1 Measuring gene expression via RNA Northern blot

1 Measuring gene expression via RNA • • • Northern blot RNase protection Primer extension • • RT-PCR Q-RT-PCR • • Microarray RNAseq
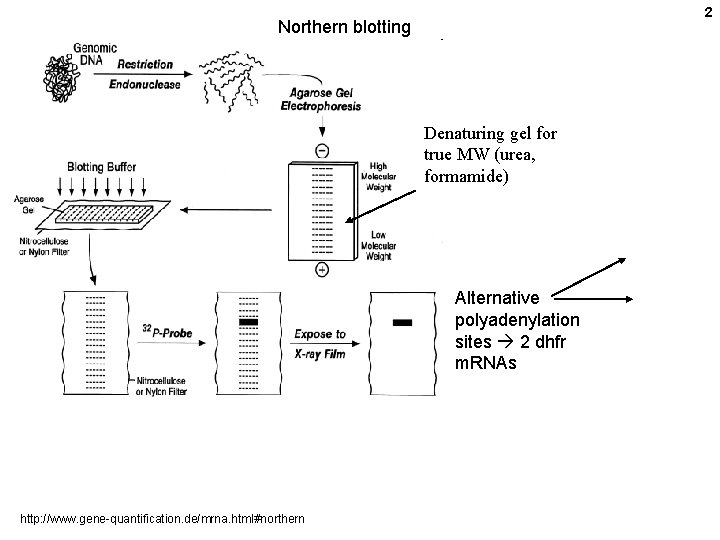
2 Northern blotting Denaturing gel for true MW (urea, formamide) Alternative polyadenylation sites 2 dhfr m. RNAs http: //www. gene-quantification. de/mrna. html#northern
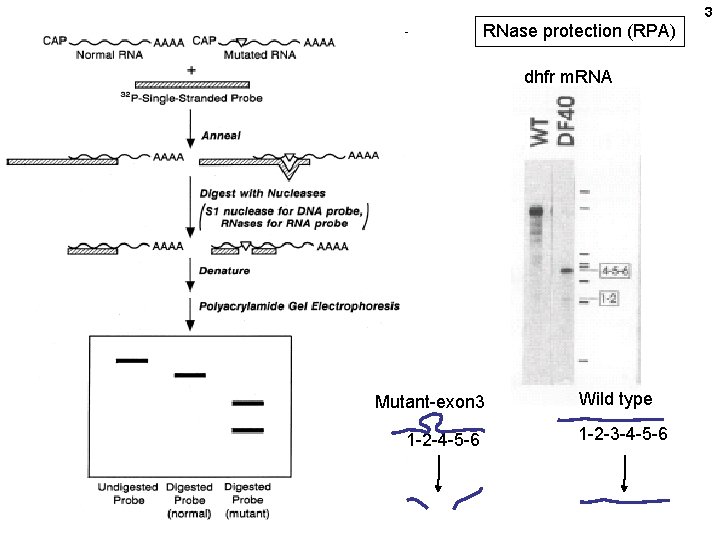
3 RNase protection (RPA) dhfr m. RNA Mutant-exon 3 1 -2 -4 -5 -6 Wild type 1 -2 -3 -4 -5 -6
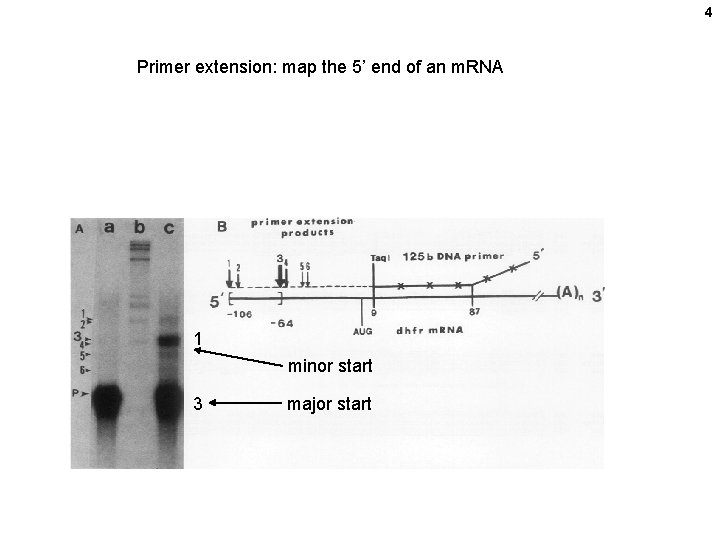
4 Primer extension: map the 5’ end of an m. RNA 1 minor start 3 major start
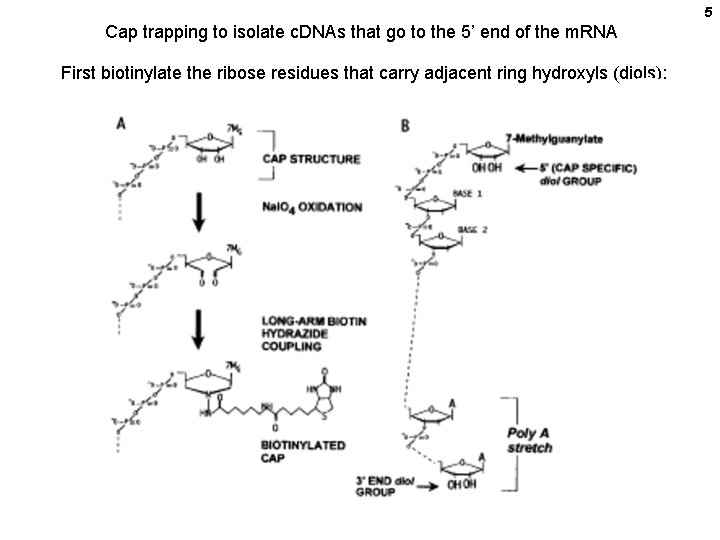
5 Cap trapping to isolate c. DNAs that go to the 5’ end of the m. RNA First biotinylate the ribose residues that carry adjacent ring hydroxyls (diols):
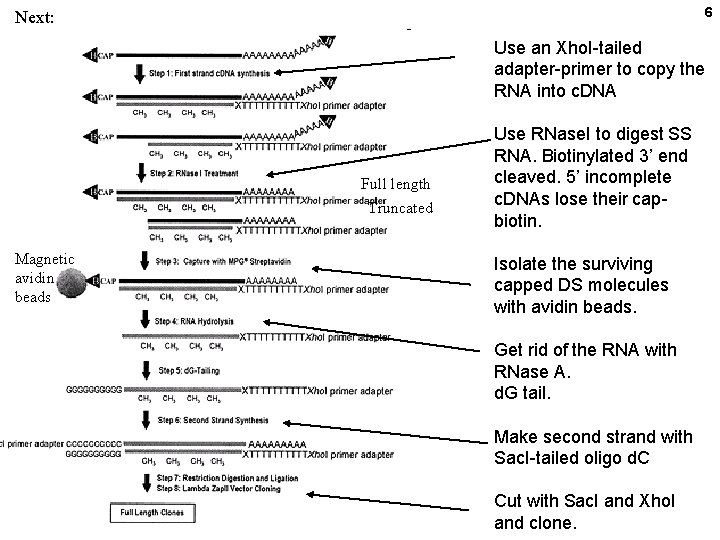
6 Next: Use an Xho. I-tailed adapter-primer to copy the RNA into c. DNA Full length Truncated Magnetic avidin beads Use RNase. I to digest SS RNA. Biotinylated 3’ end cleaved. 5’ incomplete c. DNAs lose their capbiotin. Isolate the surviving capped DS molecules with avidin beads. Get rid of the RNA with RNase A. d. G tail. Make second strand with Sac. I-tailed oligo d. C Cut with Sac. I and Xho. I and clone.
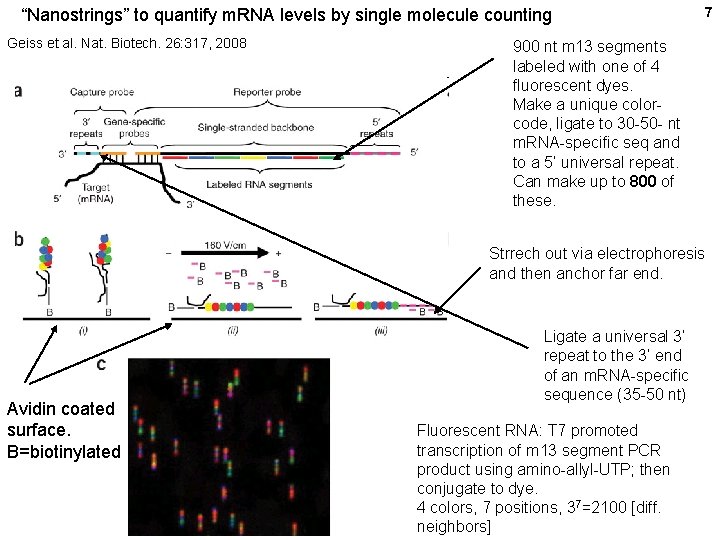
“Nanostrings” to quantify m. RNA levels by single molecule counting Geiss et al. Nat. Biotech. 26: 317, 2008 7 900 nt m 13 segments labeled with one of 4 fluorescent dyes. Make a unique colorcode, ligate to 30 -50 - nt m. RNA-specific seq and to a 5’ universal repeat. Can make up to 800 of these. Strrech out via electrophoresis and then anchor far end. Avidin coated surface. B=biotinylated Ligate a universal 3’ repeat to the 3’ end of an m. RNA-specific sequence (35 -50 nt) Fluorescent RNA: T 7 promoted transcription of m 13 segment PCR product using amino-allyl-UTP; then conjugate to dye. 4 colors, 7 positions, 37=2100 [diff. neighbors]
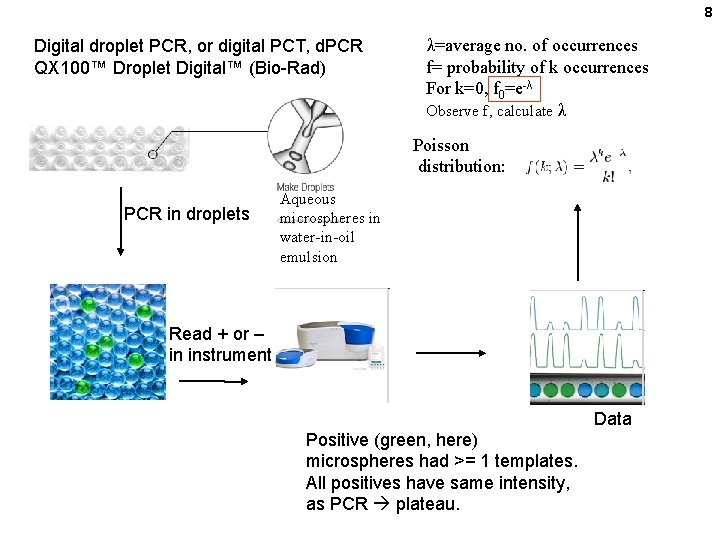
8 Digital droplet PCR, or digital PCT, d. PCR QX 100™ Droplet Digital™ (Bio-Rad) λ=average no. of occurrences f= probability of k occurrences For k=0, f 0=e-λ Observe f, calculate λ Poisson distribution: PCR in droplets Aqueous microspheres in water-in-oil emulsion Read + or – in instrument Data Positive (green, here) microspheres had >= 1 templates. All positives have same intensity, as PCR plateau.
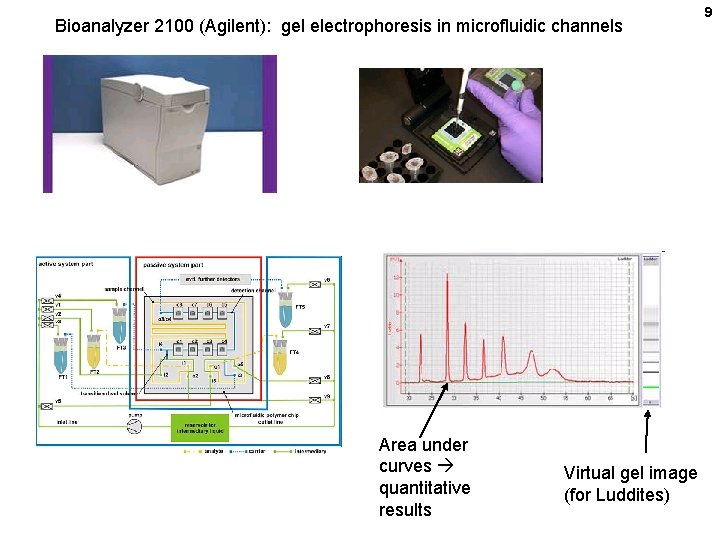
Bioanalyzer 2100 (Agilent): gel electrophoresis in microfluidic channels Area under curves quantitative results Virtual gel image (for Luddites) 9

10 Protein-protein interactions Yeast 2 -hybrid system Yeast 3 -hybrid and 1 hybrid systems Co-immunoprecipitation Pull-downs Far western blots Biacore (surface plasmon resonance, SPR) Fragment complementation
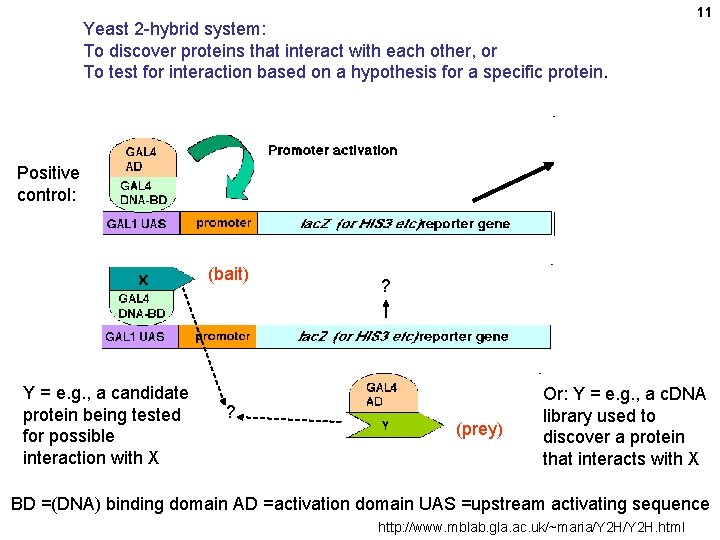
Yeast 2 -hybrid system: To discover proteins that interact with each other, or To test for interaction based on a hypothesis for a specific protein. 11 Positive control: (bait) Y = e. g. , a candidate protein being tested for possible interaction with X ? ? (prey) Or: Y = e. g. , a c. DNA library used to discover a protein that interacts with X BD =(DNA) binding domain AD =activation domain UAS =upstream activating sequence http: //www. mblab. gla. ac. uk/~maria/Y 2 H. html
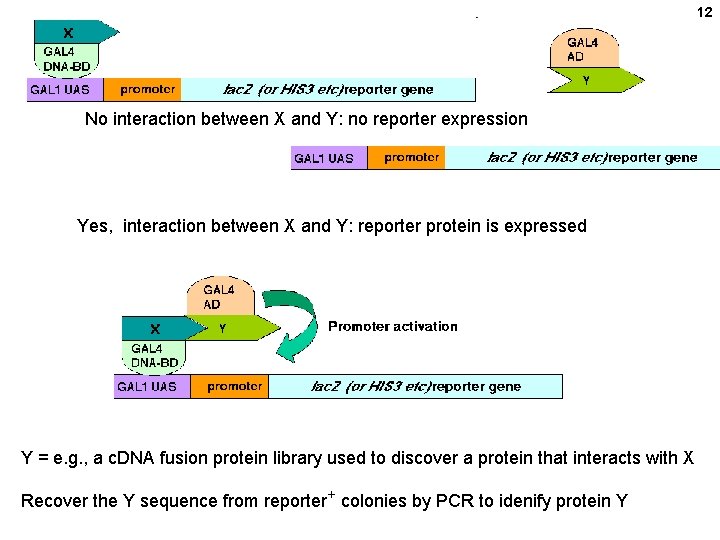
12 No interaction between X and Y: no reporter expression Yes, interaction between X and Y: reporter protein is expressed Y = e. g. , a c. DNA fusion protein library used to discover a protein that interacts with X Recover the Y sequence from reporter+ colonies by PCR to idenify protein Y
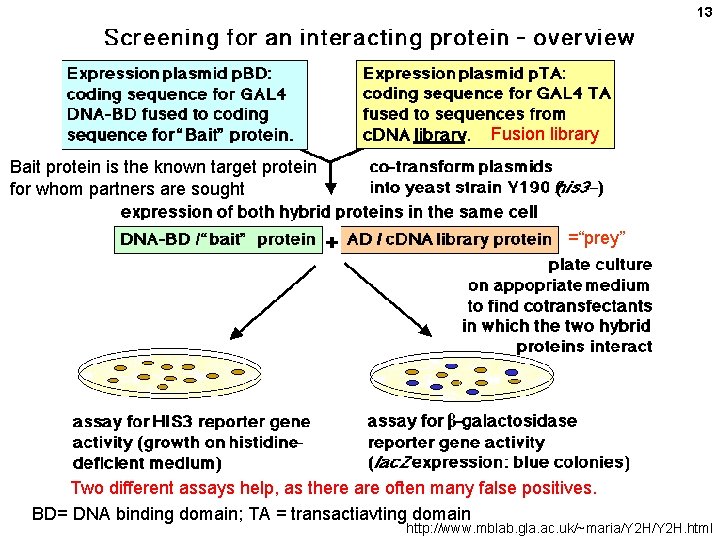
13 Fusion library Bait protein is the known target protein for whom partners are sought =“prey” Two different assays help, as there are often many false positives. BD= DNA binding domain; TA = transactiavting domain http: //www. mblab. gla. ac. uk/~maria/Y 2 H. html
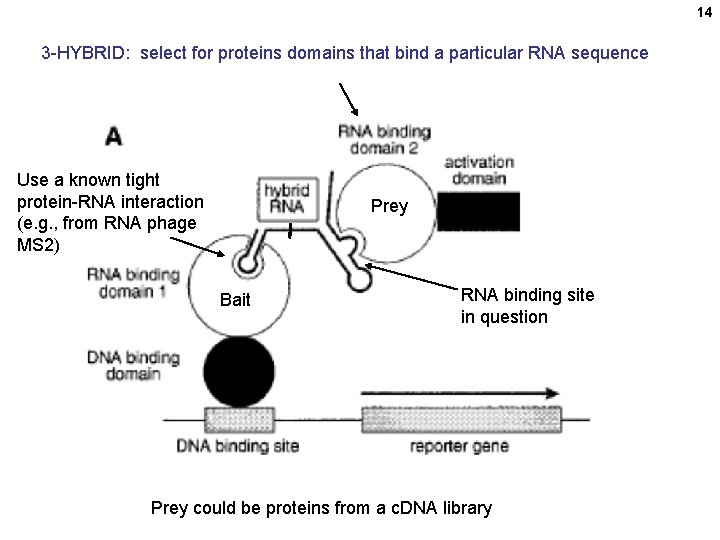
14 3 -HYBRID: select for proteins domains that bind a particular RNA sequence Use a known tight protein-RNA interaction (e. g. , from RNA phage MS 2) Prey Bait RNA binding site in question Prey could be proteins from a c. DNA library
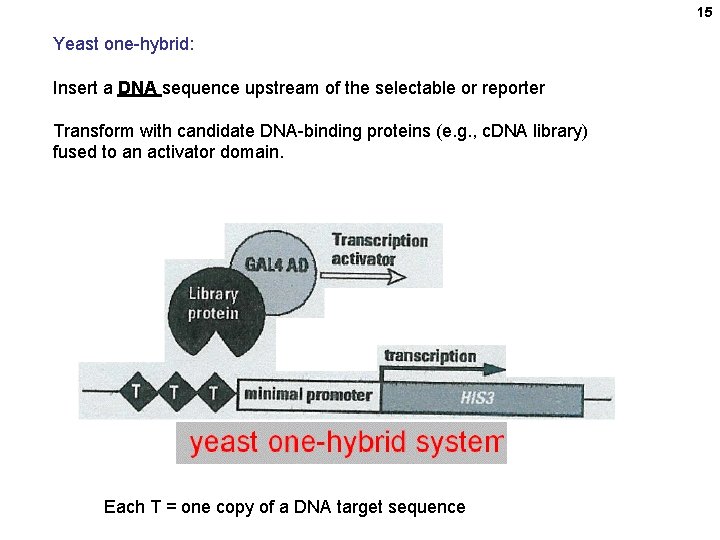
15 Yeast one-hybrid: Insert a DNA sequence upstream of the selectable or reporter Transform with candidate DNA-binding proteins (e. g. , c. DNA library) fused to an activator domain. Each T = one copy of a DNA target sequence
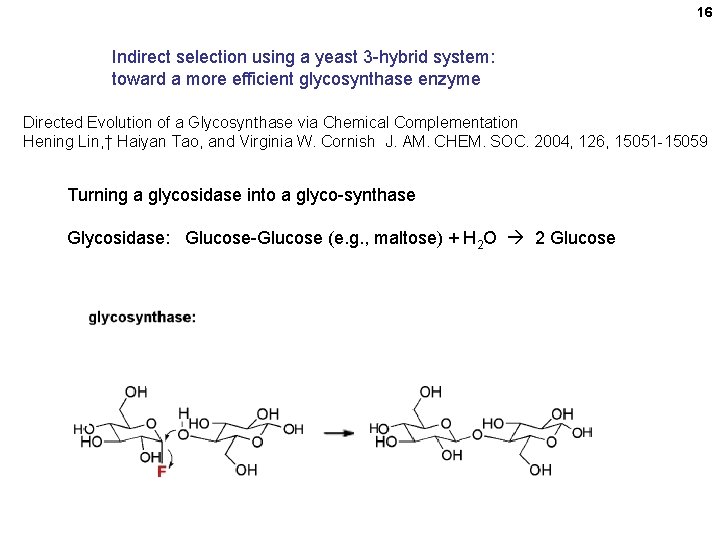
16 Indirect selection using a yeast 3 -hybrid system: toward a more efficient glycosynthase enzyme Directed Evolution of a Glycosynthase via Chemical Complementation Hening Lin, † Haiyan Tao, and Virginia W. Cornish J. AM. CHEM. SOC. 2004, 126, 15051 -15059 Turning a glycosidase into a glyco-synthase Glycosidase: Glucose-Glucose (e. g. , maltose) + H 2 O 2 Glucose
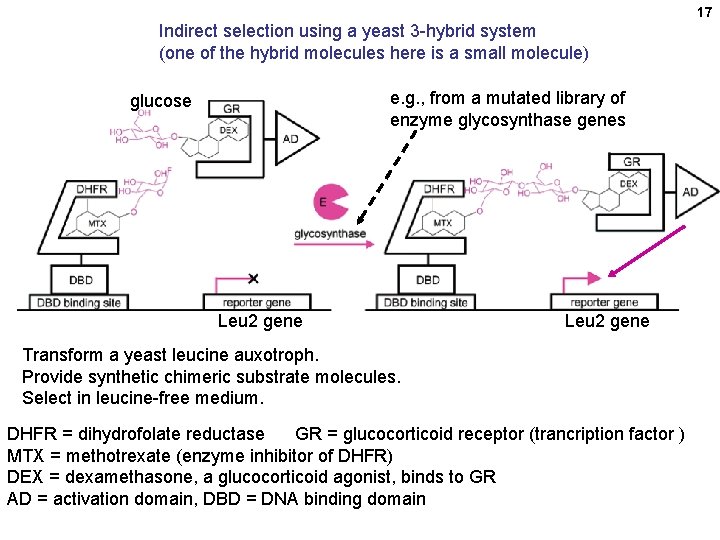
17 Indirect selection using a yeast 3 -hybrid system (one of the hybrid molecules here is a small molecule) e. g. , from a mutated library of enzyme glycosynthase genes glucose Leu 2 gene Transform a yeast leucine auxotroph. Provide synthetic chimeric substrate molecules. Select in leucine-free medium. DHFR = dihydrofolate reductase GR = glucocorticoid receptor (trancription factor ) MTX = methotrexate (enzyme inhibitor of DHFR) DEX = dexamethasone, a glucocorticoid agonist, binds to GR AD = activation domain, DBD = DNA binding domain
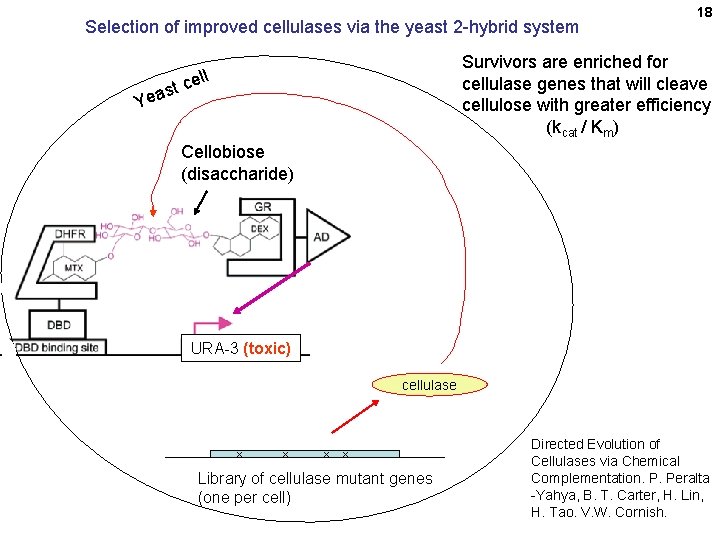
Selection of improved cellulases via the yeast 2 -hybrid system 18 Survivors are enriched for cellulase genes that will cleave cellulose with greater efficiency (kcat / Km) ll ce t s Yea Cellobiose (disaccharide) URA-3 (toxic) cellulase x x Library of cellulase mutant genes (one per cell) Directed Evolution of Cellulases via Chemical Complementation. P. Peralta -Yahya, B. T. Carter, H. Lin, H. Tao. V. W. Cornish.
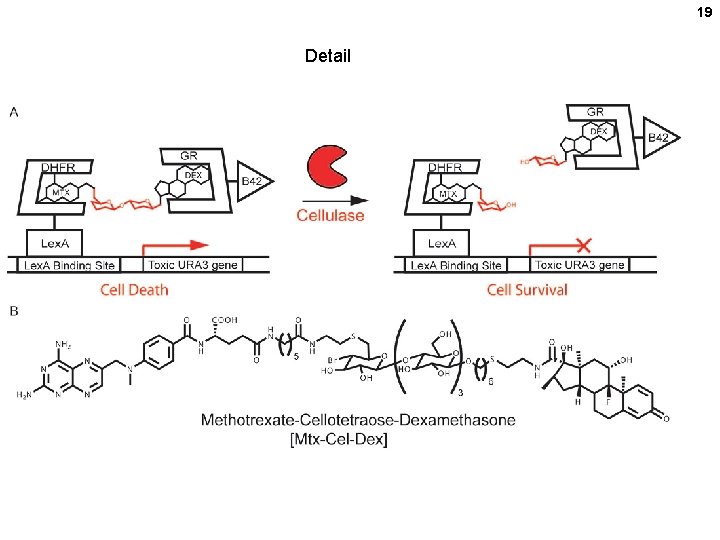
19 Detail
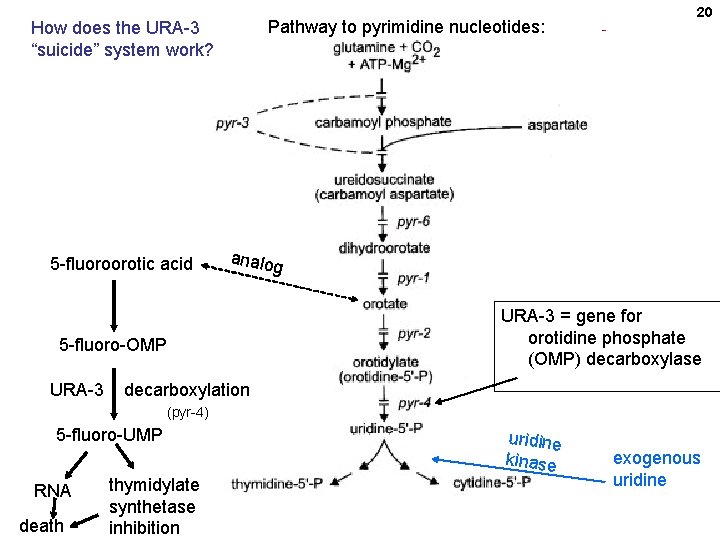
Pathway to pyrimidine nucleotides: How does the URA-3 “suicide” system work? 5 -fluoroorotic acid analo g URA-3 = gene for orotidine phosphate (OMP) decarboxylase 5 -fluoro-OMP URA-3 20 decarboxylation (pyr-4) 5 -fluoro-UMP RNA death thymidylate synthetase inhibition uridine kinase exogenous uridine

21 Measuring protein-protein interactions in vitro X=one protein Y= another protein Pull-downs: Binding between defined purified proteins, at least one being purified. Tag each protein differently by making the appropriate c. DNA clone. Examples: His 6 -X + HA-Y; Bind to nickel ion column via X, elute (his), Western with HA Ab for Y GST-X + HA-Y; Bind to glutathione ion column, elute (glutathione), Western with HA Ab His 6 -X + 35 S-Y (made in vitro); Bind Ni column, elute (his), gel + autoradiography. No antibody needed. GST = glutathione-S-transferase (HA = flu hemagglutinin) glutathione = gamma-glutamyl-cysteinyl-glycine.
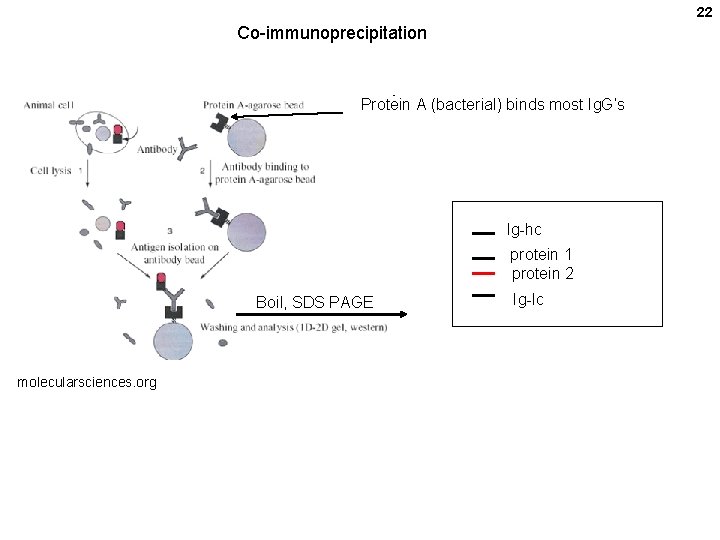
22 Co-immunoprecipitation Protein A (bacterial) binds most Ig. G’s Ig-hc protein 1 protein 2 Boil, SDS PAGE molecularsciences. org Ig-lc

23 Without a specific antibody for your target protein, express it as a fusion protein carrying a standard epitope tag such as FLAG, myc, hemagglutinin (HA), etc. Or, do a pulldown, without antibodies using pairs of strongly interacting molecules: His 6 + chelated nickel or cobalt on beads glutathione-s-tranferase (a small enzyme, GST) + Glutathione (tetrapeptide) on beads maltose binding protein (MBP) + amylose (poly-alpha-glucose) on beads
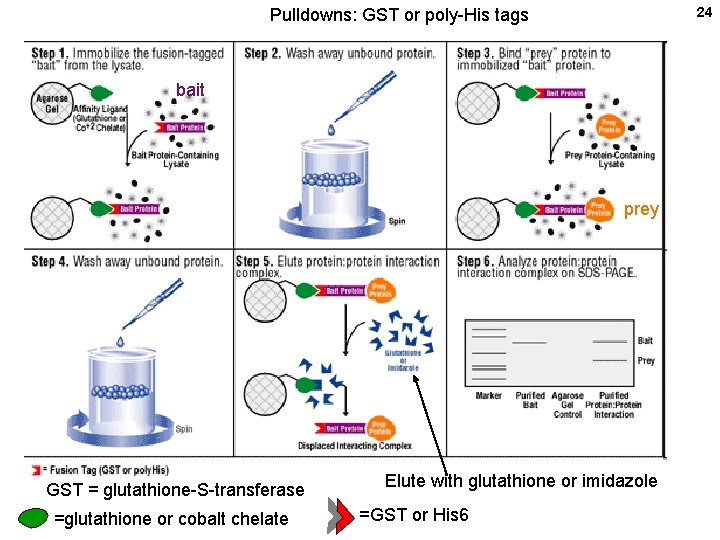
24 Pulldowns: GST or poly-His tags bait prey GST = glutathione-S-transferase =glutathione or cobalt chelate Elute with glutathione or imidazole =GST or His 6
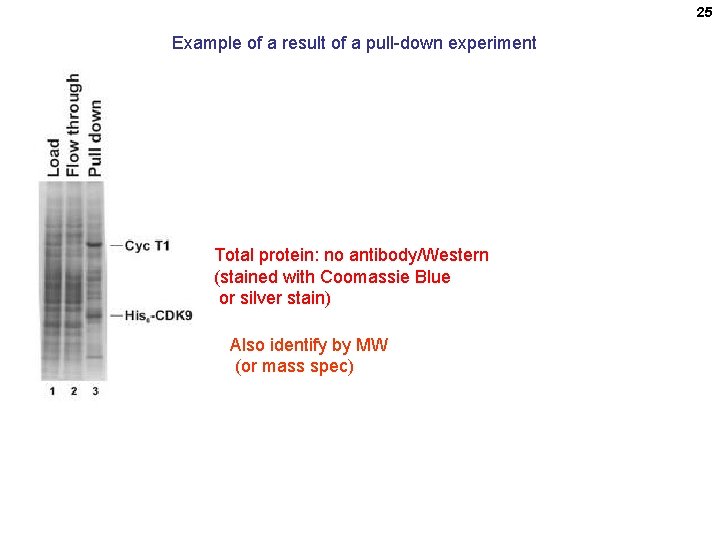
25 Example of a result of a pull-down experiment Total protein: no antibody/Western (stained with Coomassie Blue or silver stain) Also identify by MW (or mass spec)
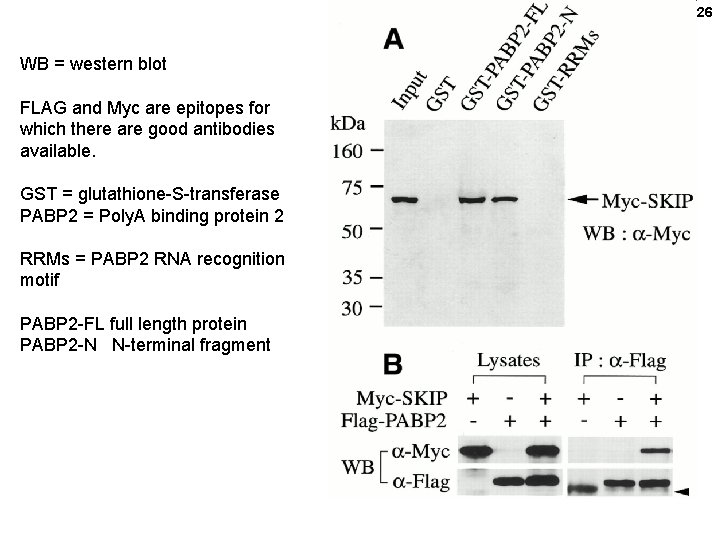
26 WB = western blot FLAG and Myc are epitopes for which there are good antibodies available. GST = glutathione-S-transferase PABP 2 = Poly. A binding protein 2 RRMs = PABP 2 RNA recognition motif PABP 2 -FL full length protein PABP 2 -N N-terminal fragment
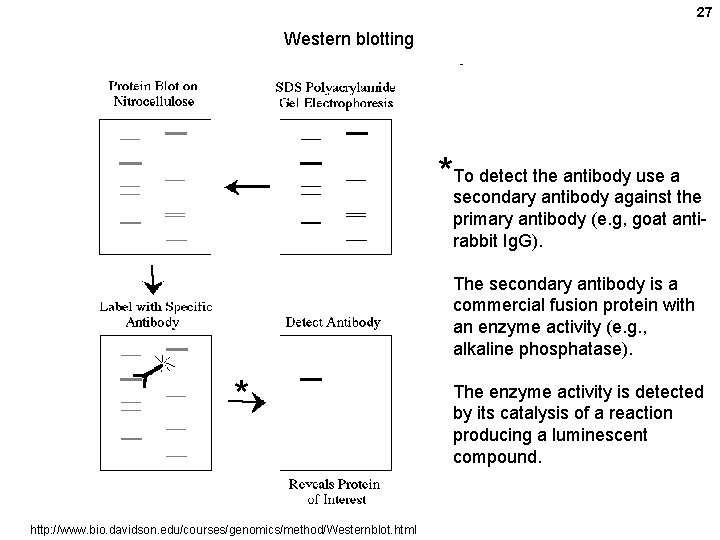
27 Western blotting detect the antibody use a *To secondary antibody against the primary antibody (e. g, goat antirabbit Ig. G). The secondary antibody is a commercial fusion protein with an enzyme activity (e. g. , alkaline phosphatase). * http: //www. bio. davidson. edu/courses/genomics/method/Westernblot. html The enzyme activity is detected by its catalysis of a reaction producing a luminescent compound.
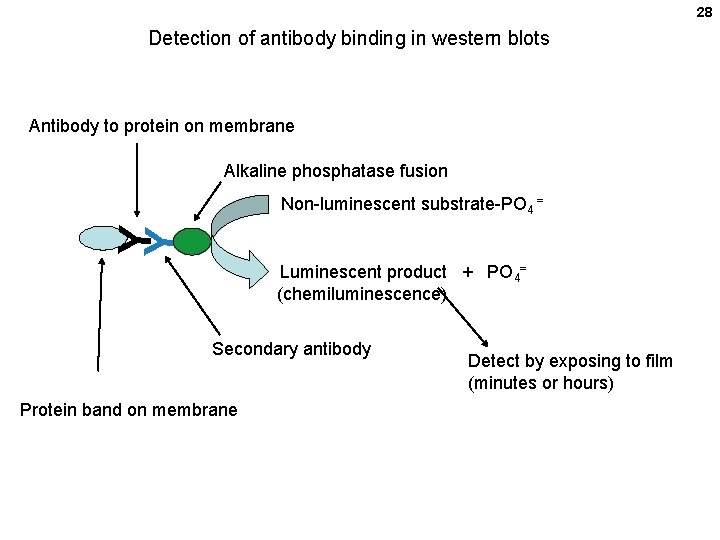
28 Detection of antibody binding in western blots Antibody to protein on membrane Alkaline phosphatase fusion Y Y Non-luminescent substrate-PO 4 = Luminescent product + PO 4= (chemiluminescence) Secondary antibody Protein band on membrane Detect by exposing to film (minutes or hours)
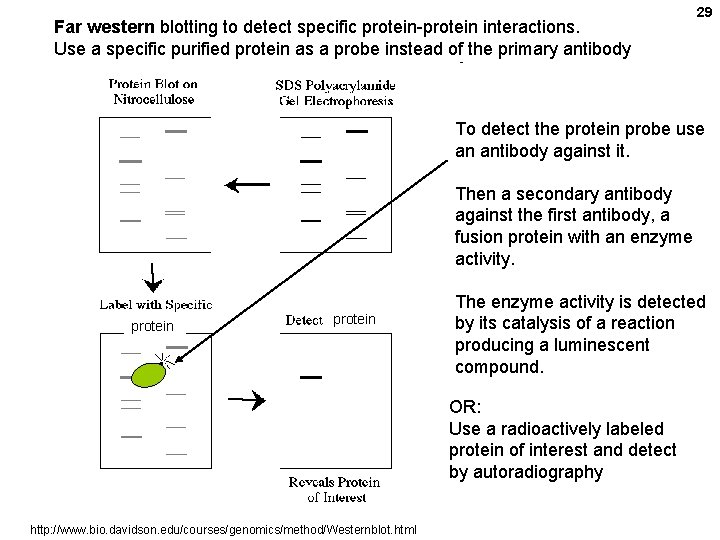
Far western blotting to detect specific protein-protein interactions. Use a specific purified protein as a probe instead of the primary antibody 29 To detect the protein probe use an antibody against it. Then a secondary antibody against the first antibody, a fusion protein with an enzyme activity. protein The enzyme activity is detected by its catalysis of a reaction producing a luminescent compound. OR: Use a radioactively labeled protein of interest and detect by autoradiography http: //www. bio. davidson. edu/courses/genomics/method/Westernblot. html

How to make a radioactively labeled protein: Expression via in vitro transcription followed by in vitro translation 30 T 7 RNA polymerase binding site (17 -21 nt) c. DNA VECTOR …. ACCATGG…. . Radioactively labeled protein 1. Transcription to m. RNA via the T 7 promoter + T 7 polymerase 2. Add a translation system: rabbit reticulocyte lysate or wheat germ lysate Or: E. coli lysate (combined transcription + translation, Tn. T) All commercially available as kits Add ATP, GTP, t. RNAs, amino acids, label (35 S-met), May need to add RNase (Ca++-dependent, stop with EGTA) to remove endogenous m. RNA In lysate NOTE: Protein is NOT at all pure (1000 s of lysate proteins present), just ~“radio-pure”
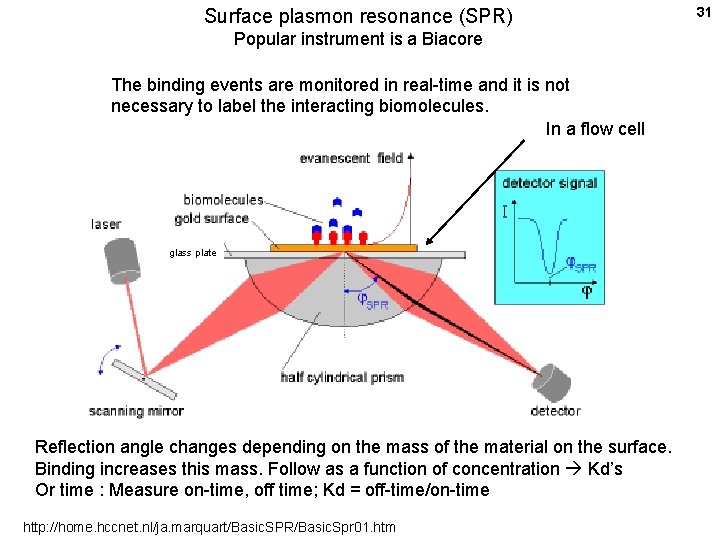
Surface plasmon resonance (SPR) Popular instrument is a Biacore The binding events are monitored in real-time and it is not necessary to label the interacting biomolecules. In a flow cell glass plate Reflection angle changes depending on the mass of the material on the surface. Binding increases this mass. Follow as a function of concentration Kd’s Or time : Measure on-time, off time; Kd = off-time/on-time http: //home. hccnet. nl/ja. marquart/Basic. SPR/Basic. Spr 01. htm 31
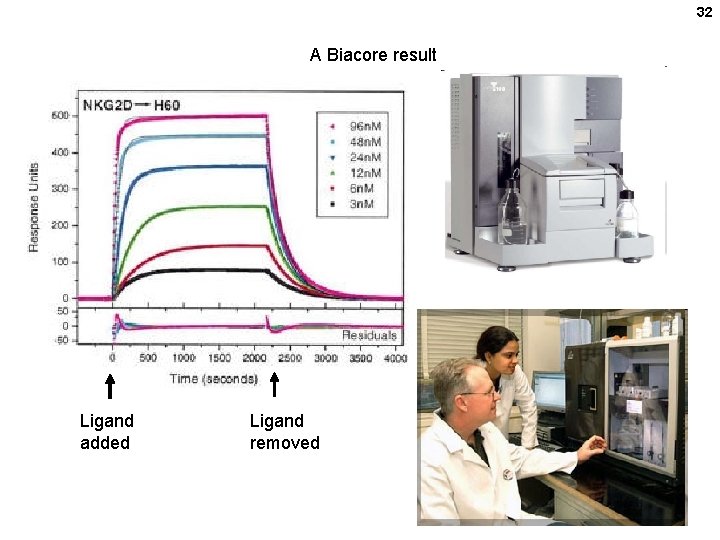
32 A Biacore result Ligand added Ligand removed
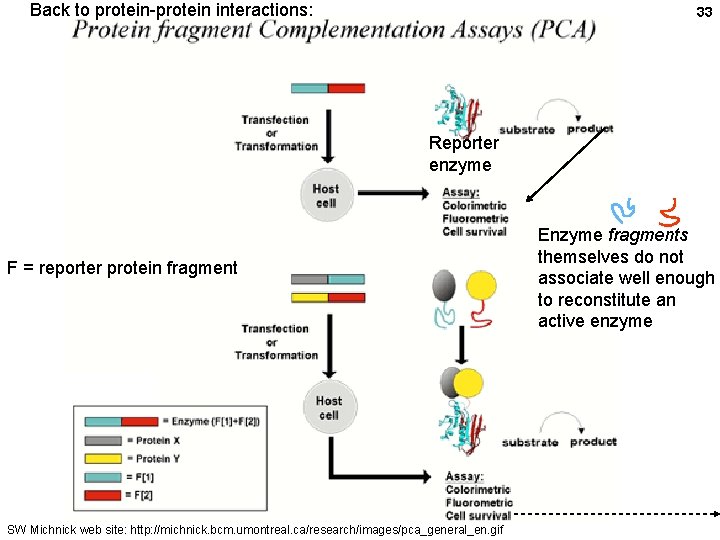
Back to protein-protein interactions: 33 Reporter enzyme F = reporter protein fragment SW Michnick web site: http: //michnick. bcm. umontreal. ca/research/images/pca_general_en. gif Enzyme fragments themselves do not associate well enough to reconstitute an active enzyme
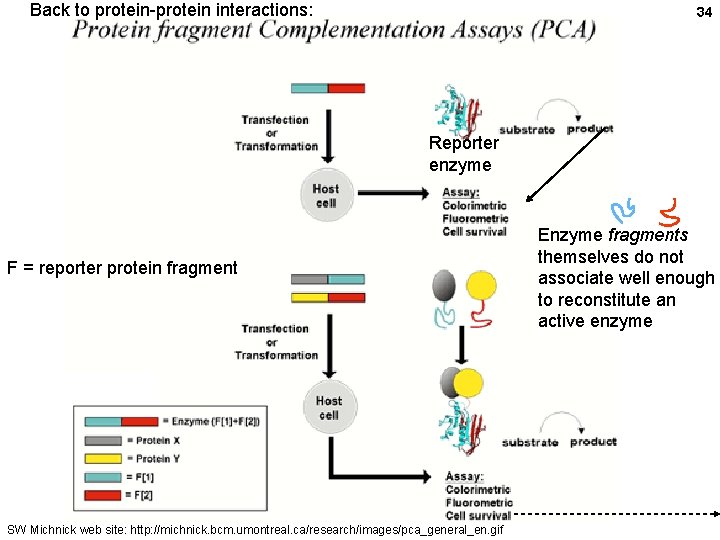
Back to protein-protein interactions: 34 Reporter enzyme F = reporter protein fragment SW Michnick web site: http: //michnick. bcm. umontreal. ca/research/images/pca_general_en. gif Enzyme fragments themselves do not associate well enough to reconstitute an active enzyme
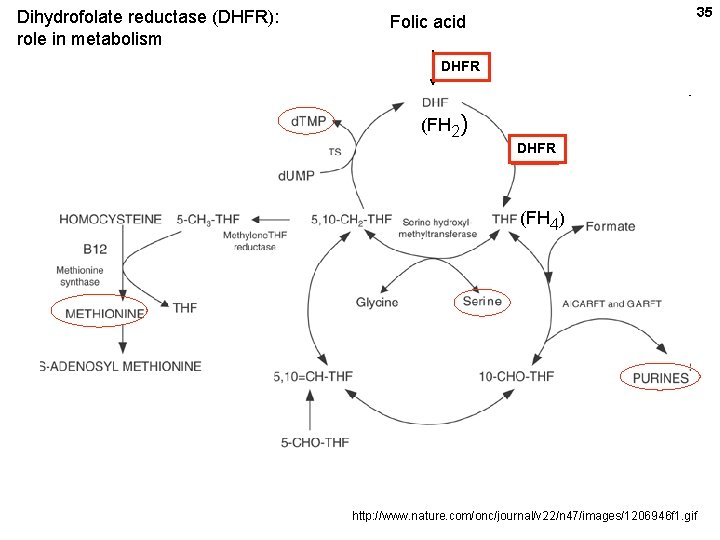
Dihydrofolate reductase (DHFR): role in metabolism 35 Folic acid DHFR (FH 2) DHFR (FH 4) http: //www. nature. com/onc/journal/v 22/n 47/images/1206946 f 1. gif
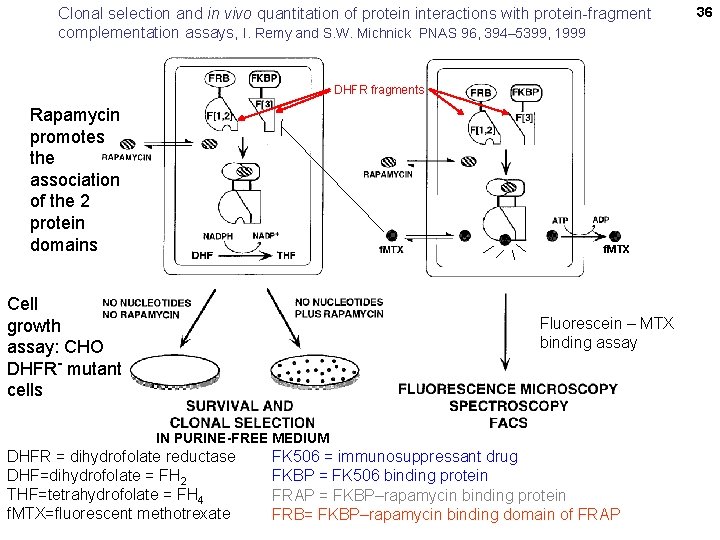
Clonal selection and in vivo quantitation of protein interactions with protein-fragment complementation assays, I. Remy and S. W. Michnick PNAS 96, 394– 5399, 1999 DHFR fragments Rapamycin promotes the association of the 2 protein domains f. MTX Cell growth assay: CHO DHFR- mutant cells Fluorescein – MTX binding assay IN PURINE-FREE MEDIUM DHFR = dihydrofolate reductase DHF=dihydrofolate = FH 2 THF=tetrahydrofolate = FH 4 f. MTX=fluorescent methotrexate FK 506 = immunosuppressant drug FKBP = FK 506 binding protein FRAP = FKBP–rapamycin binding protein FRB= FKBP–rapamycin binding domain of FRAP 36
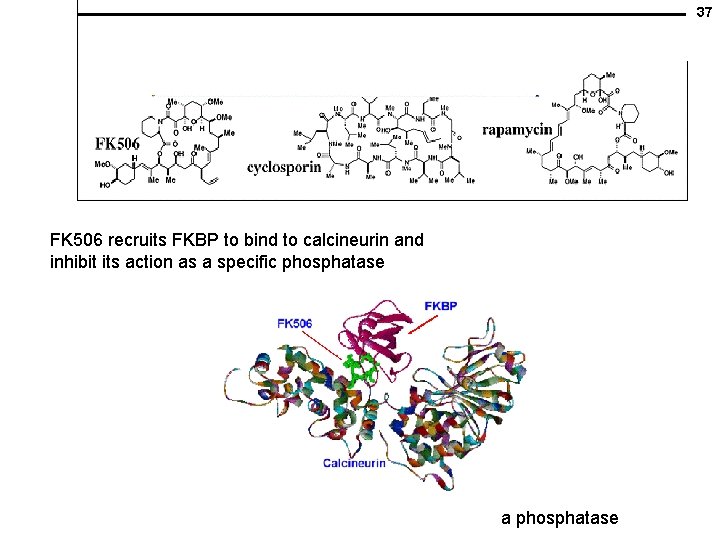
37 FK 506 recruits FKBP to bind to calcineurin and inhibit its action as a specific phosphatase a phosphatase
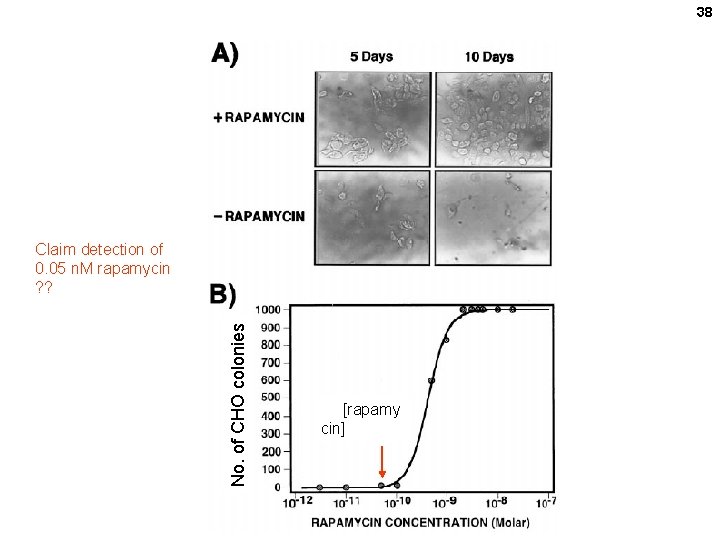
38 No. of CHO colonies Claim detection of 0. 05 n. M rapamycin ? ? [rapamy cin]
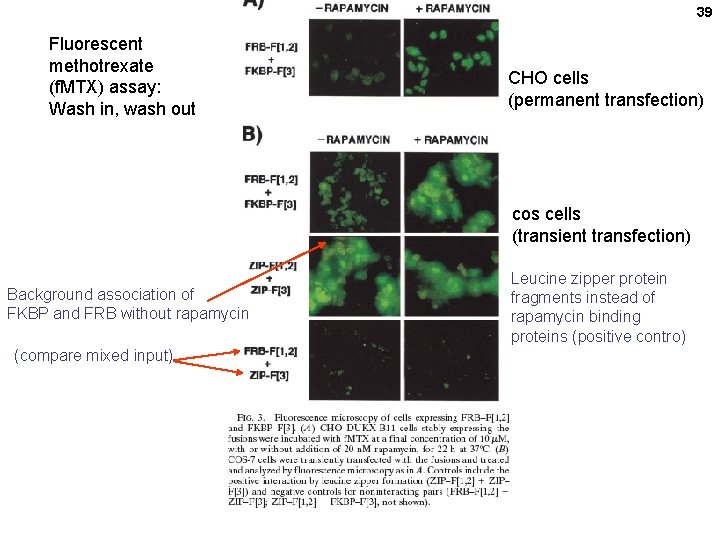
39 Fluorescent methotrexate (f. MTX) assay: Wash in, wash out CHO cells (permanent transfection) cos cells (transient transfection) Background association of FKBP and FRB without rapamycin (compare mixed input) Leucine zipper protein fragments instead of rapamycin binding proteins (positive contro)
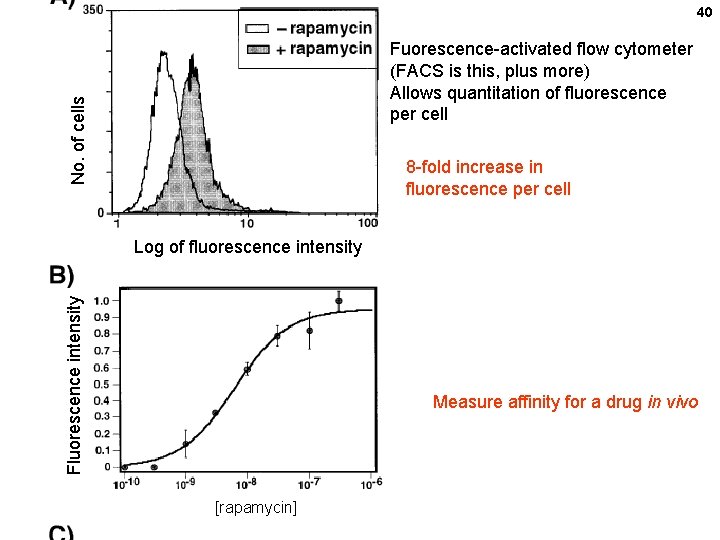
40 No. of cells Fuorescence-activated flow cytometer (FACS is this, plus more) Allows quantitation of fluorescence per cell 8 -fold increase in fluorescence per cell Fluorescence intensity Log of fluorescence intensity Measure affinity for a drug in vivo [rapamycin]
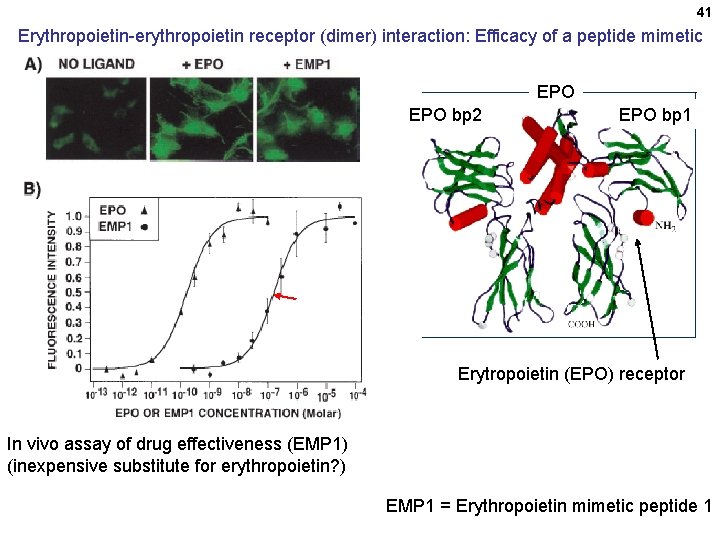
41 Erythropoietin-erythropoietin receptor (dimer) interaction: Efficacy of a peptide mimetic EPO bp 2 EPO bp 1 Erytropoietin (EPO) receptor In vivo assay of drug effectiveness (EMP 1) (inexpensive substitute for erythropoietin? ) EMP 1 = Erythropoietin mimetic peptide 1 Erythropoietin
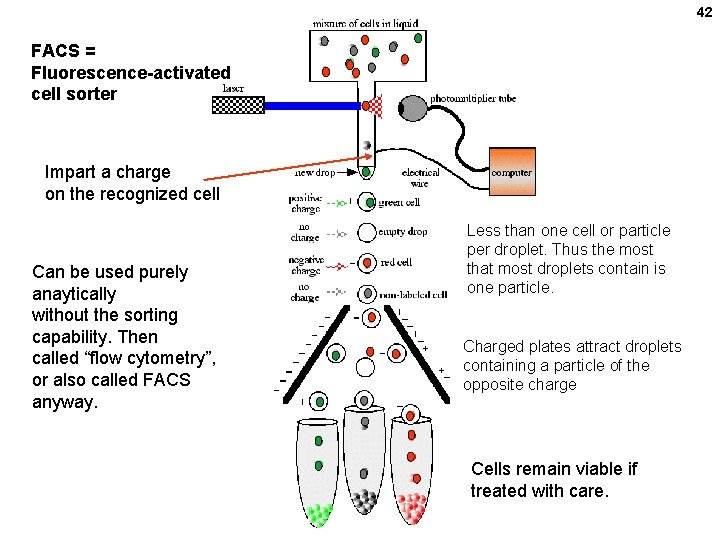
42 FACS = Fluorescence-activated cell sorter Impart a charge on the recognized cell Can be used purely anaytically without the sorting capability. Then called “flow cytometry”, or also called FACS anyway. Less than one cell or particle per droplet. Thus the most that most droplets contain is one particle. Charged plates attract droplets containing a particle of the opposite charge Cells remain viable if treated with care.
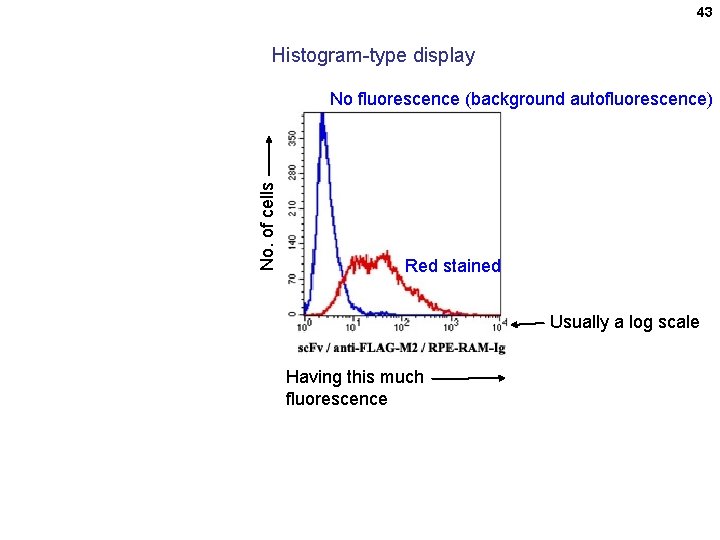
43 Histogram-type display No. of cells No fluorescence (background autofluorescence) Red stained Usually a log scale Having this much fluorescence
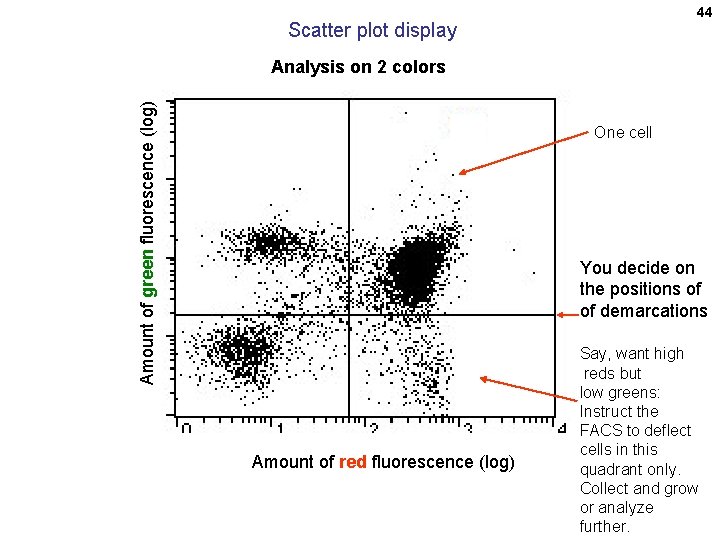
44 Scatter plot display Amount of green fluorescence (log) Analysis on 2 colors One cell You decide on the positions of of demarcations Amount of red fluorescence (log) Say, want high reds but low greens: Instruct the FACS to deflect cells in this quadrant only. Collect and grow or analyze further.

45
- Slides: 45