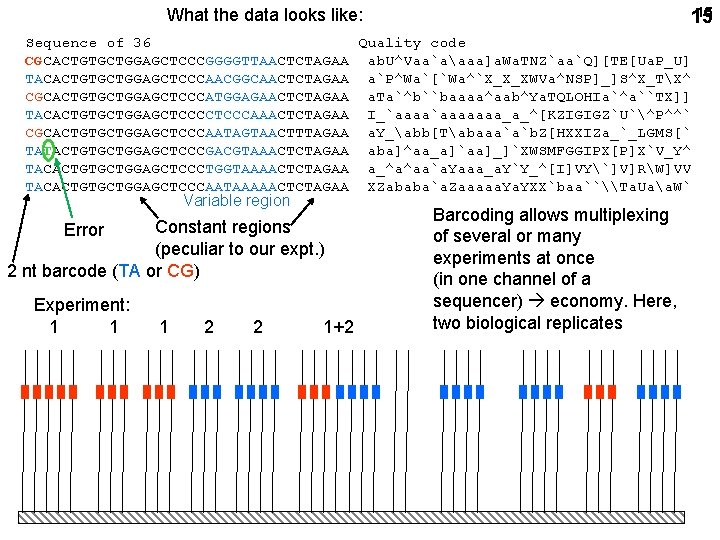
15 15 What the data looks like: Sequence of 36 Quality code CGCACTGTGCTGGAGCTCCCGGGGTTAACTCTAGAA ab. U^Vaa`aaaa]a. Wa. TNZ`aa`Q][TE[Ua. P_U] TACACTGTGCTGGAGCTCCCAACGGCAACTCTAGAA a`P^Wa`[`Wa^`X_X_XWVa^NSP]_]S^X_TX^ CGCACTGTGCTGGAGCTCCCATGGAGAACTCTAGAA a. Ta`^b``baaaa^aab^Ya. TQLOHIa`^a``TX]] TACACTGTGCTGGAGCTCCCAAACTCTAGAA I_`aaaaaaa_a_^[KZIGIGZ`U`^P^^` CGCACTGTGCTGGAGCTCCCAATAGTAACTTTAGAA a. Y_abb[Tabaaa`a`b. Z[HXXIZa_`_LGMS[` TATACTGTGCTGGAGCTCCCGACGTAAACTCTAGAA aba]^aa_a]`aa]_]`XWSMFGGIPX[P]X`V_Y^ TACACTGTGCTGGAGCTCCCTGGTAAAACTCTAGAA a_^a^aa`a. Yaaa_a. Y`Y_^[I]VY`]V]RW]VV TACACTGTGCTGGAGCTCCCAATAAAAACTCTAGAA XZababa`a. Zaaaaa. YXX`baa``\Ta. Uaa. W` Variable region Constant regions (peculiar to our expt. ) 2 nt barcode (TA or CG) Error Experiment: 1 1 1 2 2 1+2 Barcoding allows multiplexing of several or many experiments at once (in one channel of a sequencer) economy. Here, two 2 biological 2 replicates 1 2
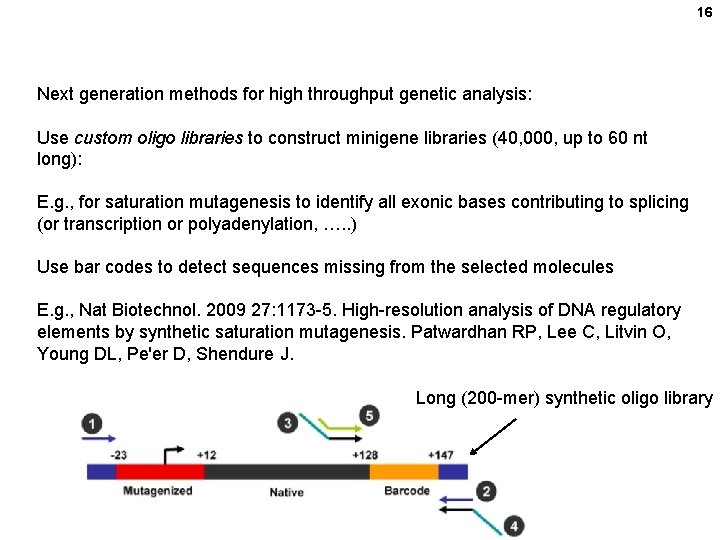
16 Next generation methods for high throughput genetic analysis: Use custom oligo libraries to construct minigene libraries (40, 000, up to 60 nt long): E. g. , for saturation mutagenesis to identify all exonic bases contributing to splicing (or transcription or polyadenylation, …. . ) Use bar codes to detect sequences missing from the selected molecules E. g. , Nat Biotechnol. 2009 27: 1173 -5. High-resolution analysis of DNA regulatory elements by synthetic saturation mutagenesis. Patwardhan RP, Lee C, Litvin O, Young DL, Pe'er D, Shendure J. Long (200 -mer) synthetic oligo library

OUTLINE OF LECTURE TOPICS COMING UP Expression and manipulation of transgenes in the laboratory • In vitro mutagenesis to isolate variants of your protein/gene with desirable properties – – • To study the protein: Express your transgene – – – • • • Single base mutations Deletions Overlap extension PCR Cassette mutagenesis Usually in E. coli, for speed, economy Expression in eukaryotic hosts Drive it with a promoter/enhancer Purify it via a protein tag Cleave it to get the pure protein Explore protein-protein interaction Co-immunoprecipitation (co-IP) from extracts 2 -hybrid formation surface plasmon resonance FRET (Fluorescence resonance energy transfer) Complementation readout 17 17
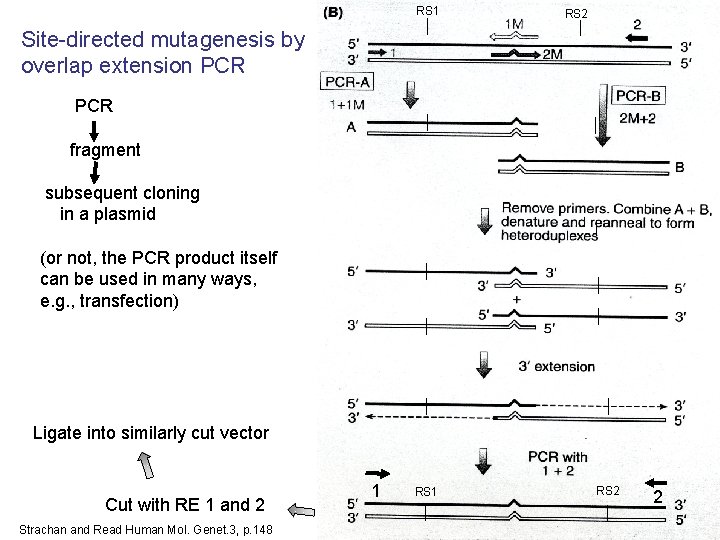
RS 1 18 18 RS 2 Site-directed mutagenesis by overlap extension PCR fragment subsequent cloning in a plasmid (or not, the PCR product itself can be used in many ways, e. g. , transfection) Ligate into similarly cut vector Cut with RE 1 and 2 Strachan and Read Human Mol. Genet. 3, p. 148 1 RS 2 2

19 19 Cassette mutagenesis = random mutagenesis but in a limited region: 1) by error-prone PCR ----------------------------------------------------------- Original sequence coding for, e. g. , a transcription enhancer region PCR fragment with high Taq polymerase and Mn+2 instead of Mg+2 errors ------*--*-**--------*------*--*--------------*-*-*------------*-- Cut in primer sites and clone upstream of a reporter protein sequence. Pick colonies Analyze phenotypes Sequence

Cassette mutagenesis = random mutagenesis but in a limited region: 20 20 2) by “doped” synthesis Target = e. g. , an enhancer element -----------------------------Original enhancer sequence ------------------------------*------------*-*-*------------*--------*--*-**--------*------*--*------ Clone upstream of a reporter. Pick colonies Analyze phenotypes Sequence Buy 2 doped oligos; anneal OK for up to ~80 nt. Doping = e. g. , 90% G, 3. 3% A, 3. 3% C, 3. 3% T at each position

21 21 E. coli as a host • PROs: Easy, flexible, high tech, fast, cheap; but problems • • • CONs Folding (can misfold) Sorting within the cell -> can form inclusion bodies Purification -- endotoxins Modifications -- not done (glycosylation, phosphorylation, etc. ) • • • Modifications: Glycoproteins Acylation: acetylation, myristoylation Methylation (arg, lys) Phosphorylation (ser, thr, tyr) Sulfation (tyr) Prenylation (farnesyl, geranyl on cys) Vitamin C-Dependent Modifications (hydroxylation of proline and lysine) Vitamin K-Dependent Modifications (gamma carboxylation of glu) Selenoproteins (seleno-cys t. RNA at UGA stop)

E. coli expression vectors Promoter examples: 1) Lac promoter (with operator)-YFG, + lac repressor (I gene): Induce expression by inactivationof thelac repressor with IPTG or lactose 2) As above but with a hybrid Tac promoter (tryptophan operon + lac operon): Stronger. Use iq mutant of lac I gene, which prodices high levels of the lac repressor. Expression regulatatable over several orders of magnitude. 3) BAD promoter-YFG. Arabinose utilization operon. Inducible by arabinose via the endogenous ara. C gene for a transciptional activator. Background levels driven down by including glucose. 4) Phage T 7 promoter-YFG. Vector carries gene for T 7 polymerase, under control of the lac promoter. Add IPTG or lactose to induce T 7 polymerase and thence YFG. IPTG = isoproplthiogalactoside (non-metabolizable indicer) YFG = your favorite gene
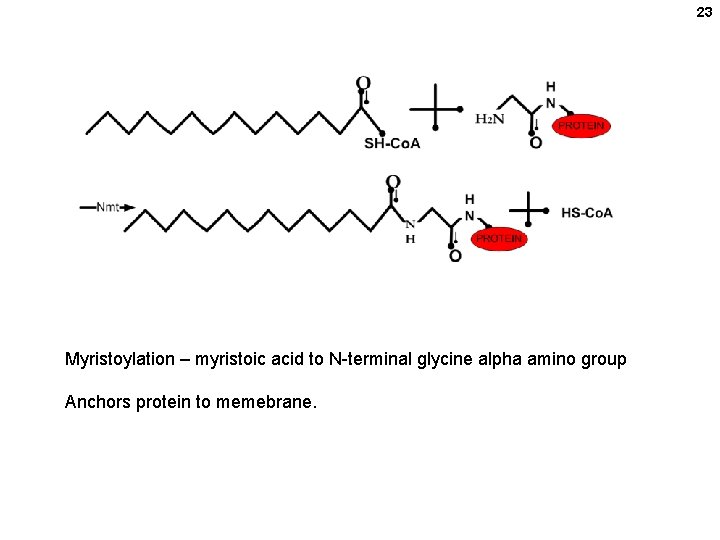
23 Myristoylation – myristoic acid to N-terminal glycine alpha amino group Anchors protein to memebrane.
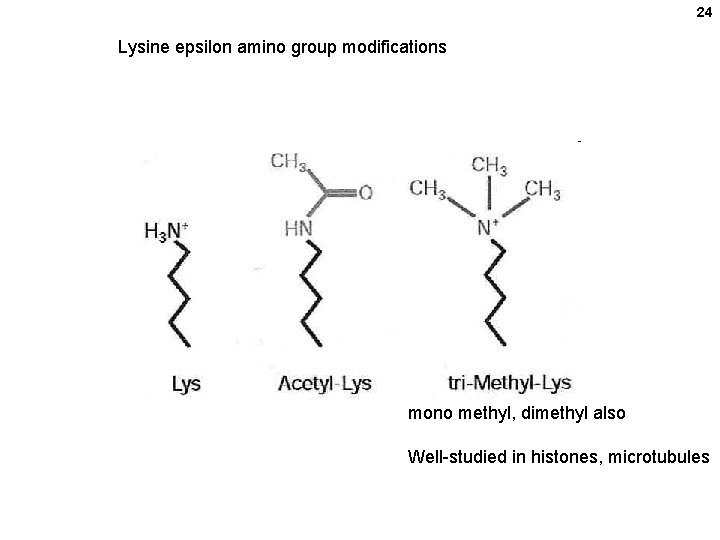
24 Lysine epsilon amino group modifications mono methyl, dimethyl also Well-studied in histones, microtubules
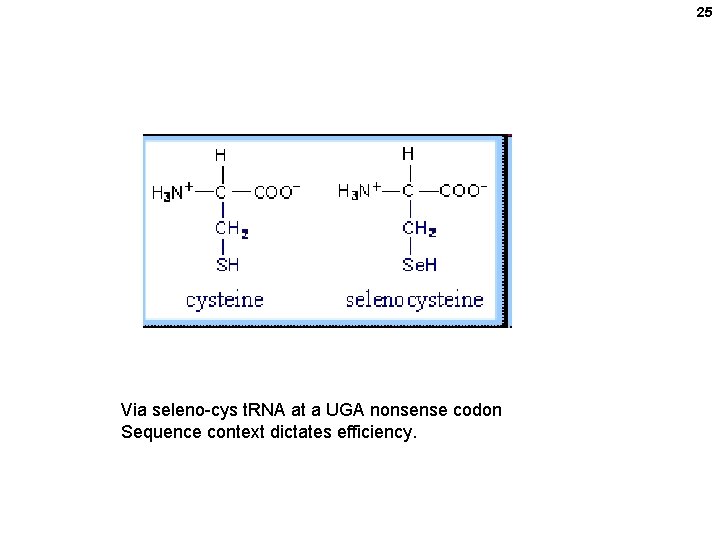
25 Via seleno-cys t. RNA at a UGA nonsense codon Sequence context dictates efficiency.
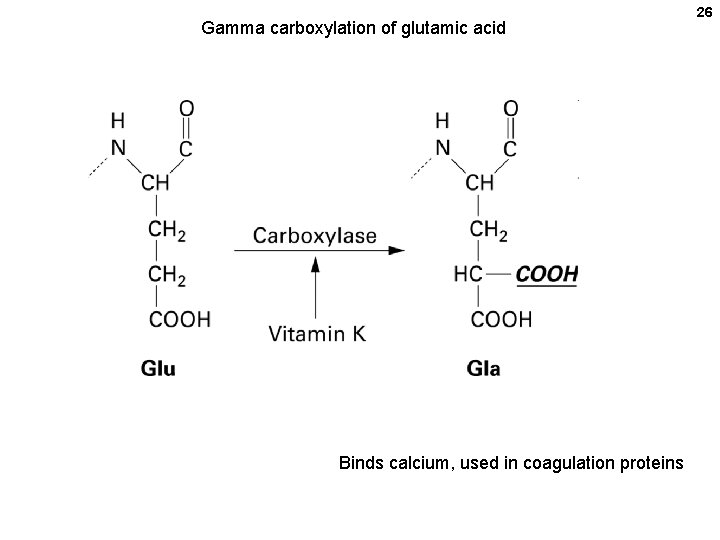
Gamma carboxylation of glutamic acid Binds calcium, used in coagulation proteins 26

27 27 Some alternative hosts • • Yeasts (Saccharomyces , Pichia) Insect cells with baculovirus vectors Mammalian cells in culture (later) Whole organisms (mice, goats, corn) (not discussed) • In vitro (cell-free), for analysis only, not preparatively (good for radiolabeled proteins, discussed later)

Some popular yeast promoters Selectable marker ori http: //biochemie. web. med. unimuenchen. de/Yeast_Biol/04 Yeast Molecular Techniques. pdf ARS = autonomously replicating sequence element
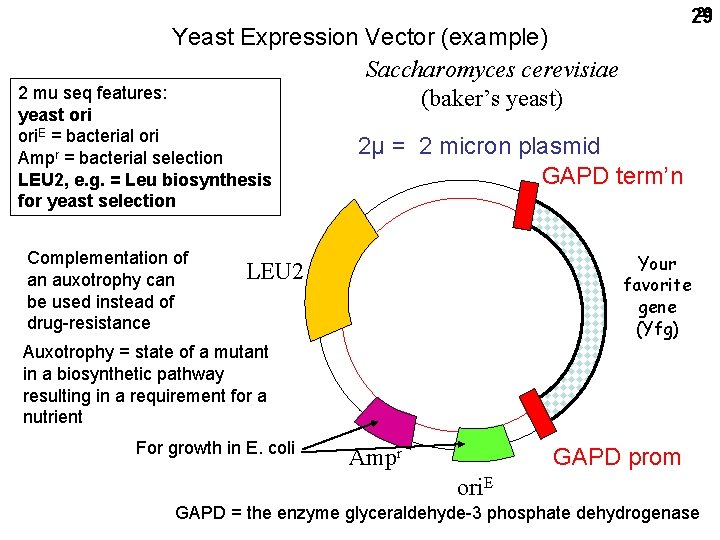
29 29 Yeast Expression Vector (example) Saccharomyces cerevisiae 2 mu seq features: (baker’s yeast) yeast ori. E = bacterial ori Ampr = bacterial selection LEU 2, e. g. = Leu biosynthesis for yeast selection Complementation of an auxotrophy can be used instead of drug-resistance 2μ = 2 micron plasmid GAPD term’n Your favorite gene (Yfg) LEU 2 Auxotrophy = state of a mutant in a biosynthetic pathway resulting in a requirement for a nutrient For growth in E. coli GAPD prom Ampr ori. E GAPD = the enzyme glyceraldehyde-3 phosphate dehydrogenase

Got this far
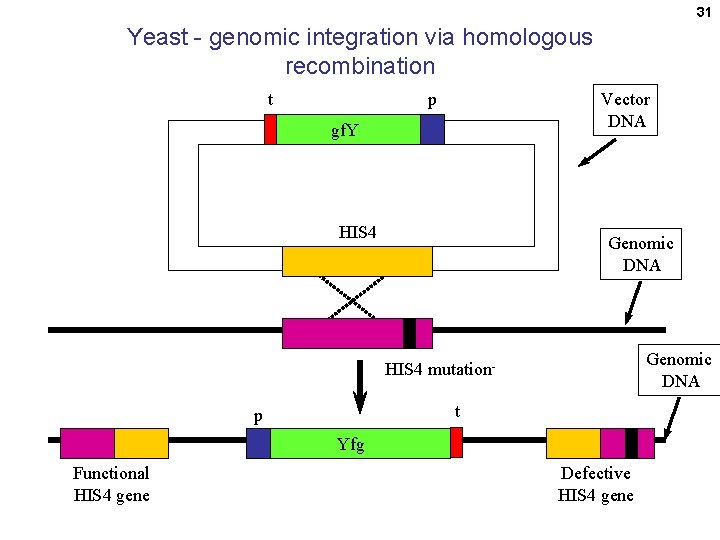
31 Yeast - genomic integration via homologous recombination t Vector DNA p gf. Y HIS 4 Genomic DNA HIS 4 mutationt p Yfg Functional HIS 4 gene Defective HIS 4 gene
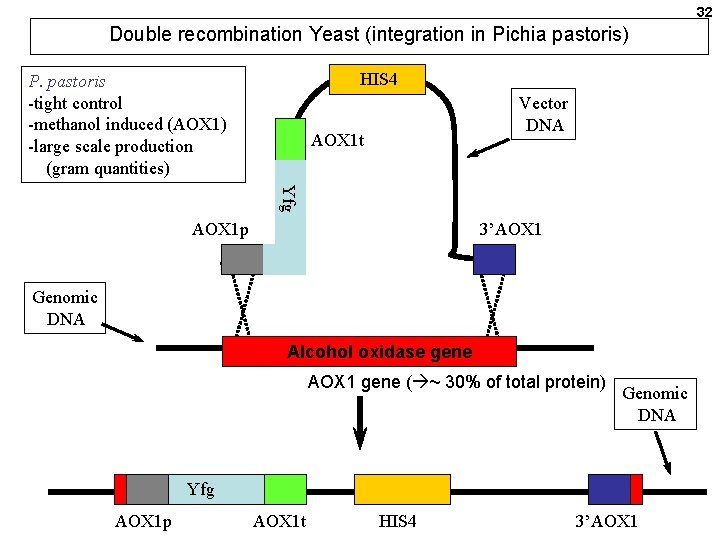
32 Double recombination Yeast (integration in Pichia pastoris) HIS 4 P. pastoris -tight control -methanol induced (AOX 1) -large scale production (gram quantities) Vector DNA AOX 1 t Yfg AOX 1 p 3’AOX 1 Genomic DNA Alcohol oxidase gene AOX 1 gene ( ~ 30% of total protein) Genomic DNA Yfg AOX 1 p AOX 1 t HIS 4 3’AOX 1
Expression in mammalian cells Lab examples of immortal cell lines: HEK 293 Human embyonic kidney (high transfection efficiency) He. La Human cervical carcinoma (historical, low RNase) CHO Chinese hamster ovary (hardy, diploid DNA content, mutants) Cos Monkey cells with SV 40 replication proteins (-> high transgene copies) 3 T 3 Mouse or human exhibiting ~regulated (normal-like) growth + various others, many differentiated to different degrees, e. g. : BHK Baby hamster kidney Hep. G 2 Human hepatoma GH 3 Rat pituitary cells PC 12 Mouse neuronal-like tumor cells MCF 7 Human breast cancer HT 1080 Human fibroblastic cells with near diploid karyotype IPS induced pluripotent stem cells and: Primary cells cultured with a limited lifetime. E. g. , MEF = mouse embryonic fibroblasts, HDF = Human diploid fibroblasts Common in industry: NS 1 m. Abs Vero vaccines CHO m. Abs, otherapeutic proteins PER 6 m. Abs, otherapeutic proteins Mouse plasma cell tumor cells African greem monkey cells Chinese hamster ovary cells Human retinal cells
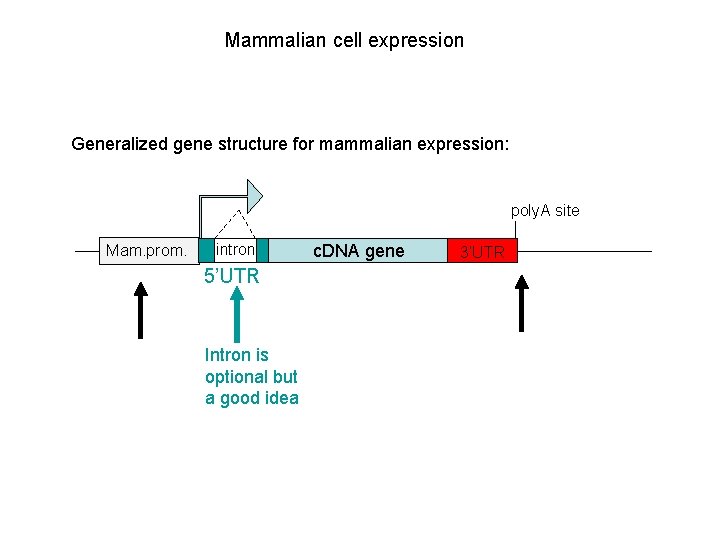
Mammalian cell expression Generalized gene structure for mammalian expression: poly. A site Mam. prom. intron 5’UTR Intron is optional but a good idea c. DNA gene 3’UTR

Popular mammalian cell promoters • • • SV 40 Large. T Ag (Simian Virus 40) RSV LTR (Rous sarcoma virus) MMTV (steroid inducible) (Mouse mammary tumor virus) HSV TK (low expression) (Herpes simplex virus) Metallothionein (metal inducible, Cd++) CMV early (Cytomegalovirus) Actin EIF 2 alpha Engineered inducible / repressible: tet, ecdysone, glucocorticoid (tet = tetracycline)
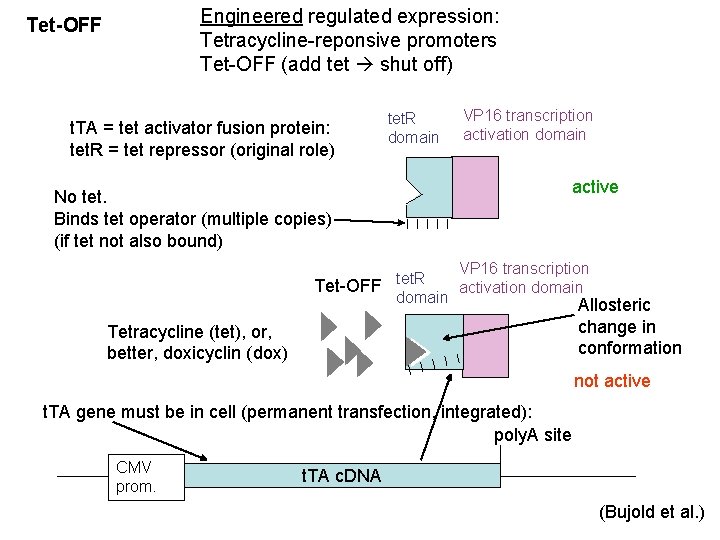
Engineered regulated expression: Tetracycline-reponsive promoters Tet-OFF (add tet shut off) Tet-OFF t. TA = tet activator fusion protein: tet. R = tet repressor (original role) tet. R domain VP 16 transcription activation domain No tet. Binds tet operator (multiple copies) (if tet not also bound) active VP 16 transcription tet. R Tet-OFF activation domain Allosteric change in conformation Tetracycline (tet), or, better, doxicyclin (dox) not active t. TA gene must be in cell (permanent transfection, integrated): poly. A site CMV prom. t. TA c. DNA (Bujold et al. )
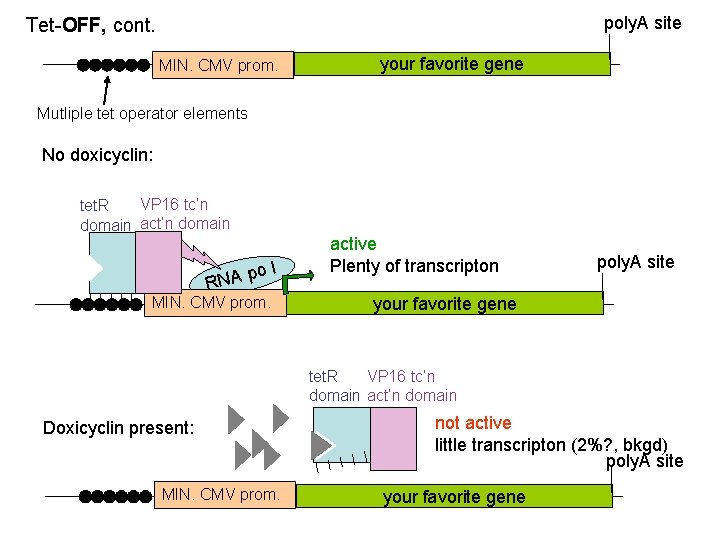
poly. A site Tet-OFF, cont. MIN. CMV prom. your favorite gene Mutliple tet operator elements No doxicyclin: VP 16 tc’n tet. R domain act’n domain po RNA l MIN. CMV prom. active Plenty of transcripton poly. A site your favorite gene tet. R VP 16 tc’n domain act’n domain Doxicyclin present: MIN. CMV prom. not active little transcripton (2%? , bkgd) poly. A site your favorite gene
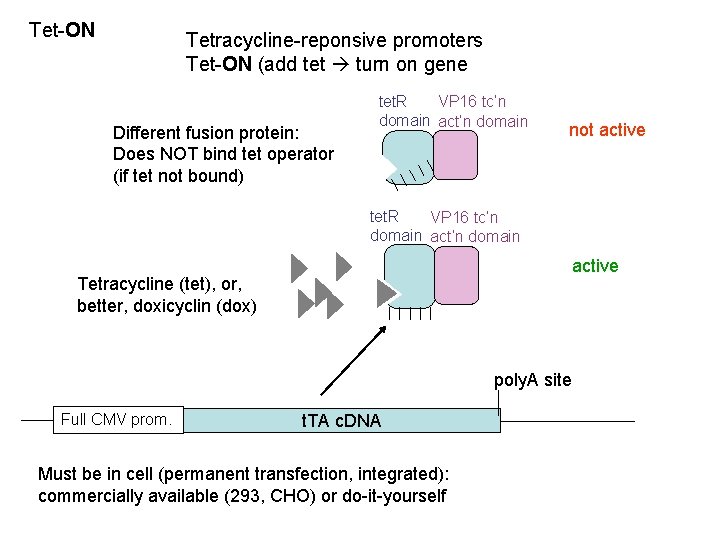
Tet-ON Tetracycline-reponsive promoters Tet-ON (add tet turn on gene Different fusion protein: Does NOT bind tet operator (if tet not bound) tet. R VP 16 tc’n domain act’n domain not active tet. R VP 16 tc’n domain active Tetracycline (tet), or, better, doxicyclin (dox) poly. A site Full CMV prom. t. TA c. DNA Must be in cell (permanent transfection, integrated): commercially available (293, CHO) or do-it-yourself
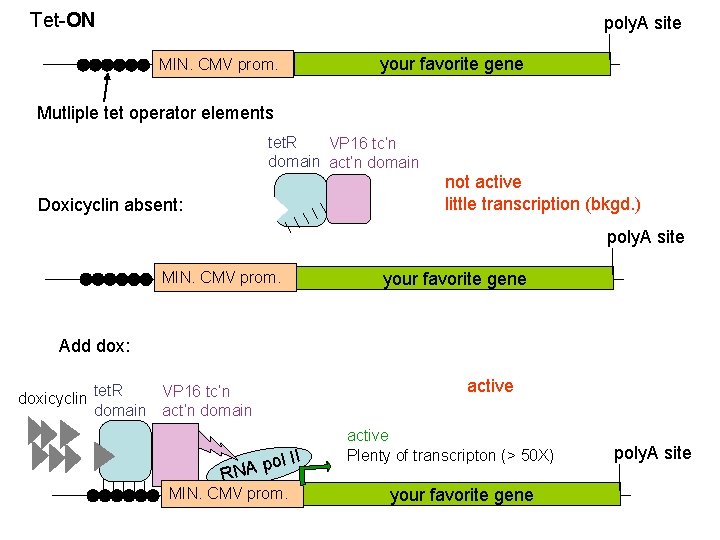
Tet-ON poly. A site MIN. CMV prom. your favorite gene Mutliple tet operator elements tet. R VP 16 tc’n domain act’n domain not active little transcription (bkgd. ) Doxicyclin absent: poly. A site MIN. CMV prom. your favorite gene Add dox: VP 16 tc’n doxicyclin tet. R domain act’n domain R ol II p A N MIN. CMV prom. active Plenty of transcripton (> 50 X) your favorite gene poly. A site
- Slides: 39