1 Genome sequence assembly Assembly concepts and methods

1

Genome sequence assembly Assembly concepts and methods Mihai Pop Center for Bioinformatics and Computational Biology University of Maryland 2

Building a library • Break DNA into random fragments (8 -10 x coverage) Actual situation 3

Building a library • Break DNA into random fragments (8 -10 x coverage) • Sequence the ends of the fragments – Amplify the fragments in a vector – Sequence 800 -1000 (500 -700) bases at each end of the fragment 4
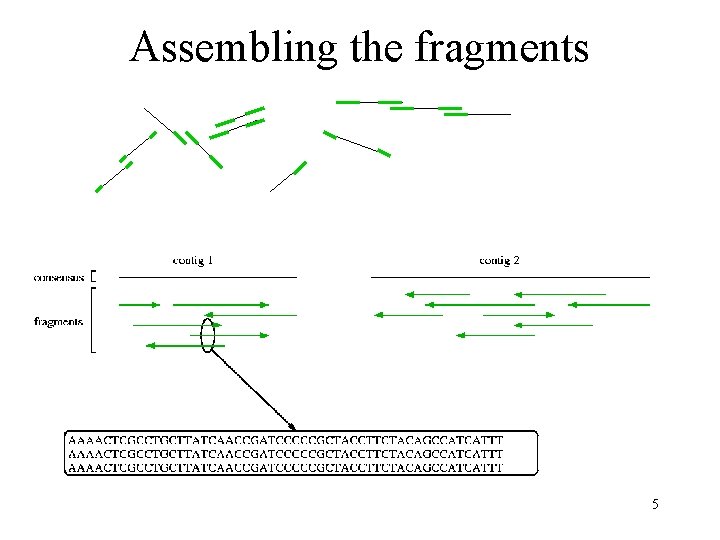
Assembling the fragments 5
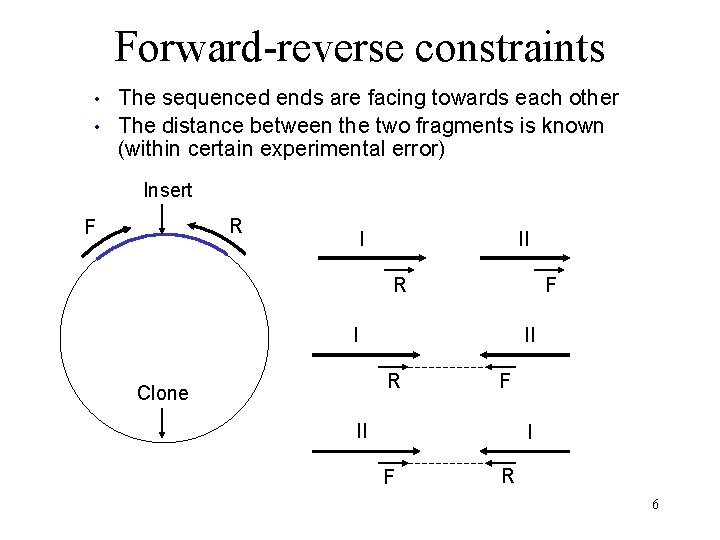
Forward-reverse constraints • • The sequenced ends are facing towards each other The distance between the two fragments is known (within certain experimental error) Insert R F I II R Clone F II I F R 6

Building Scaffolds • • Break DNA into random fragments (8 -10 x coverage) Sequence the ends of the fragments Assemble the sequenced ends Build scaffolds 7
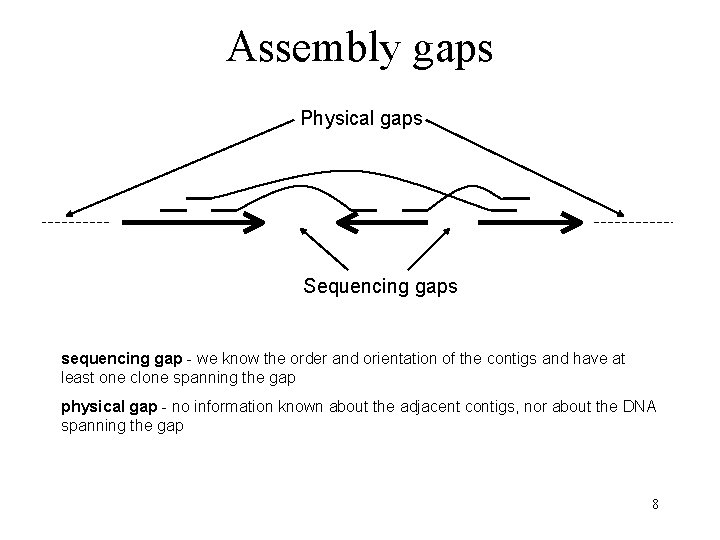
Assembly gaps Physical gaps Sequencing gaps sequencing gap - we know the order and orientation of the contigs and have at least one clone spanning the gap physical gap - no information known about the adjacent contigs, nor about the DNA spanning the gap 8

Unifying view of assembly Assembly Scaffolding 9

Shotgun sequencing statistics 10

Typical contig coverage Imagine raindrops on a sidewalk 11

Lander-Waterman statistics L = read length T = minimum detectable overlap G = genome size N = number of reads c = coverage (NL / G) σ = 1 – T/L E(#islands) = Ne-cσ E(island size) = L((ecσ – 1) / c + 1 – σ) contig = island with 2 or more reads 12
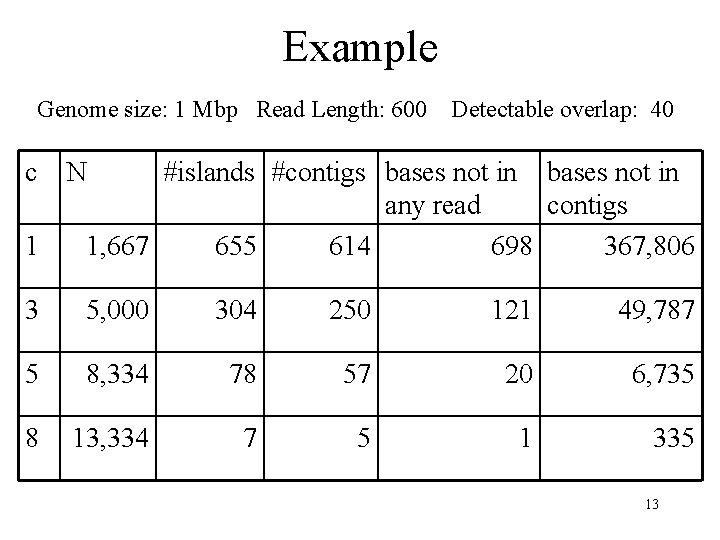
Example Genome size: 1 Mbp Read Length: 600 c Detectable overlap: 40 N 1 #islands #contigs bases not in any read contigs 1, 667 655 614 698 367, 806 3 5, 000 304 250 121 49, 787 5 8, 334 78 57 20 6, 735 8 13, 334 7 5 1 335 13
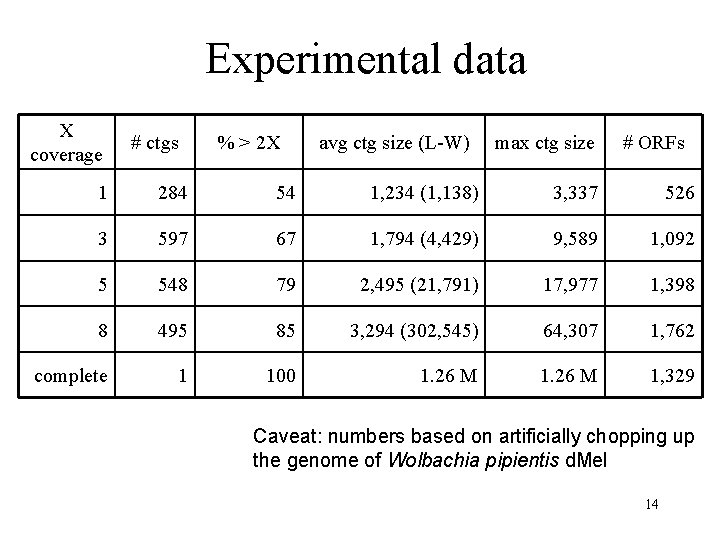
Experimental data X coverage # ctgs % > 2 X avg ctg size (L-W) max ctg size # ORFs 1 284 54 1, 234 (1, 138) 3, 337 526 3 597 67 1, 794 (4, 429) 9, 589 1, 092 5 548 79 2, 495 (21, 791) 17, 977 1, 398 8 495 85 3, 294 (302, 545) 64, 307 1, 762 complete 1 100 1. 26 M 1, 329 Caveat: numbers based on artificially chopping up the genome of Wolbachia pipientis d. Mel 14

Read coverage vs. Clone coverage 4 kbp 1 kbp Read coverage = 8 X Clone (insert) coverage = 16 2 X coverage in BAC-ends implies 100 x coverage by BACs (1 BAC clone = approx. 100 kbp) 15

Assembly paradigms • Overlap-layout-consensus – greedy (TIGR Assembler, phrap, CAP 3. . . ) – graph-based (Celera Assembler, Arachne) • Eulerian path (especially useful for short read sequencing) 16

TIGR Assembler/phrap Greedy • Build a rough map of fragment overlaps • Pick the largest scoring overlap • Merge the two fragments • Repeat until no more merges can be done 17
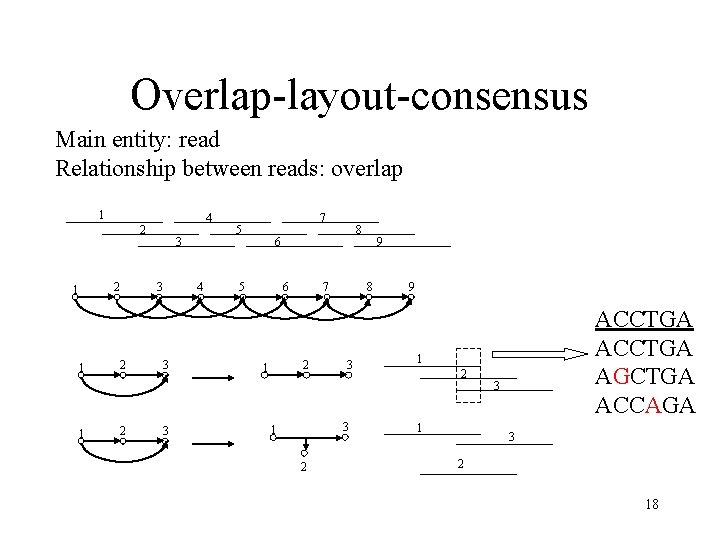
Overlap-layout-consensus Main entity: read Relationship between reads: overlap 1 4 2 2 1 1 1 3 3 2 2 4 3 3 7 5 8 6 5 6 7 2 1 8 3 3 1 2 9 9 ACCTGA AGCTGA ACCAGA 1 2 1 3 3 2 18

Paths through graphs and assembly • Hamiltonian circuit: visit each node (city) exactly once, returning to the start Genome 19

Implementation details 20

Overlap between two sequences overlap (19 bases) overhang (6 bases) …AGCCTAGACCTACAGGATGCGCGGACACGTAGC CAGGAC CAGTACTTGGATGCGCTGACACGTAGCTTATCCGGT… overhang % identity = 18/19 % = 94. 7% overlap - region of similarity between regions overhang - un-aligned ends of the sequences The assembler screens merges based on: • length of overlap • % identity in overlap region • maximum overhang size. 21

All pairs alignment • Needed by the assembler • Try all pairs – must consider ~ n 2 pairs • Smarter solution: only n x coverage (e. g. 8) pairs are possible – Build a table of k-mers contained in sequences (single pass through the genome) – Generate the pairs from k-mer table (single pass through k-mer table) k-mer 22

23

REPEATS 24
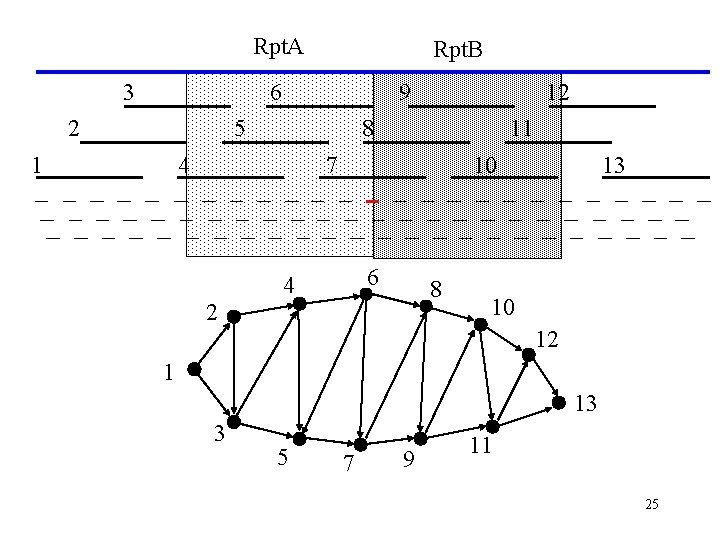
Rpt. A 3 Rpt. B 6 2 9 5 1 12 8 4 11 7 10 6 4 8 2 13 10 12 1 13 3 5 7 9 11 25
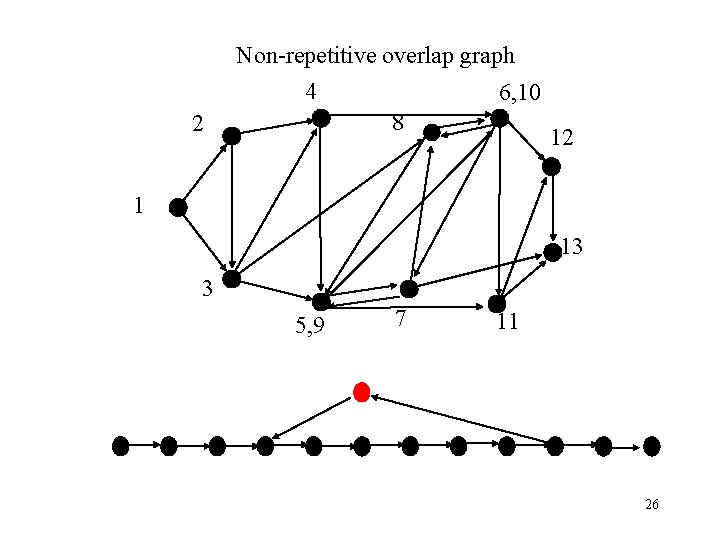
Non-repetitive overlap graph 4 6, 10 8 2 12 1 13 3 5, 9 7 11 26

Handling repeats 1. Repeat detection – pre-assembly: find fragments that belong to repeats • • – – statistically (most existing assemblers) repeat database (Repeat. Masker) during assembly: detect "tangles" indicative of repeats (Pevzner, Tang, Waterman 2001) post-assembly: find repetitive regions and potential mis-assemblies. • • Reputer, Repeat. Masker "unhappy" mate-pairs (too close, too far, mis-oriented) 2. Repeat resolution – – find DNA fragments belonging to the repeat determine correct tiling across the repeat 27

Statistical repeat detection Significant deviations from average coverage flagged as repeats. - frequent k-mers are ignored - “arrival” rate of reads in contigs compared with theoretical value (e. g. , 800 bp reads & 8 x coverage - reads "arrive" every 100 bp) Problem 1: assumption of uniform distribution of fragments - leads to false positives non-random libraries poor clonability regions Problem 2: repeats with low copy number are missed - leads to false negatives 28

Mis-assembled repeats collapsed tandem excision rearrangement 29

SASA repeat (4776 AA, 14 Kb) from Streptococcus pneumoniae MTETVEDKVSHSITGLDILKGIVAAGAVISGTVATQTKVFTNESAVLEKTVEKTDALATNDTVVLGTISTSNSASSTSLSASESASTSASTSASESASTSISASSTVVGSQTAAA TEATAKKVEEDRKKPASDYVASVTNVNLQSYAKRRKRSVDSIEQLLASIKNAAVFSGNTIVNGAPAINASLNIAKSETKVYTGEGVDSVYRVPIYYKLKVTNDGSKLTFTYTVTYVNPKTNDLGNISSMRP GYSIYNSGTSTQTMLTLGSDLGKPSGVKNYITDKNGRQVLSYNTSTMTTQGSGYTWGNGAQMNGFFAKKGYGLTSSWTVPITGTDTSFTFTPYAARTDRIGINYFNGGGKVVESSTTSQSLSQSKSLSVSA SQSASASASTSASASASTSASVSASTSASASASTSASASASTSASESASTSASASASTSASESASTSASASASTS ASGSASTSTSASASASTSASASASISASESASTSTSASASTSASESASTSASASASTSASASASTSASASASTSASASA STSASVSASTSASASASTSASASASTSASESASTSASASASTSASASASTSASASASTSASASASISASESASTS ASASASTSASASASTSASESASTSASASASTSASASASTSASASASTSASESASTSASASASTSASASASTSASASASTSASASASTSASASASISASESASTSASASASTSASVSASTS ASASASTSASESASTSASASASTSASASASISASESASTSASASASTSASASASTSASESASTSTSASASTSASESASTSASASASTSASASASTSASASASTSASASASTSASESASTSASASASTSASASASTSASVSASTS ASESASTSASASASTSASESASTSASASASTSASASASTSASASASTSASASASTSASASASTSASASASISASESASTSASASASTSASVSASTSASASASTSASASASISASESASTSASASASTSASASASTSASASASTSASASASTSASESASTSASASASTSASASASTSASVSASTSASESASTSASASASTSASASASTSASASASTSASASASTSASASASISASESASTSTSASASTSASESASTSASASASTSASASTSASESASTSASASASTSASASASTSASASASTSASVSASTSASASASTSASESASTSASASASTSASASASTSASASASISASESASTSASASASTSASASASTSASASASTSASASASTSASESASTSASASASTSASASASTSASASASTSASASASTSASESASTSASASASTSASASASTSASESAST SASASASTSASASASTSASASASISASESASTSASASASTSASVSASTSASASASTSASESASTSASASASISASESASTSASASASTSASESASTSTSASASTSASESASTSASASASTSASASASTSASASASTSASASASTSASASASTSASESASTSASASASTSASASASTSASVSASTSASESASTSASASASTSASASASTSASESASTSASASASTSASASASASTSASASASTSASASASISASESASTSASASASTSASASASISASESASTST SASASTSASESASTSASASASTSASASASTSASASASTSASASASTSASVSASTSASASASTSASESASTSASASTS ASESASTSASASASTSASASASTSASESASTSASASASTSASASASTSASESASTSASASASTSASASASTSASG SASTSTSASASASTSASASASISASESASTSTSASASTSASESASTSASASASTSASASASTSASASASTSASASASTS ASVSASTSASASASTSASASASTSASESASTSASASASTSASASASTSASASASTSASASASISASESASTSASASASTSASESASTSASASASTSASASASTSASASASTSASESASTSASASASTSASASASTSASASASTSASASTSASESASTSASASASTSASASASTSASASASTSASASASISASESASTSASASASTSASVSASTSASA SASTSASESASTSASASASTSASASASISASESASTSASASASTSASASASTSASESASTSTSASASTSASESASTSASASASTSASASASTSASASASTSASASASTSASESASTSASASASTSASASASTSASVSASTSASE SASTSASASASTSASESASTSASASASTSASASASTSASASASTSASASASTSASASASTSASASASISASESASTSASASASTSASVSASTSASASASTSASASASISASESASTSASASASTSASASASTSASASASTSASASASTSASESASTSASASASISASESASTSASASASTSASASASTSASESASTSTSASASTSASES ASTSASASASTSASASASTSASESASTSASASASTSASASASTSASASASTSASASASTSASESASTSASASASTSASASASTSASVSASTSASESASTSASASASTSASESASTSASASASTSASASASTSASASASTSASASASISASESASTSASASASTSASVSASTSASASASISASESASTSASASASTSASASASTSASASASTSASASASTSASASASTSASASASTSASASASTSVS NSANHSNSQVGNTSGSTGKSQKELPNTGTESSIGSVLLGVLAAVTGIGLVAKRRKRDEEE 30
- Slides: 30