1 MTHFD 1 controls DNA methylation in Arabidopsis

1 MTHFD 1 controls DNA methylation in Arabidopsis Speaker:Kai-Ya Chang Advisor:Shih-Feng Fu Date: 2016. 10. 21

2 DNA methylation DNA methyltransferases (DNMT) G A C T G A CH 2 O N SAM C 5 N 6 H 4 M S M C C 3 H 5 NH 2 COOH CH 3 M 。transposable elements (TEs) 。repetitive DNA G C C G M M

3 Methylation site CG CHH ( H:A、T、G ) gene bodies pericentromeric transposable elements (TEs)
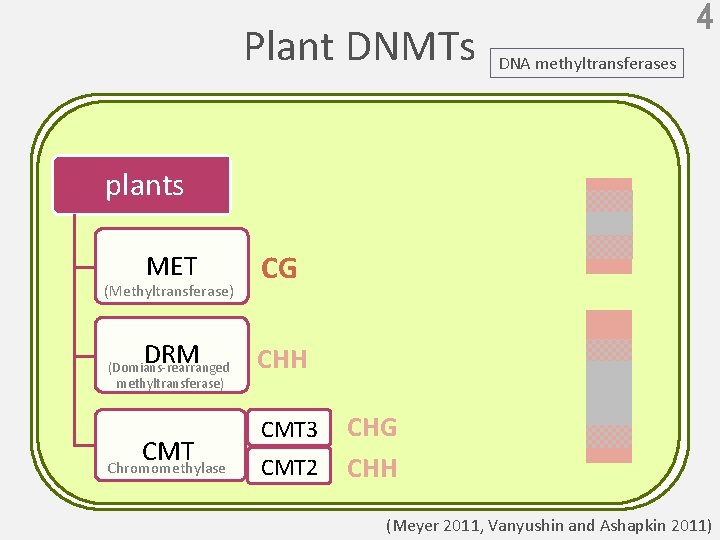
Plant DNMTs 4 DNA methyltransferases plants MET CG DRM CHH (Methyltransferase) (Domians-rearranged methyltransferase) CMT Chromomethylase CMT 3 CMT 2 CHG CHH (Meyer 2011, Vanyushin and Ashapkin 2011)
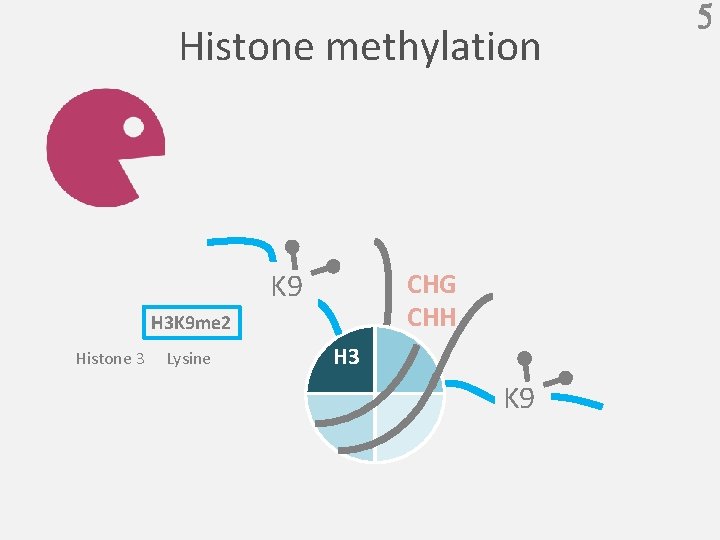
Histone methylation K 9 CHG CHH H 3 K 9 me 2 Histone 3 Lysine H 3 K 9 5
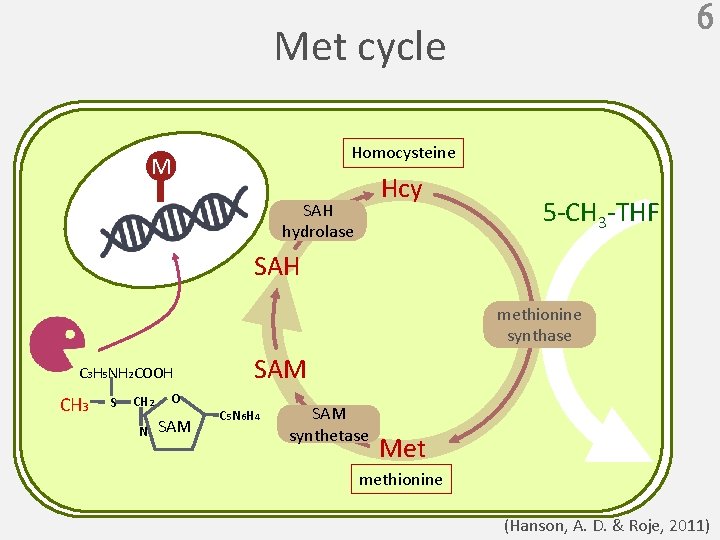
6 Met cycle Homocysteine M Hcy SAH hydrolase 5 -CH 3 -THF SAH methionine synthase C 3 H 5 NH 2 COOH CH 3 S CH 2 O N SAM C 5 N 6 H 4 SAM synthetase Met methionine (Hanson, A. D. & Roje, 2011)
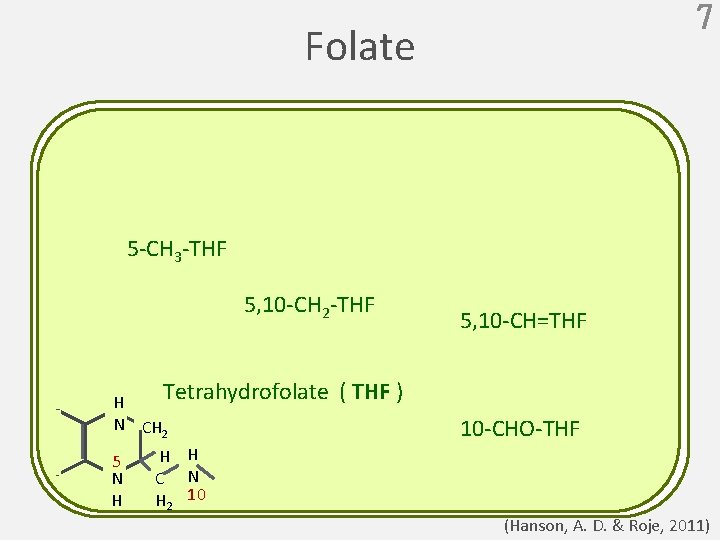
7 Folate 5 -CH 3 -THF 5, 10 -CH 2 -THF 5, 10 -CH=THF Tetrahydrofolate ( THF ) H N CH 2 5 N H 10 -CHO-THF H H C N H 2 10 (Hanson, A. D. & Roje, 2011)
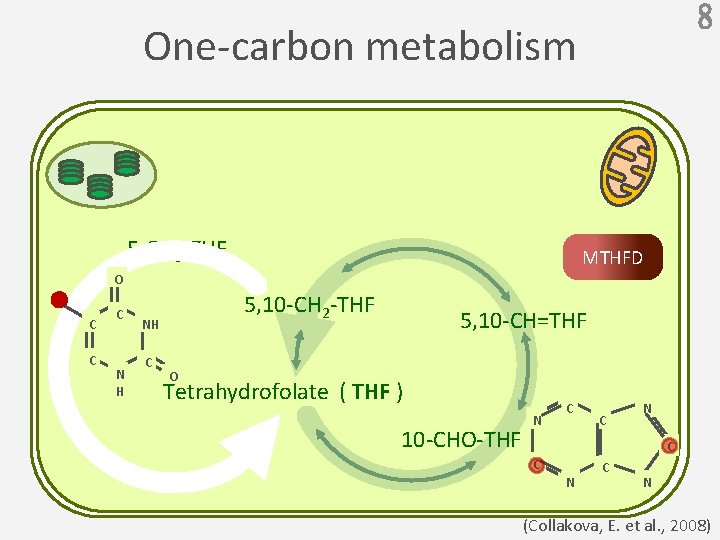
8 One-carbon metabolism 5 -CH 3 -THF MTHFD O C C C N H 5, 10 -CH 2 -THF NH C 5, 10 -CH=THF O Tetrahydrofolate ( THF ) 10 -CHO-THF N C C N C N (Collakova, E. et al. , 2008)

9 MTHFD methylenetetrahydrofolate dehydrogenase D methenyltetrahydrofolate cyclohydrolase C NADP+ 5 -CH 3 -THF NAD(P+) binding catalytic 5, 10 -CH 2 -THF NADPH D 5, 10 -CH=THF C Tetrahydrofolate ( THF ) 10 -CHO-THF MTHFD 1 MTHFD

10 Aim DNA methylation ? M MTHFD 1 Hcy SAH 5 -CH 3 -THF D 5, 10 -CH 2 -THF 5, 10 -CH=THF C Tetrahydrofolate ( THF ) 10 -CHO-THF SAM Met cycle Folate cycle
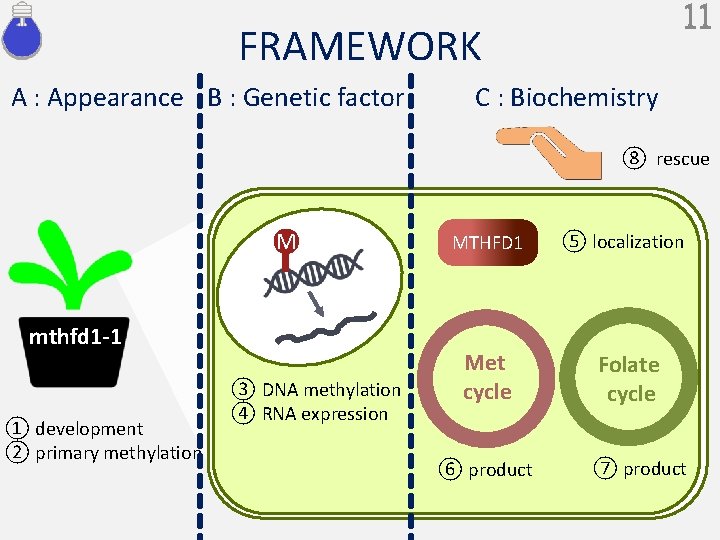
11 FRAMEWORK A : Appearance B : Genetic factor C : Biochemistry ⑧ rescue M mthfd 1 -1 ① development ② primary methylation ③ DNA methylation ④ RNA expression MTHFD 1 Met cycle ⑥ product ⑤ localization Folate cycle ⑦ product

12 MATERIAL

13 Material Kingdom Plantae Phylum Magnoliophyta Class Magnoliopsida Order Brassicales Family Brassicaceae Genus Arabidopsis thaliana WT SUPPRESSOR OF DRM 2 CMT 3 (Moissiard, G. et al. , 2012)

14 RESULT
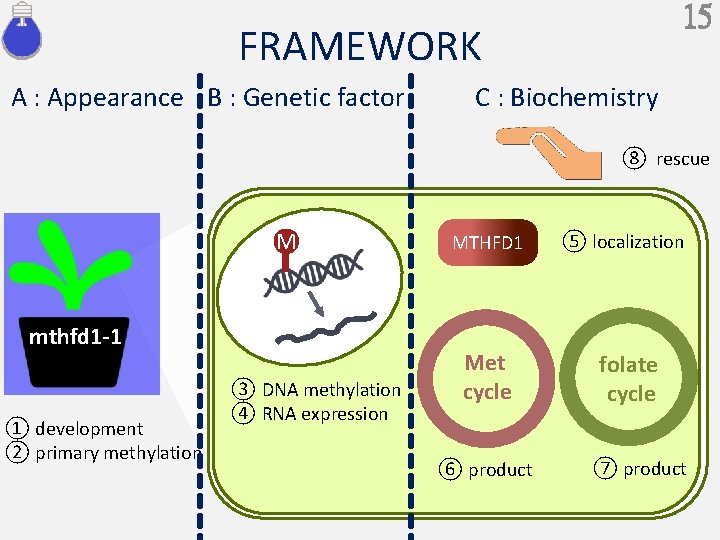
15 FRAMEWORK A : Appearance B : Genetic factor C : Biochemistry ⑧ rescue M mthfd 1 -1 ① development ② primary methylation ③ DNA methylation ④ RNA expression MTHFD 1 Met cycle ⑥ product ⑤ localization folate cycle ⑦ product

16 Mutant mthfd 1 -1 WT F 1 EMS Ler #162 F 2 Sequence/Mapping F 1 (Fig. 1 b)
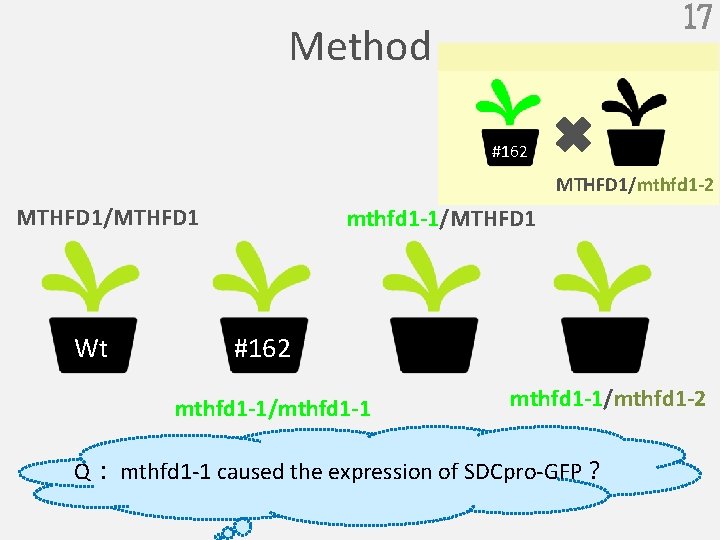
17 Method #162 MTHFD 1/mthfd 1 -2 MTHFD 1/MTHFD 1 Wt mthfd 1 -1/MTHFD 1 #162 mthfd 1 -1/mthfd 1 -1/mthfd 1 -2 Q: mthfd 1 -1 caused the expression of SDCpro-GFP?

18 Expression of SDCpro-GFP +/+ -/MTHFD 1 mthfd 1 -1/mthfd 1 -1 -/+ -/- mthfd 1 -1 caused the expression of SDCpro-GFP in #162 (Fig. 1 a)
19 FRAMEWORK A : Appearance B : Genetic factor C : Biochemistry ⑧ rescue M mthfd 1 -1 ① development ② primary methylation ③ DNA methylation ④ RNA expression MTHFD 1 Met cycle ⑥ product ⑤ localization folate cycle ⑦ product
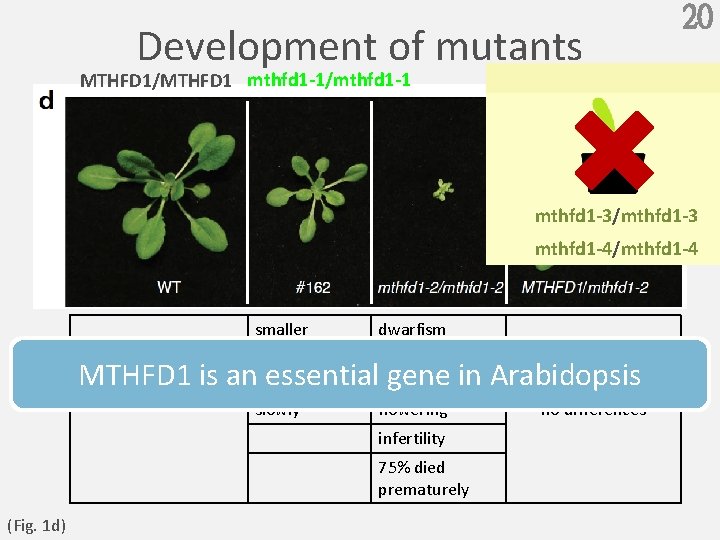
20 Development of mutants MTHFD 1/MTHFD 1 mthfd 1 -1/mthfd 1 -1 10 mm mthfd 1 -3/mthfd 1 -3 mthfd 1 -4/mthfd 1 -4 smaller dwarfism pale MTHFD 1 is an essential gene in Arabidopsis developed delayed -- slowly flowering infertility 75% died prematurely (Fig. 1 d) no differences
21 FRAMEWORK A : Appearance B : Genetic factor C : Biochemistry ⑧ rescue M mthfd 1 -1 √ ① development ② primary methylation ③ DNA methylation ④ RNA expression MTHFD 1 Met cycle ⑥ product ⑤ localization folate cycle ⑦ product
Question 22 WT SUPPRESSOR OF DRM 2 CMT 3 Q:What is happen to DNA methylation?
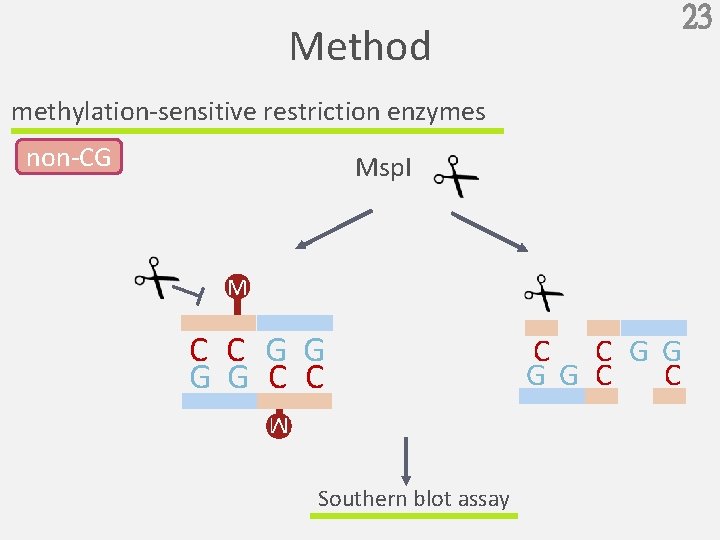
23 Method methylation-sensitive restriction enzymes non-CG Msp. I M C C G G C C M Southern blot assay C C G G C C
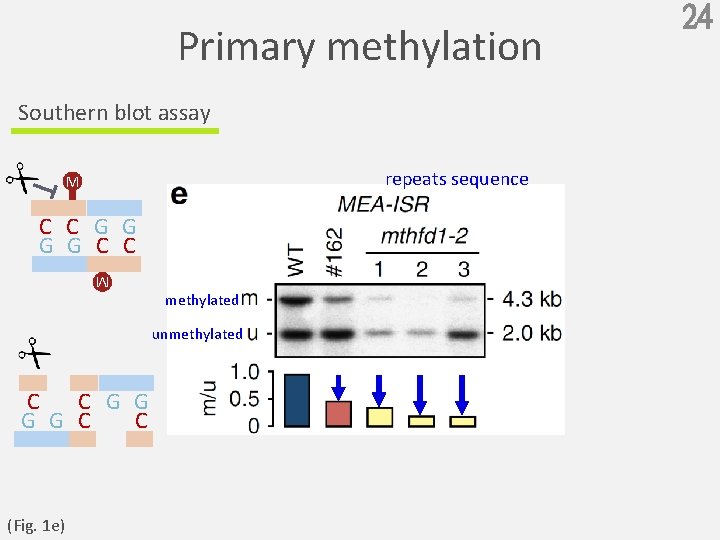
Primary methylation Southern blot assay repeats sequence M C C G G C C methylated M unmethylated C C G G C C (Fig. 1 e) 24
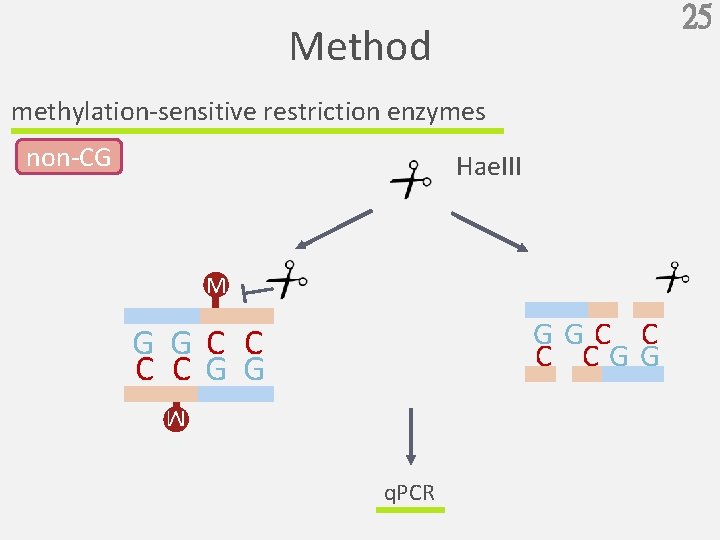
25 Method methylation-sensitive restriction enzymes non-CG Hae. III M GGC C C CGG G GC C C CG G M q. PCR
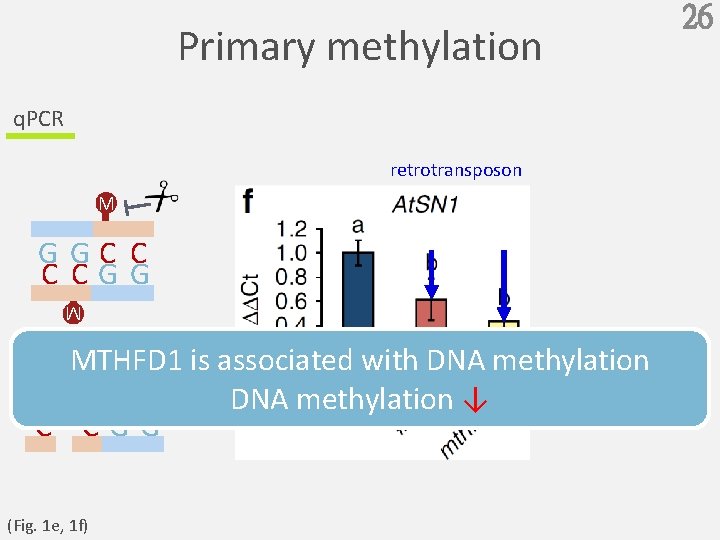
Primary methylation q. PCR retrotransposon M GGC C C CGG M MTHFD 1 is associated with DNA methylation ↓ GGC C C CGG (Fig. 1 e, 1 f) 26
27 FRAMEWORK A : Appearance B : Genetic factor C : Biochemistry ⑧ rescue M mthfd 1 -1 √ ① development √ ② primary methylation ③ DNA methylation ④ RNA expression MTHFD 1 Met cycle ⑥ product ⑤ localization folate cycle ⑦ product
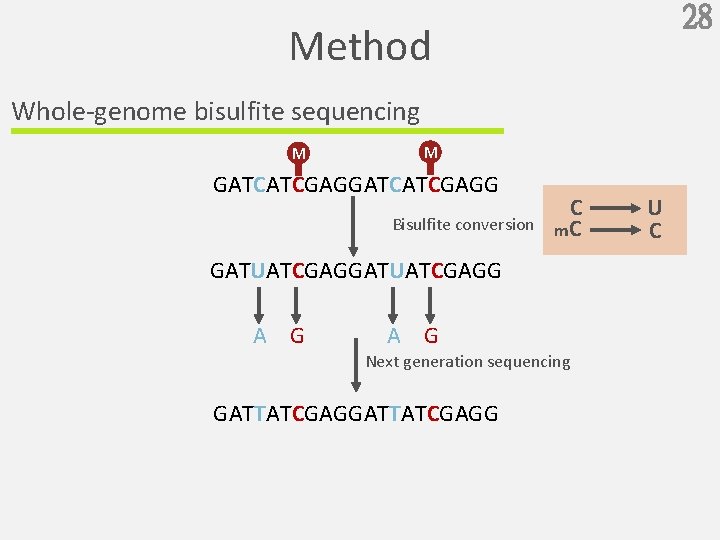
28 Method Whole-genome bisulfite sequencing M M GATCATCGAGG Bisulfite conversion C m. C GATUATCGAGG A G Next generation sequencing GATTATCGAGG U C
FRAMEWORK 29 B : Genetic factor M ? ③ DNA methylation ■ Average global Cs □ □ Chromosomes DNA methylation □ TE & PCG DNA methylation □ CG, CHH methylation

Average global DNA methylation for Cs 24% 40% CHG > CHH > CG (Fig. 2 a) 62% 50% 30
FRAMEWORK 31 B : Genetic factor M ? ③ DNA methylation □ Average global Cs □ ■ Chromosomes DNA methylation □ TE & PCG DNA methylation □ CG, CHH methylation
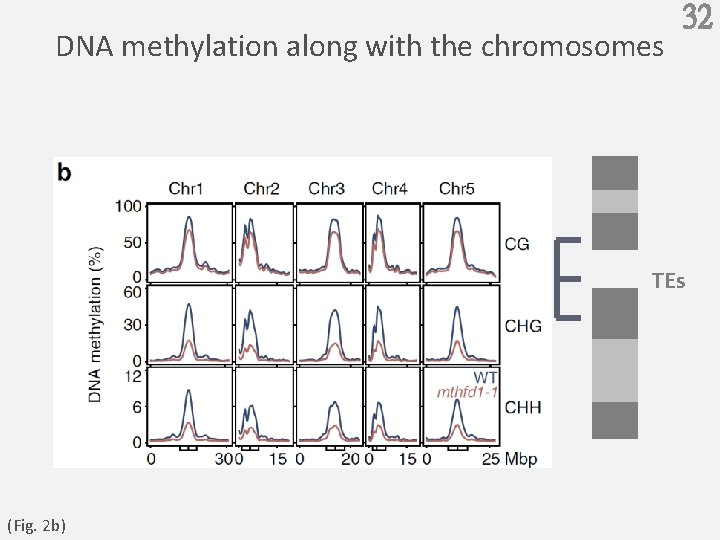
DNA methylation along with the chromosomes 32 TEs (Fig. 2 b)
FRAMEWORK 33 B : Genetic factor M ? ③ DNA methylation □ Average global Cs □ Chromosomes DNA methylation ■ TE & PCG DNA methylation □ □ CG, CHH methylation
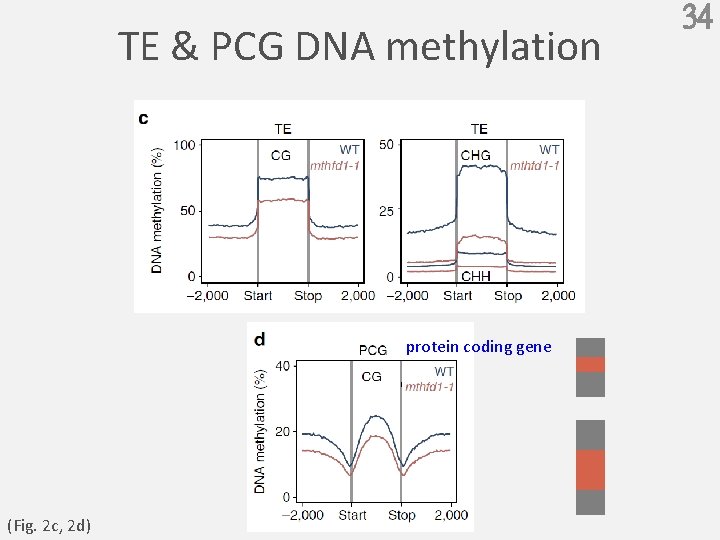
TE & PCG DNA methylation protein coding gene (Fig. 2 c, 2 d) 34
FRAMEWORK 35 B : Genetic factor M ? ③ DNA methylation □ Average global Cs □ Chromosomes DNA methylation □ TE & PCG DNA methylation ■ CG, CHH methylation □
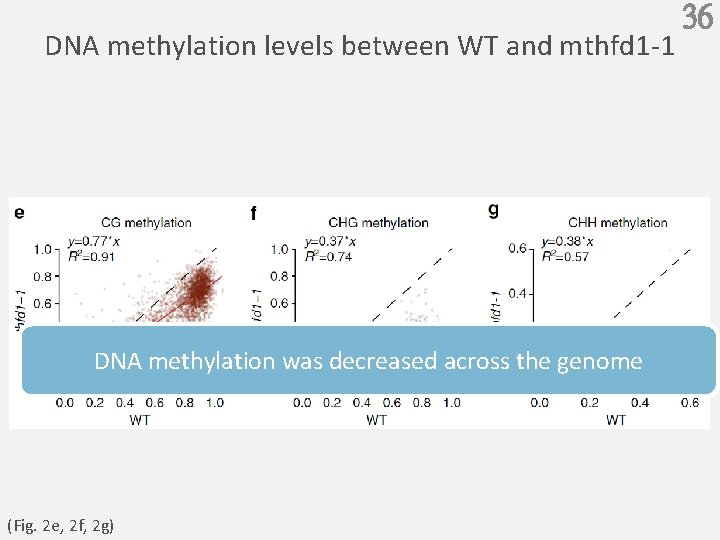
DNA methylation levels between WT and mthfd 1 -1 DNA methylation was decreased across the genome (Fig. 2 e, 2 f, 2 g) 36

Question 37 ( H:A、T、G ) CG CHH met MET cmt 3 CMT 3 cmt 2 CMT 2 drm 1, 2 DRM mthfd 1 -1 Q:mthfd 1 -1 vs DNMT mutant hypo-DMRs differentially methylated regions
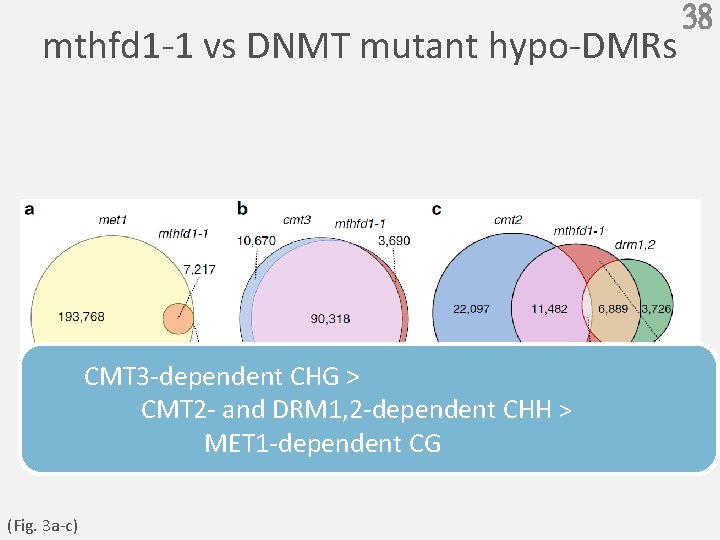
mthfd 1 -1 vs DNMT mutant hypo-DMRs CMT 3 -dependent CHG > CMT 2 - and DRM 1, 2 -dependent CHH > MET 1 -dependent CG (Fig. 3 a-c) 38
39 Question K 9 CHG CHH H 3 K 9 me 2 Histone 3 Lysine H 3 K 9 Q:What is the effect on H 3 K 9 me 2 ?
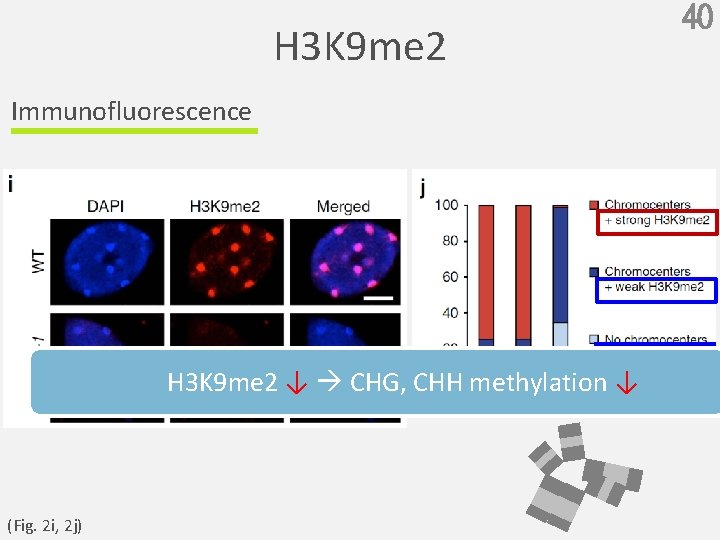
H 3 K 9 me 2 Immunofluorescence H 3 K 9 me 2 ↓ CHG, CHH methylation ↓ (Fig. 2 i, 2 j) 40
41 FRAMEWORK A : Appearance B : Genetic factor C : Biochemistry ⑧ rescue M mthfd 1 -1 √ ① development √ ② primary methylation √ ③ DNA methylation ④ RNA expression MTHFD 1 Met cycle ⑥ product ⑤ localization folate cycle ⑦ product
FRAMEWORK B : Genetic factor M ? ■ retrotransposons expression □ ④ RNA expression □ CHH and CHG hypo-DMRs □ Chromosomes transcript □ Up-and down-regulated gene 42
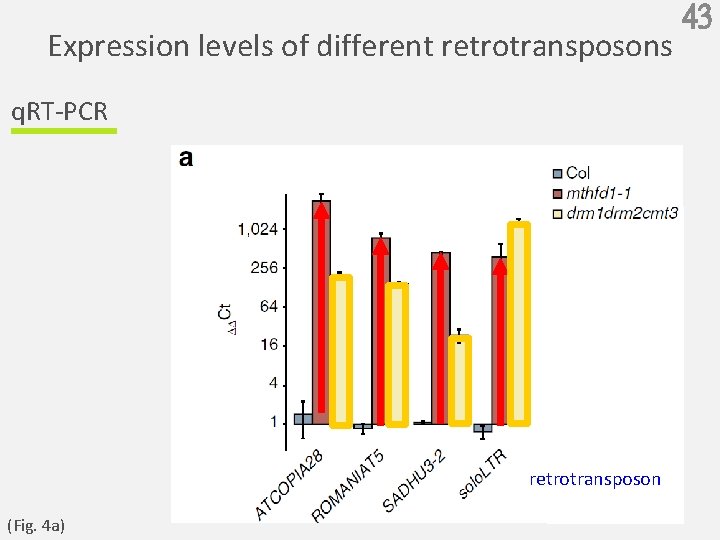
Expression levels of different retrotransposons q. RT-PCR retrotransposon (Fig. 4 a) 43
FRAMEWORK B : Genetic factor M ? □ retrotransposons expression ④ RNA expression □ CHH and CHG hypo-DMRs ■ □ Chromosomes transcript □ Up-and down-regulated gene 44
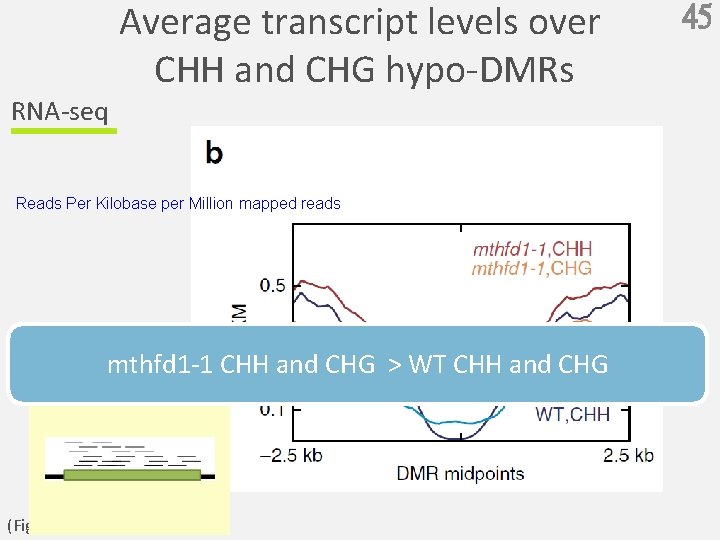
RNA-seq Average transcript levels over CHH and CHG hypo-DMRs Reads Per Kilobase per Million mapped reads mthfd 1 -1 CHH and CHG > WT CHH and CHG (Fig. 4 b) 45
FRAMEWORK B : Genetic factor M ? □ retrotransposons expression ④ RNA expression □ CHH and CHG hypo-DMRs ■ Chromosomes transcript □ □ Up-and down-regulated gene 46
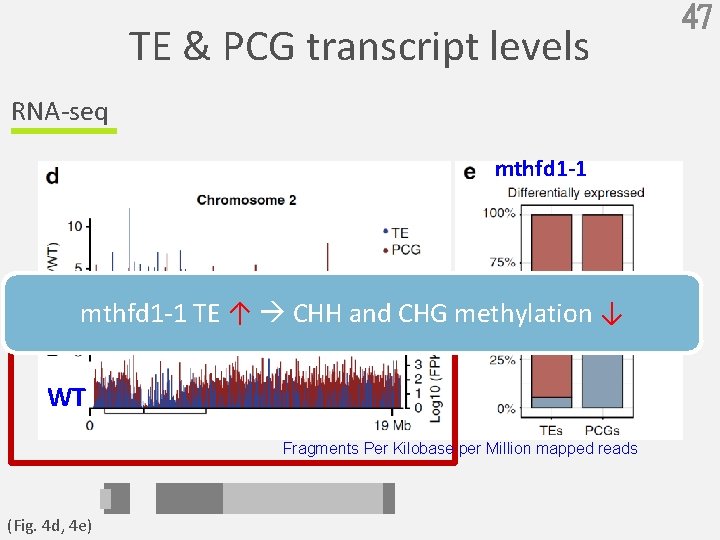
TE & PCG transcript levels RNA-seq mthfd 1 -1 TE ↑ CHH and CHG methylation ↓ WT Fragments Per Kilobase per Million mapped reads (Fig. 4 d, 4 e) 47
FRAMEWORK B : Genetic factor M ? □ retrotransposons expression ④ RNA expression □ CHH and CHG hypo-DMRs □ Chromosomes transcript ■ Up-and down-regulated gene □ 48
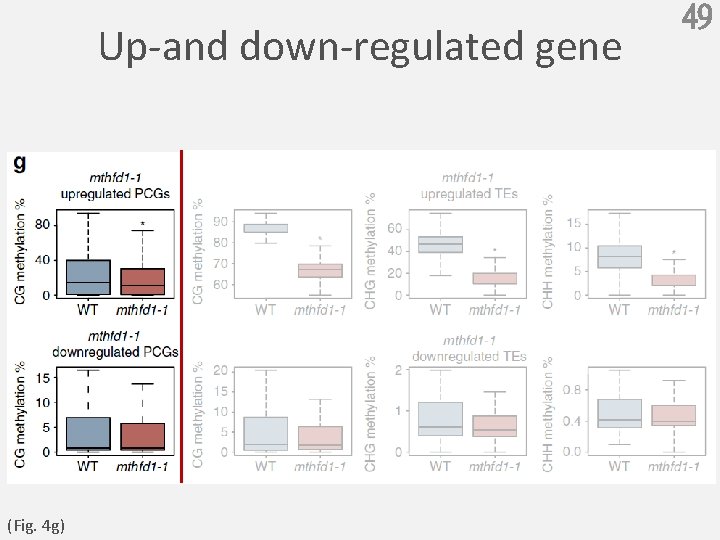
Up-and down-regulated gene (Fig. 4 g) 49
50 FRAMEWORK A : Appearance B : Genetic factor C : Biochemistry ⑧ rescue M mthfd 1 -1 √ ① development √ ② primary methylation √ ③ DNA methylation √ ④ RNA expression MTHFD 1 Met cycle ⑥ product ⑤ localization folate cycle ⑦ product
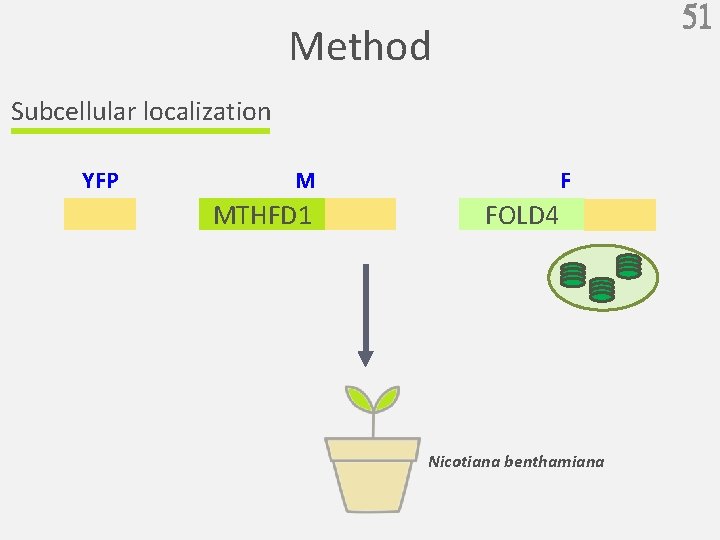
51 Method Subcellular localization YFP F M MTHFD 1 FOLD 4 Nicotiana benthamiana

MTHFD 1 localization Subcellular localization MTHFD 1 is in the cytoplasm FOLD 4 (Fig. 6 a, 6 b) 52
53 FRAMEWORK A : Appearance B : Genetic factor C : Biochemistry ⑧ rescue M mthfd 1 -1 √ ① development √ ② primary methylation √ ③ DNA methylation √ ④ RNA expression MTHFD 1 Met cycle ⑥ product √ ⑤ localization folate cycle ⑦ product

The levels of SAM and SAH HPLC SAM MI = SAH (Fig. 7 a, b, c ) 54
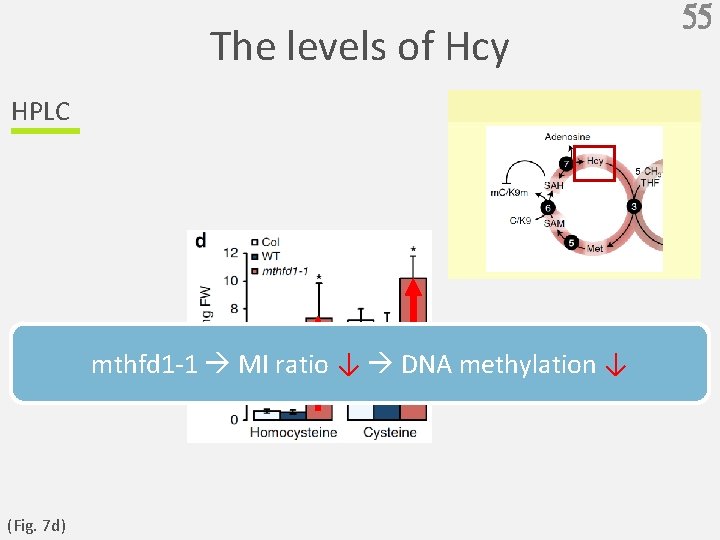
The levels of Hcy HPLC mthfd 1 -1 MI ratio ↓ DNA methylation ↓ (Fig. 7 d) 55
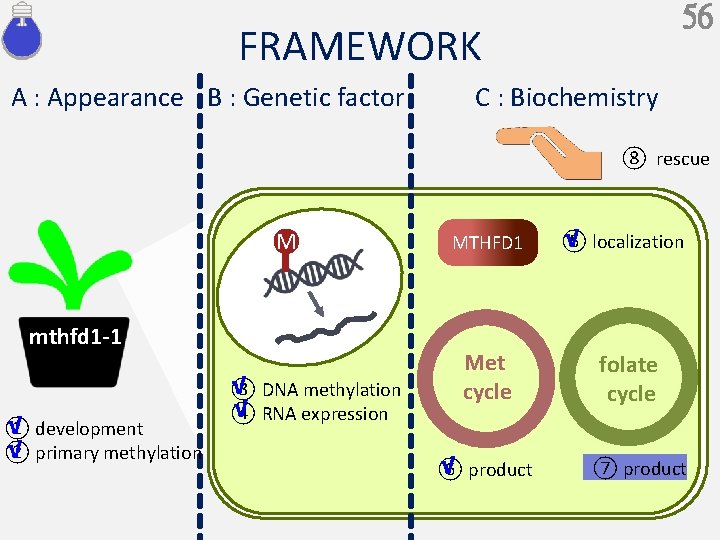
56 FRAMEWORK A : Appearance B : Genetic factor C : Biochemistry ⑧ rescue M mthfd 1 -1 √ ① development √ ② primary methylation √ ③ DNA methylation √ ④ RNA expression MTHFD 1 Met cycle √ ⑥ product √ ⑤ localization folate cycle ⑦ product
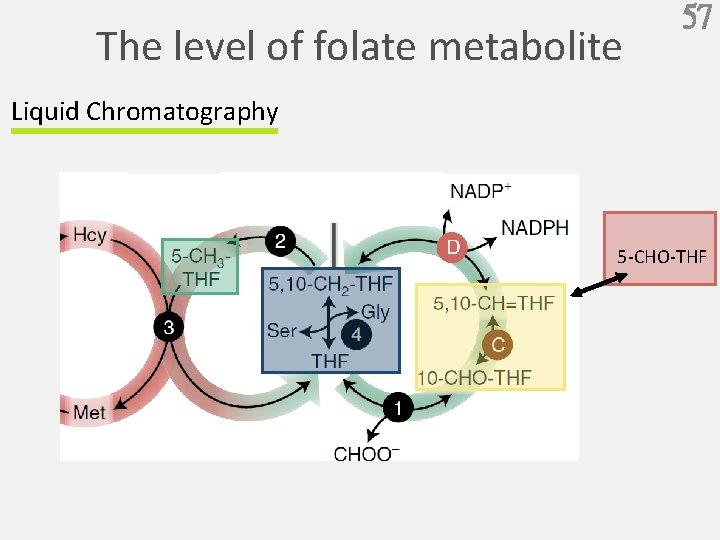
The level of folate metabolite 57 Liquid Chromatography 5 -CHO-THF
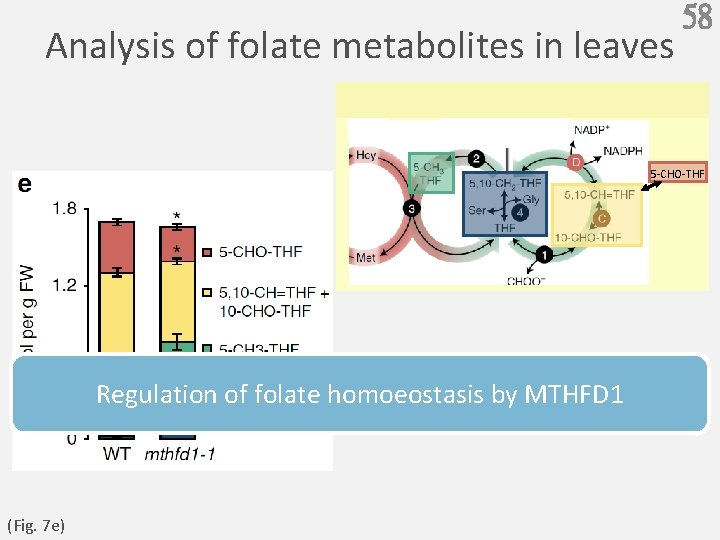
Analysis of folate metabolites in leaves 58 5 -CHO-THF Regulation of folate homoeostasis by MTHFD 1 (Fig. 7 e)
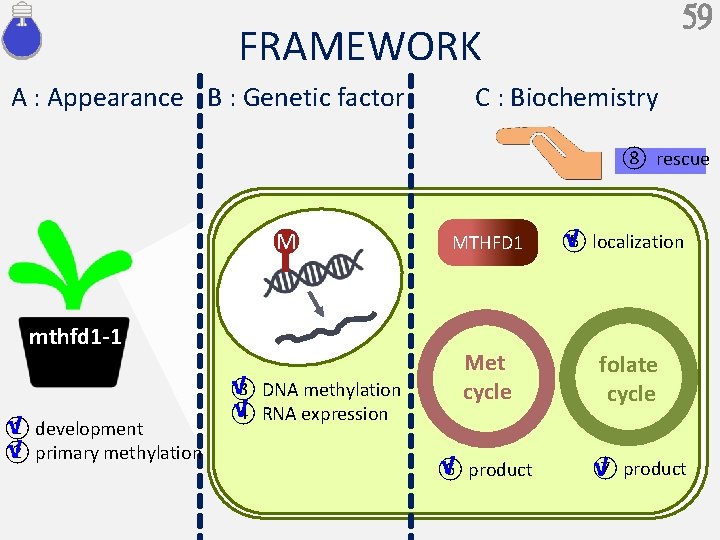
59 FRAMEWORK A : Appearance B : Genetic factor C : Biochemistry ⑧ rescue M mthfd 1 -1 √ ① development √ ② primary methylation √ ③ DNA methylation √ ④ RNA expression MTHFD 1 Met cycle √ ⑥ product √ ⑤ localization folate cycle ⑦ product √
FRAMEWORK C : Biochemistry ⑧ rescue ■ Root growth □ □ DNA methylation 60
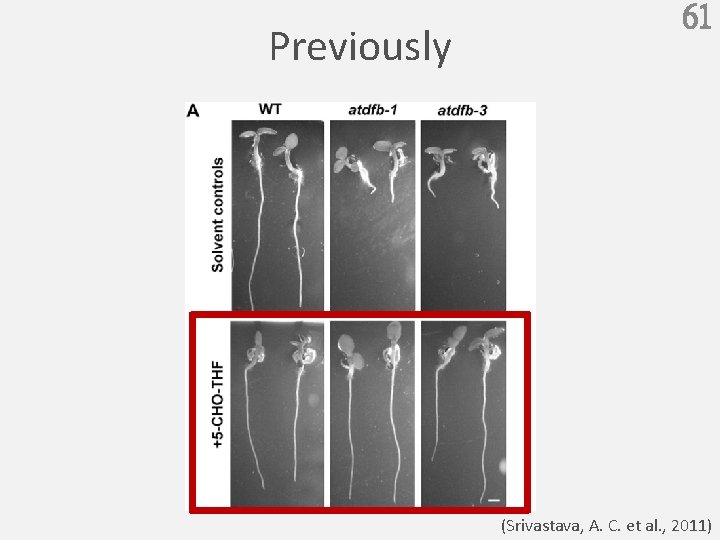
Previously 61 (Srivastava, A. C. et al. , 2011)
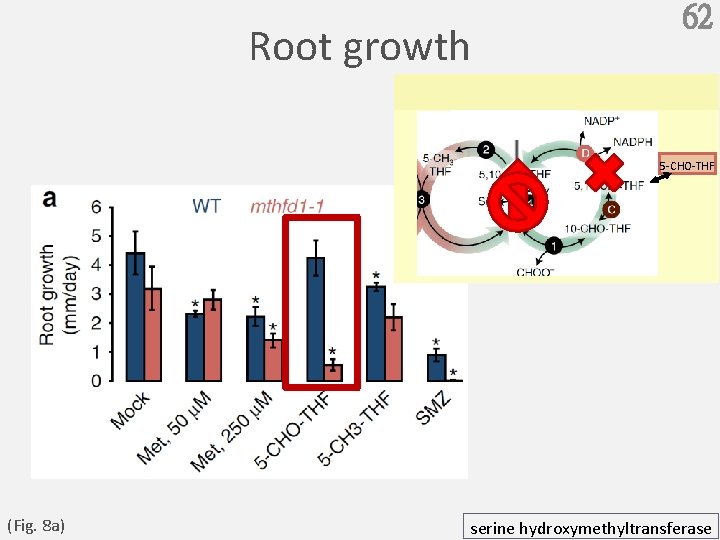
Root growth 62 5 -CHO-THF (Fig. 8 a) serine hydroxymethyltransferase
FRAMEWORK C : Biochemistry ⑧ rescue □ Root growth □ ■ DNA methylation 63
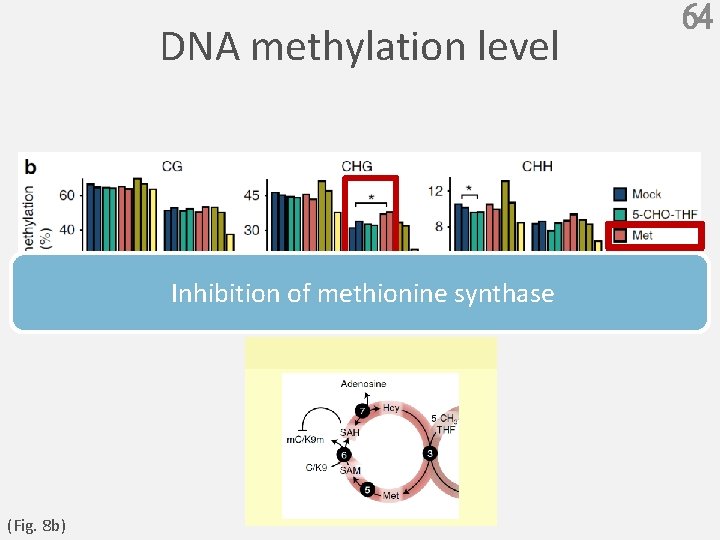
DNA methylation level Inhibition of methionine synthase (Fig. 8 b) 64
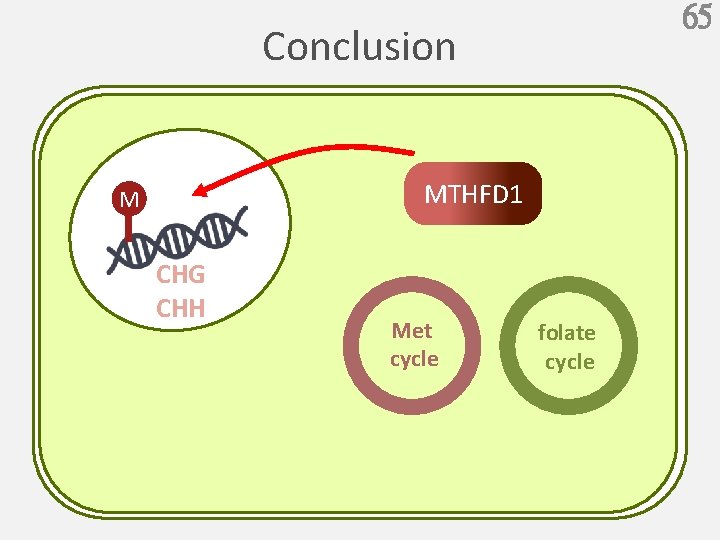
65 Conclusion MTHFD 1 M CHG CHH Met cycle folate cycle

66 ~~ Thanks for listening ~~

67

One-carbon metabolism 68 (Hanson, A. D. & Roje, S. , 2001)

RPKM & FPKM 69
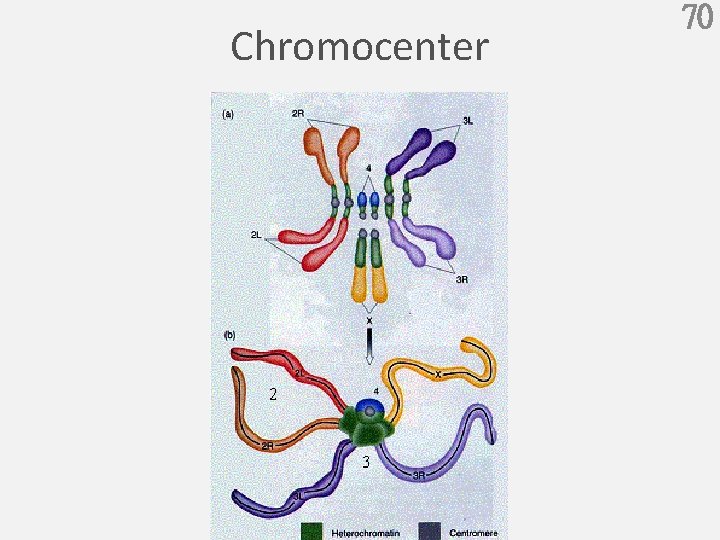
Chromocenter 70
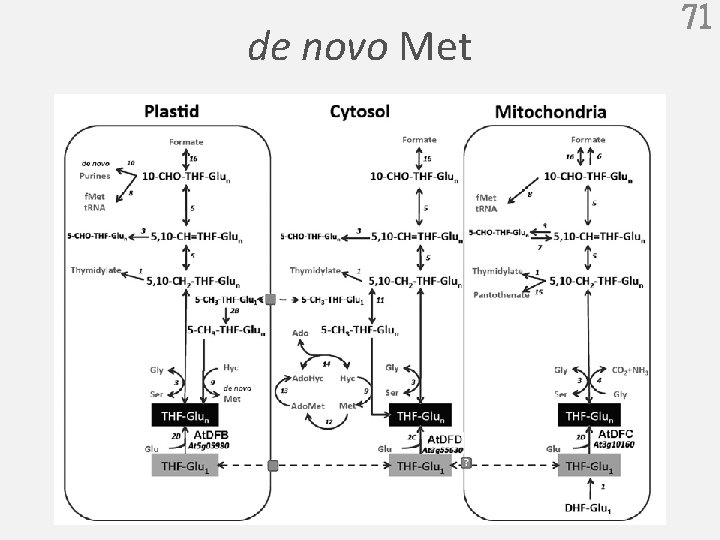
de novo Met 71
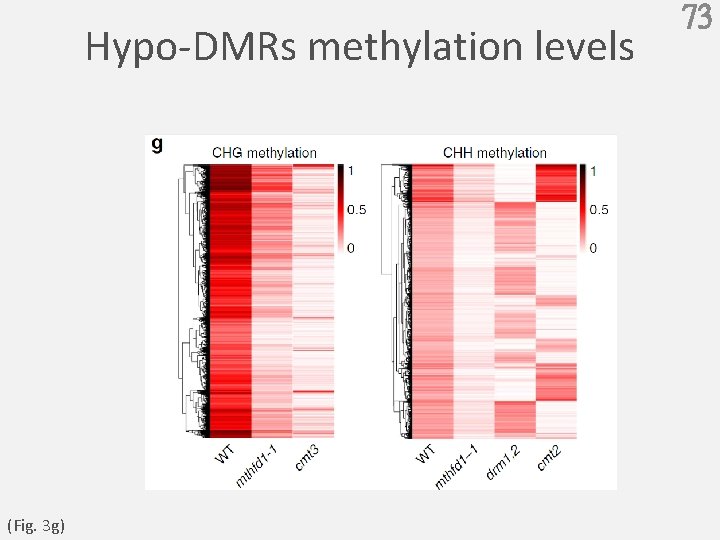
Hypo-DMRs methylation levels (Fig. 3 g) 73
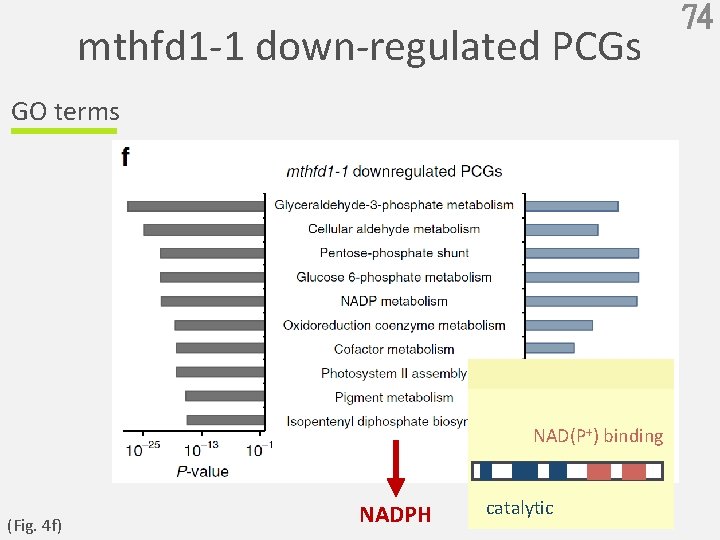
mthfd 1 -1 down-regulated PCGs GO terms NAD(P+) binding (Fig. 4 f) NADPH catalytic 74
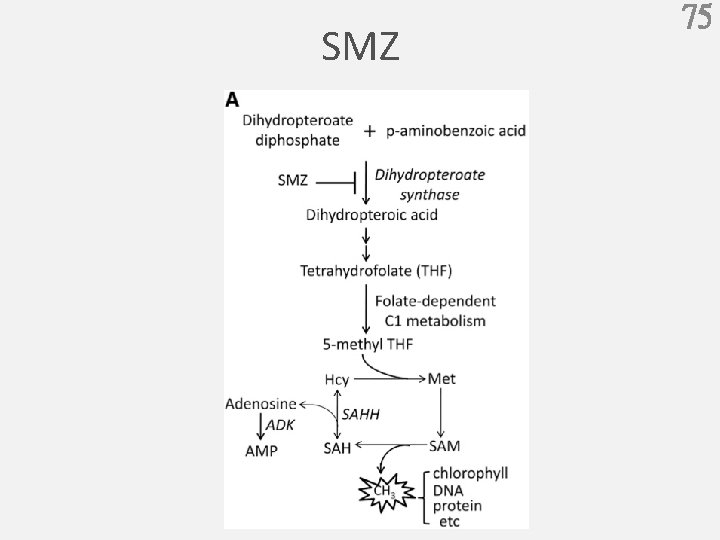
SMZ 75

78 • FPGS 1 = atdfb
- Slides: 75