1 CONTENTS Requirements for translation Genetic code Mutations




























- Slides: 70

1

CONTENTS ØRequirements for translation ØGenetic code ØMutations – types and their effects ØProcess of translation ØInhibitors of translation ØPost-translational modification of proteins ØRegulation of gene expression in Prokaryotes and Eukaryotes 2

TRANSLATION A cytoplasmic process where the genetic information present in the form of nucleotide sequences in m–RNA is translated into the language of amino acid sequence (proteins). 3

Translation……. . Genetic information (genes in DNA) Transcription Genetic information (m-RNA) Translation Amino acid sequence in proteins. 4

Translation……. Site of protein biosynthesis Sub-cellular site : Cytosol. : Ribosomes (free), rough endoplasmic reticulum. Template for protein synthesis : m. RNA. Amino acids – all the 20 amino acids should be present at the same time in the cytosol (amino acid pool). 5

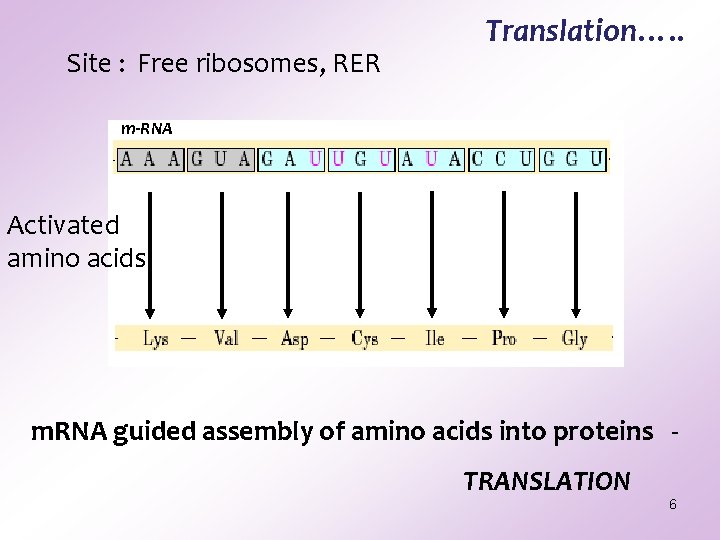
Site : Free ribosomes, RER Translation…. . m-RNA Activated amino acids m. RNA guided assembly of amino acids into proteins TRANSLATION 6

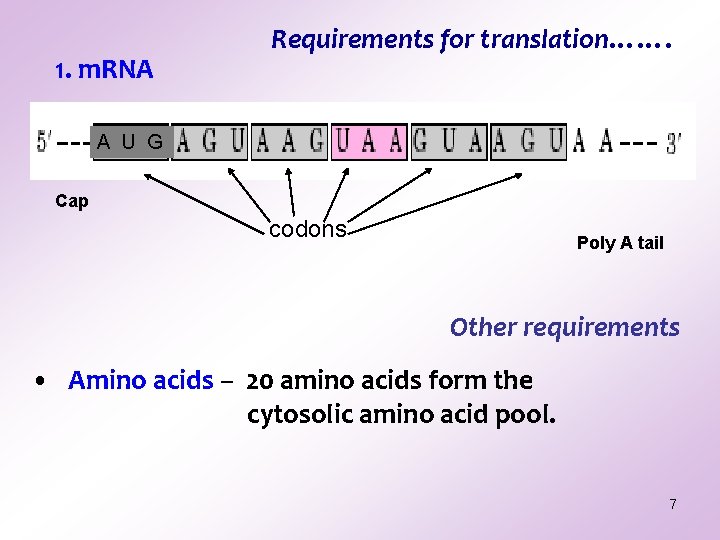
1. m. RNA Requirements for translation……. A U G Cap codons Poly A tail Other requirements • Amino acids – 20 amino acids form the cytosolic amino acid pool. 7

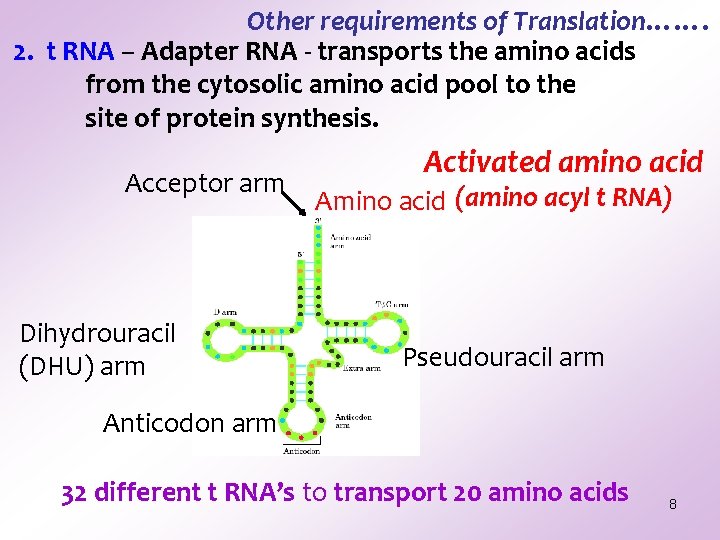
Other requirements of Translation……. 2. t RNA – Adapter RNA - transports the amino acids from the cytosolic amino acid pool to the site of protein synthesis. Acceptor arm Dihydrouracil (DHU) arm Activated amino acid Amino acid (amino acyl t RNA) Pseudouracil arm Anticodon arm 32 different t RNA’s to transport 20 amino acids 8

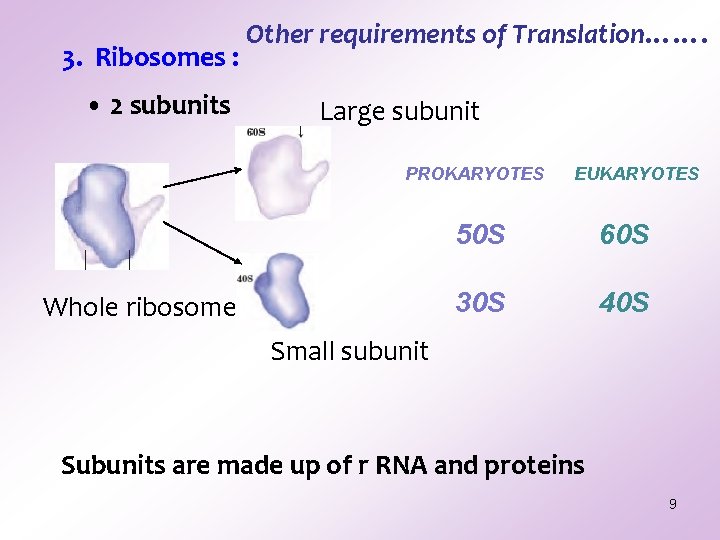
3. Ribosomes : • 2 subunits Other requirements of Translation……. Large subunit PROKARYOTES Whole ribosome EUKARYOTES 50 S 60 S 30 S 40 S Small subunit Subunits are made up of r RNA and proteins 9

Other requirements of Translation…Ribosome Subunits are - • Oddly shaped. • Fit together to form a cleft through which the m. RNA moves and the newly formed polypeptide emerges during translation process. • Size is different in prokaryotes and eukaryotes. 10

Other requirements of Translation • Enzymes • Protein factors – Initiation factors, Elongation factors, Termination factors • Energy ---- ATP, GTP • Mg++ 11

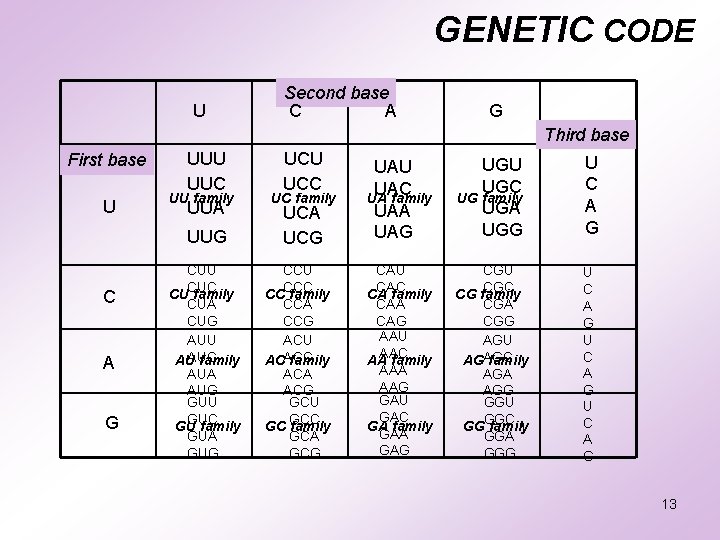
GENETIC CODE m-RNA : codes for numerous amino acids. directs the amino acid sequences in a protein. There are 4 nitrogenous bases – A, G, U, C. If Codon has 2 bases 42 = 16 codons is possible. If Codon has 3 bases 43 = 64 codons is possible Each codon is a triplet of three bases. Genetic code contains 64 codons coding for 20 amino acids. 12

GENETIC CODE U Second base C A G Third base First base U UUC UU family UUA UUG C A G CUU CUC CU family CUA CUG AUU AUC AU family AUA AUG GUU GUC GU family GUA GUG UCU UCC UC family UCA UCG CCU CCC CC family CCA CCG ACU ACACC family ACA ACG GCU GCC GC family GCA GCG UAU UAC UA family UAA UAG UGU UGC UG family UGA UGG U C A G CAU CAC CA family CAA CAG AAU AAC AA family AAA AAG GAU GAC GA family GAA GAG CGU CGC CG family CGA CGG AGU AGAGC family AGA AGG GGU GGC GG family GGA GGG U C A G 13

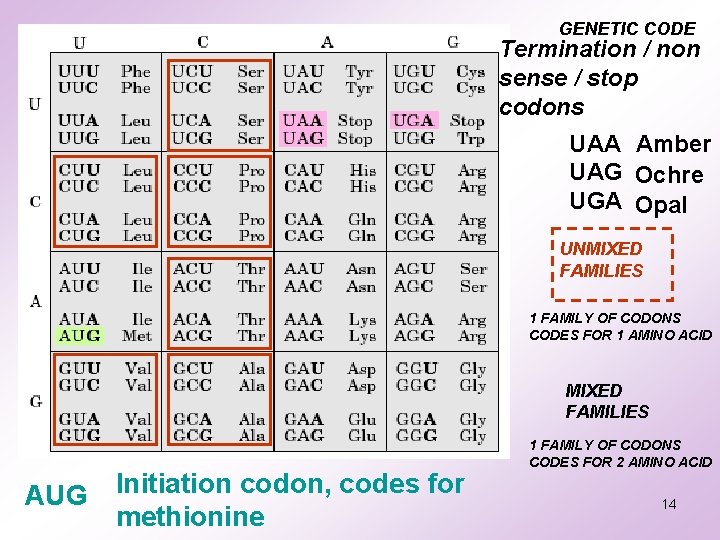
GENETIC CODE Termination / non sense / stop codons UAA Amber UAG Ochre UGA Opal UNMIXED FAMILIES 1 FAMILY OF CODONS CODES FOR 1 AMINO ACID MIXED FAMILIES AUG Initiation codon, codes for methionine 1 FAMILY OF CODONS CODES FOR 2 AMINO ACID 14

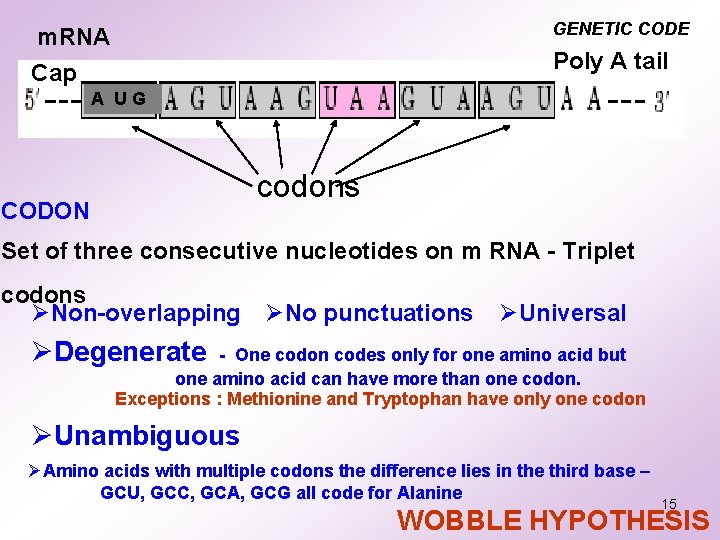
GENETIC CODE m. RNA Cap Poly A tail A UG codons CODON Set of three consecutive nucleotides on m RNA - Triplet codons ØNon-overlapping ØNo punctuations ØUniversal ØDegenerate - One codon codes only for one amino acid but one amino acid can have more than one codon. Exceptions : Methionine and Tryptophan have only one codon ØUnambiguous ØAmino acids with multiple codons the difference lies in the third base – GCU, GCC, GCA, GCG all code for Alanine 15 WOBBLE HYPOTHESIS

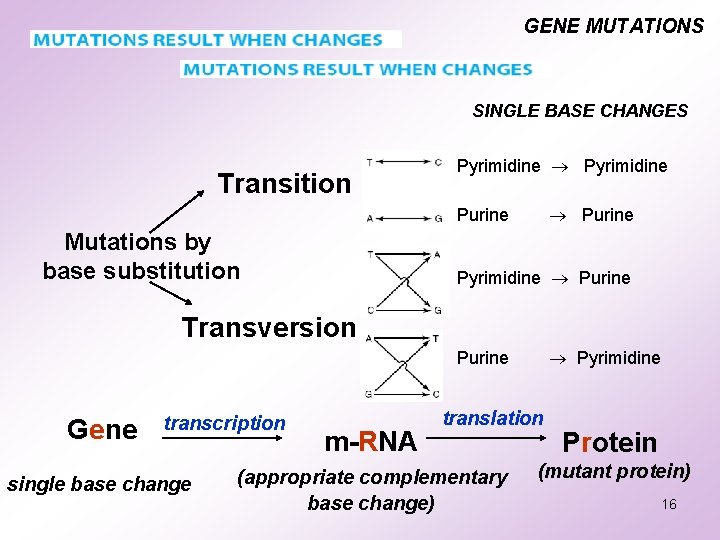
GENE MUTATIONS SINGLE BASE CHANGES Transition Pyrimidine Purine Mutations by base substitution Pyrimidine Purine Transversion Pyrimidine Purine Gene transcription single base change m-RNA translation (appropriate complementary base change) Protein (mutant protein) 16

EFFECT OF SINGLE BASE CHANGES v Silent mutation No on the function of the protein Hbeffect Milwaukee, Hb Sydney v Missense mutation Different amino acid is incorporated in the mutant protein 3 types – based on the location of the amino acid replacement in the protein Acceptable mutation Mutant Hbprotein Hikari is not recognizable from the native protein Partially acceptable mutation Hb Sprotein (Sicklehas cellpartial, hemoglobin) Mutant abnormal function Unacceptable mutation Mutant protein is Hb incapable of doing its assigned function M v Mutation leading to nonsense codon Premature termination of the protein which may or may not be functional FRAME SHIFT MUTATIONS Cause : Insertion or deletion of bases Result : Altered reading frame beyond the point of deletion / insertion Garbled protein beyond the point of mutation Terminated protein if a nonsense codon appears 17

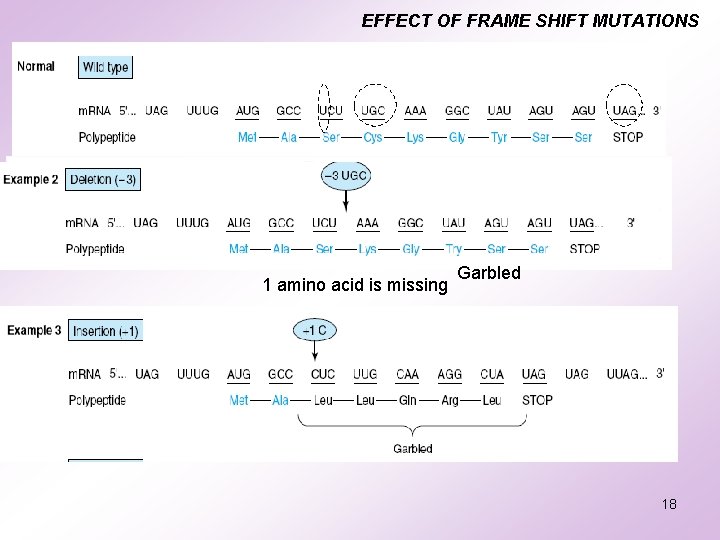
EFFECT OF FRAME SHIFT MUTATIONS 1 amino acid is missing Garbled 18

TRANSLATION Direction of protein synthesis : N-terminal to C-terminal Direction in which m-RNA is read : 5’ to 3’ direction STAGES OF TRANSLATION – 5 stages AMINOACYL-t RNA SYNTHESIS – Activation of amino acids. INITIATION ELONGATION TERMINATION POST-TRANSLATIONAL MODIFICATIONS 19

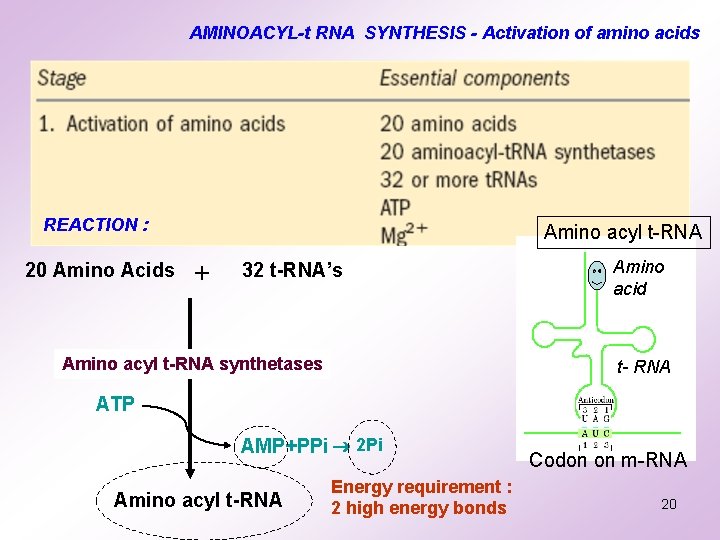
AMINOACYL-t RNA SYNTHESIS - Activation of amino acids REACTION : 20 Amino Acids Amino acyl t-RNA + 32 t-RNA’s Amino acyl t-RNA synthetases Amino acid t- RNA ATP AMP+PPi 2 Pi Amino acyl t-RNA Energy requirement : 2 high energy bonds Codon on m-RNA 20

AMINOACYL-t RNA SYNTHESIS Proof reading Activity Binds very specifically to corresponding amino acid & t-RNA. Bigger / Smaller molecules are hydrolysed Amino acyl t-RNA synthetase enzyme Amino acylation site Hydrolytic site 21

INITIATION OF PROTEIN BIOSYNTHESIS Synthesis of N-formyl methionine : Initiation involves 3 steps Enzyme : Transformylase 22

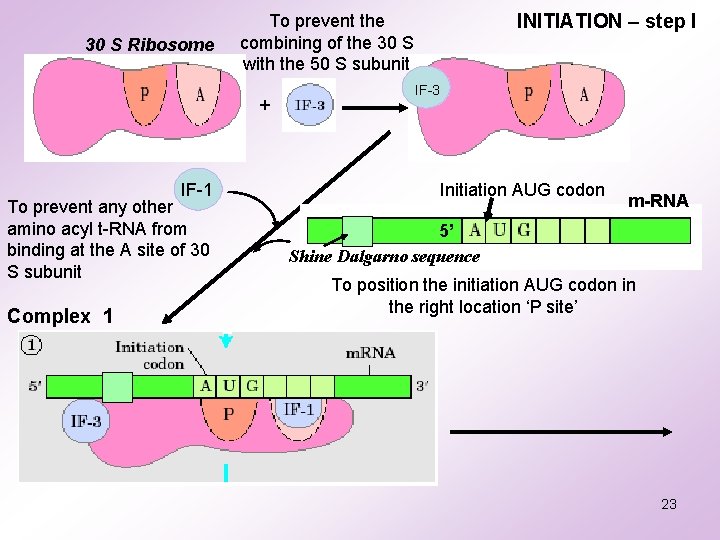
30 S Ribosome + IF-1 To prevent any other amino acyl t-RNA from binding at the A site of 30 S subunit Complex 1 INITIATION – step I To prevent the combining of the 30 S with the 50 S subunit IF-3 Initiation AUG codon m-RNA 5’ Shine Dalgarno sequence To position the initiation AUG codon in the right location ‘P site’ 23

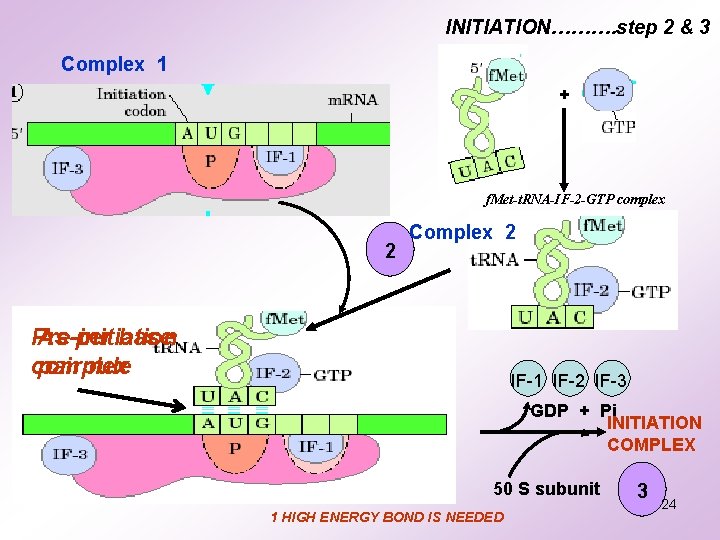
INITIATION………. step 2 & 3 Complex 1 + f. Met-t. RNA-IF-2 -GTP complex 2 Complex 2 Pre-initiation As per base complex pair rule IF-1 IF-2 IF-3 GDP + Pi INITIATION COMPLEX 50 S subunit 1 HIGH ENERGY BOND IS NEEDED 3 24

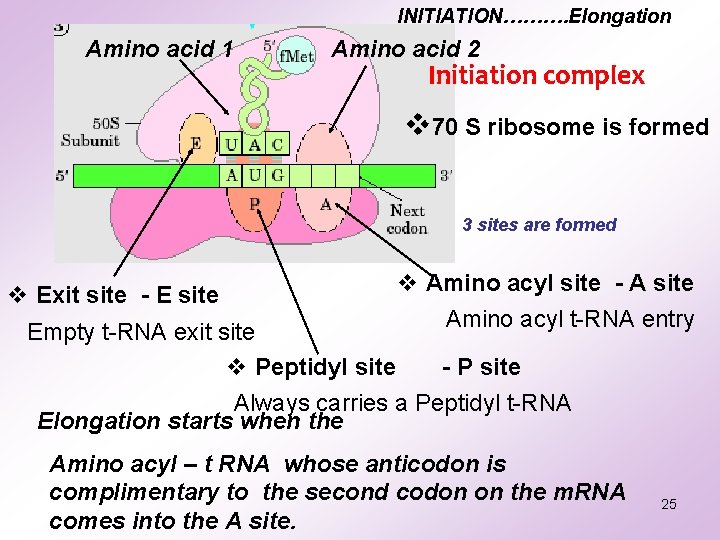
INITIATION………. Elongation Amino acid 1 Amino acid 2 Initiation complex v 70 S ribosome is formed 3 sites are formed v Exit site - E site v Amino acyl site - A site Amino acyl t-RNA entry Empty t-RNA exit site v Peptidyl site - P site Always carries a Peptidyl t-RNA Elongation starts when the Amino acyl – t RNA whose anticodon is complimentary to the second codon on the m. RNA comes into the A site. 25

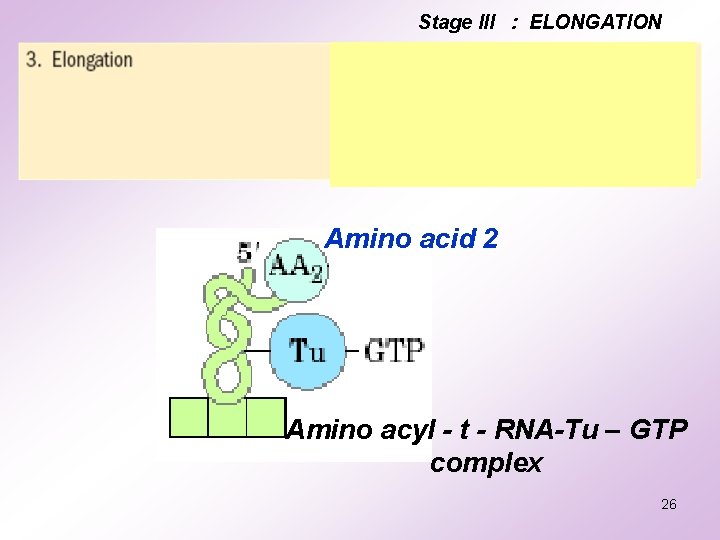
Stage III : ELONGATION Amino acid 2 Amino acyl - t - RNA-Tu – GTP complex 26

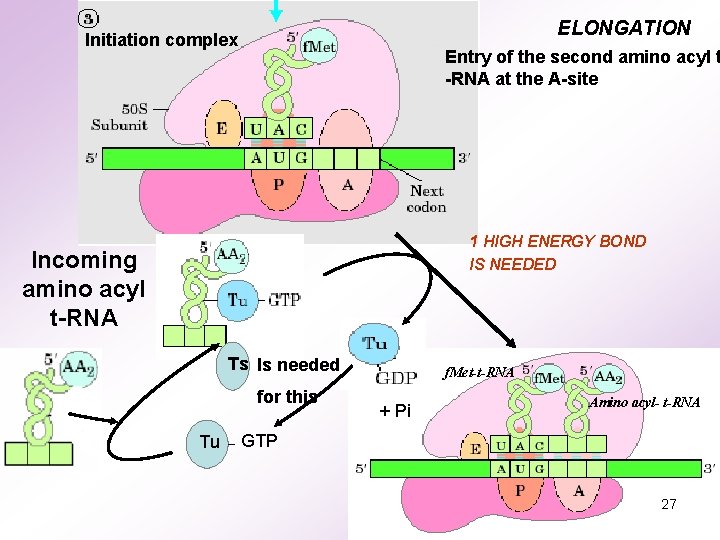
ELONGATION Initiation complex Entry of the second amino acyl t -RNA at the A-site 1 HIGH ENERGY BOND IS NEEDED Incoming amino acyl t-RNA Ts Is needed for this Tu f. Met-t-RNA + Pi Amino acyl- t-RNA GTP 27

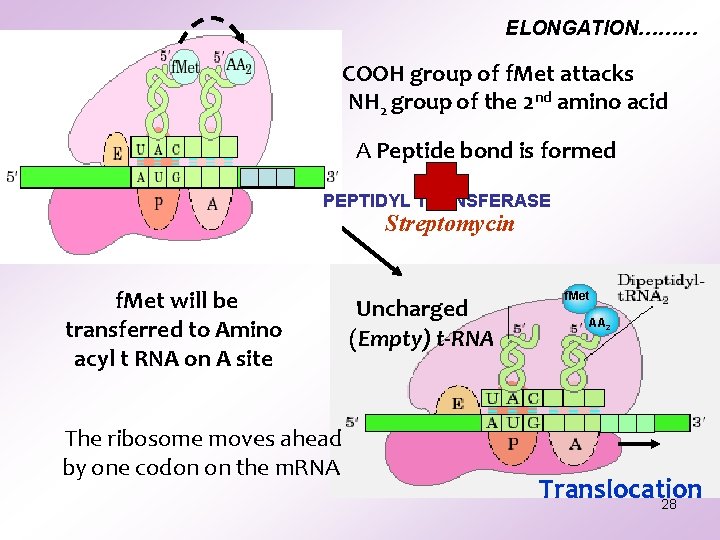
ELONGATION……… COOH group of f. Met attacks the NH 2 group of the 2 nd amino acid A Peptide bond is formed PEPTIDYL TRANSFERASE Streptomycin f. Met will be transferred to Amino acyl t RNA on A site The ribosome moves ahead by one codon on the m. RNA Uncharged (Empty) t-RNA f. Met AA 2 Translocation 28

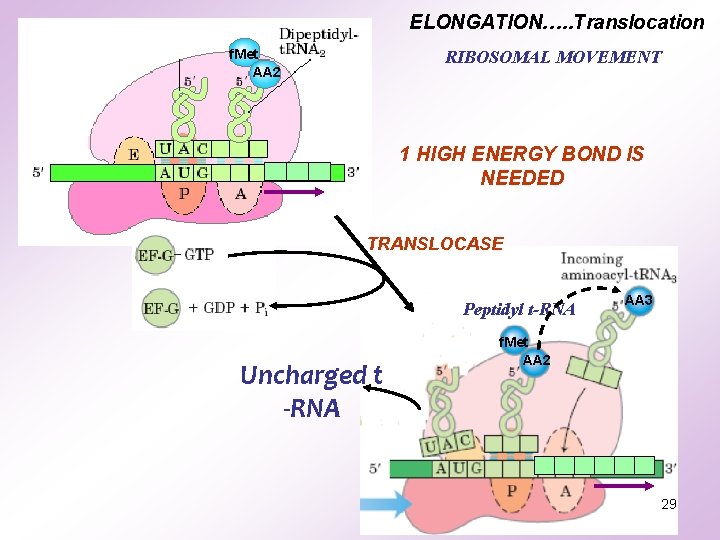
ELONGATION…. . Translocation f. Met AA 2 RIBOSOMAL MOVEMENT 1 HIGH ENERGY BOND IS NEEDED TRANSLOCASE Peptidyl t-RNA Uncharged t -RNA AA 3 f. Met AA 2 29

In elongation : Elongation ØEntry of next amino acyl-t-RNA at A-site ØFormation of a peptide bond (di-peptide) at the A-site ØTranslocation of the ribosome on the m-RNA to the next codon. ØExit of uncharged t. RNA from E-site. ØEntry of next amino acyl-t-RNA at A-site. 30

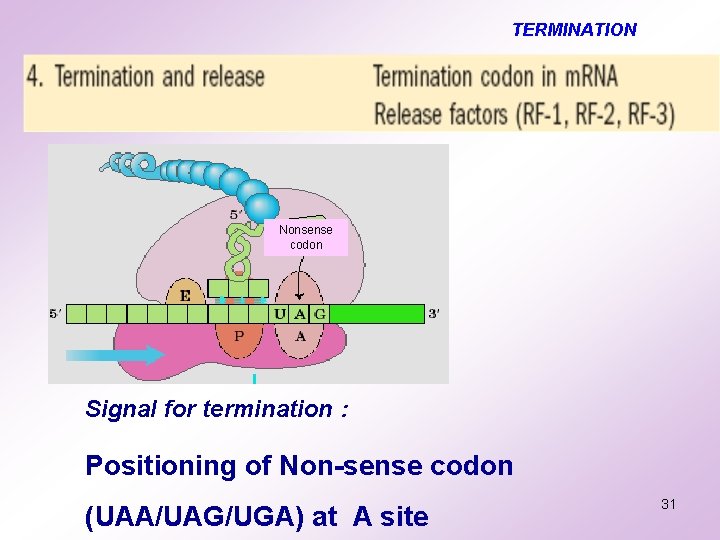
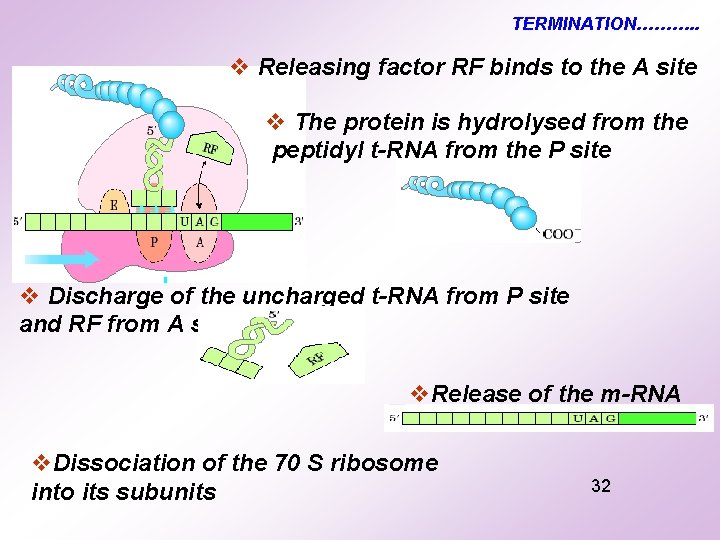
TERMINATION Nonsense codon Signal for termination : Positioning of Non-sense codon (UAA/UAG/UGA) at A site 31

TERMINATION………. . v Releasing factor RF binds to the A site v The protein is hydrolysed from the peptidyl t-RNA from the P site v Discharge of the uncharged t-RNA from P site and RF from A site v. Release of the m-RNA v. Dissociation of the 70 S ribosome into its subunits 32

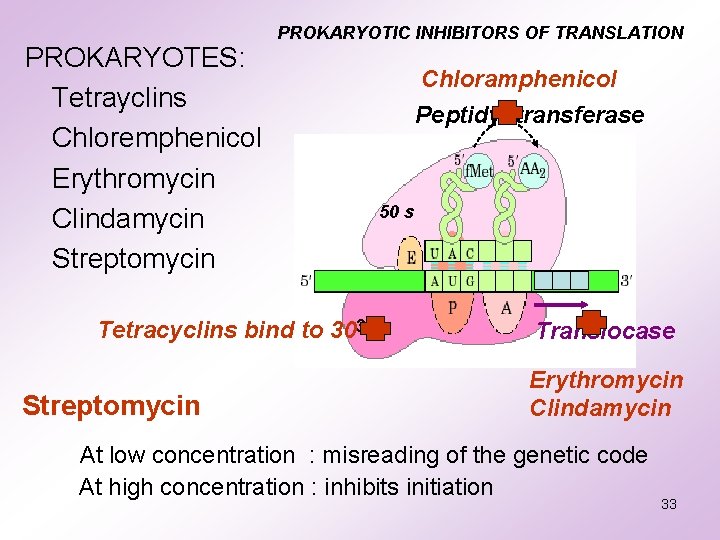
PROKARYOTES: Tetrayclins Chloremphenicol Erythromycin Clindamycin Streptomycin PROKARYOTIC INHIBITORS OF TRANSLATION Chloramphenicol Peptidyl transferase 50 s Tetracyclins bind to 3030 Ss Streptomycin Translocase Erythromycin Clindamycin At low concentration : misreading of the genetic code At high concentration : inhibits initiation 33

EUKARYOTIC INHIBITORS OF TRANSLATION Diphtheria toxin Produced by Corynebacterium Diphtheri. Attaches ADP to EF 2 & inactivates it. Ricin : Toxic protein from castor beans Diphtheria toxin Inactivates 60 S subunit. Ricin. of Eukaryotic ribosomes Puromycin: similar to aminoacyl-t. RNA Puromycin terminates Cycloheximide protein synthesis by binding to A site of ribosome. Cycloheximide: Blocks the peptidyl transferase of the 80 S ribosome and not the 70 S ribosome 34

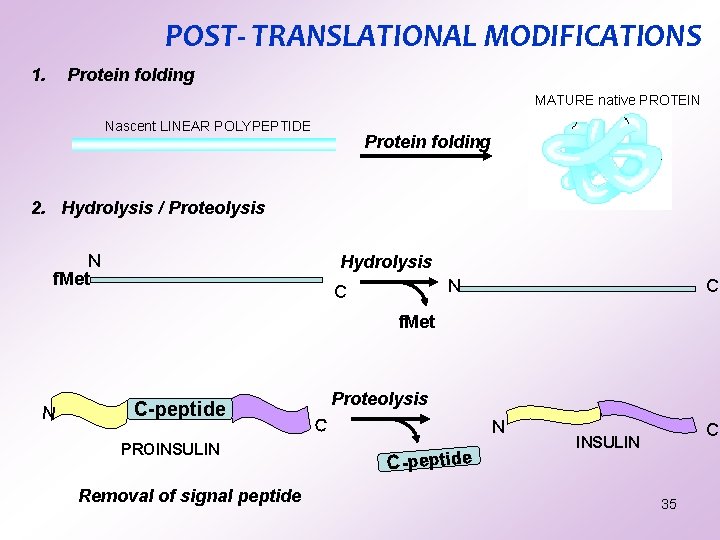
POST- TRANSLATIONAL MODIFICATIONS 1. Protein folding MATURE native PROTEIN Nascent LINEAR POLYPEPTIDE Protein folding 2. Hydrolysis / Proteolysis N f. Met Hydrolysis N C C f. Met N C-peptide PROINSULIN Removal of signal peptide Proteolysis C N C-peptide C INSULIN 35

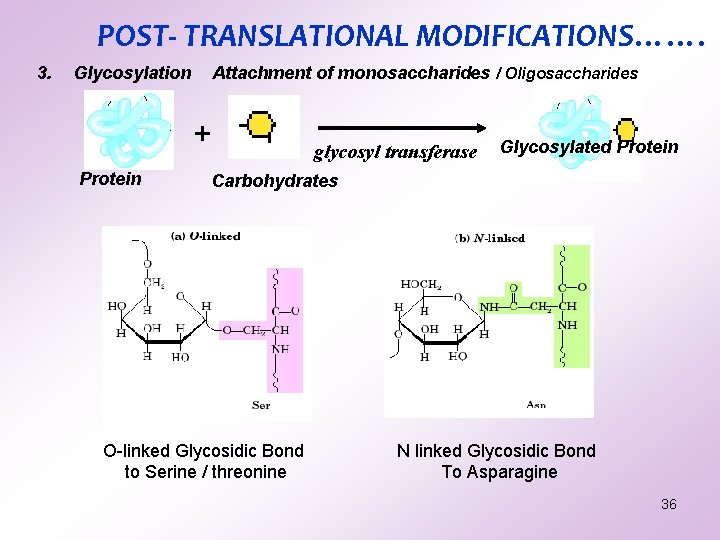
POST- TRANSLATIONAL MODIFICATIONS……. 3. Glycosylation Attachment of monosaccharides / Oligosaccharides + Protein glycosyl transferase Glycosylated Protein Carbohydrates O-linked Glycosidic Bond to Serine / threonine N linked Glycosidic Bond To Asparagine 36

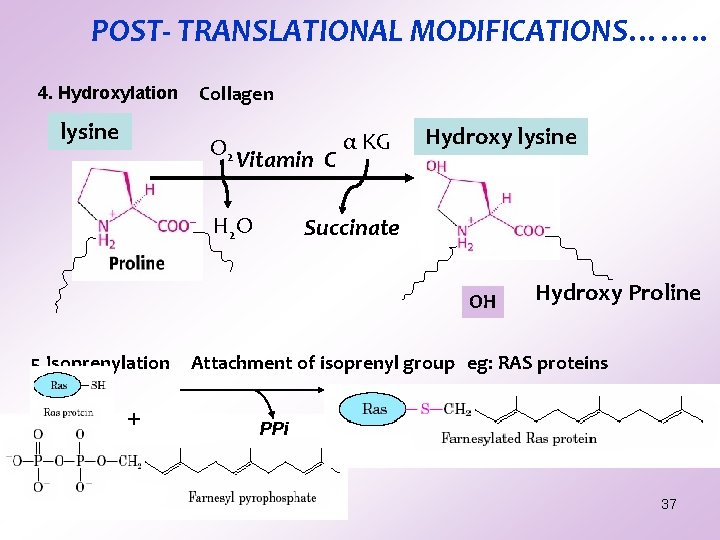
POST- TRANSLATIONAL MODIFICATIONS……. . 4. Hydroxylation lysine Collagen O 2 Vitamin C H 2 O α KG Hydroxy lysine Succinate OH 5. Isoprenylation + Hydroxy Proline Attachment of isoprenyl group eg: RAS proteins PPi 37

POST- TRANSLATIONAL MODIFICATIONS…… 6. Phosphorylation : Phosphate group is attached to the hydroxyl group of Serine, Threonine and Tyrosine. Phosphorylation – by protein kinases Eg : Glycogen synthase 7. Acetylation / Methylation : Attachment of acetyl group Eg : Histone proteins 8. Carboxylation. Eg : Gamma carboxylation of coagulation factors 2, 7, 9, 10 9. Addition of Prosthetic group : Eg : Biotin to Carboxylase enzymes 10. Formation of di-sulfide bridges : Eg : common in proteins to be exported from the ce 11. Modification of N- and C-teminal residues of the protein 38

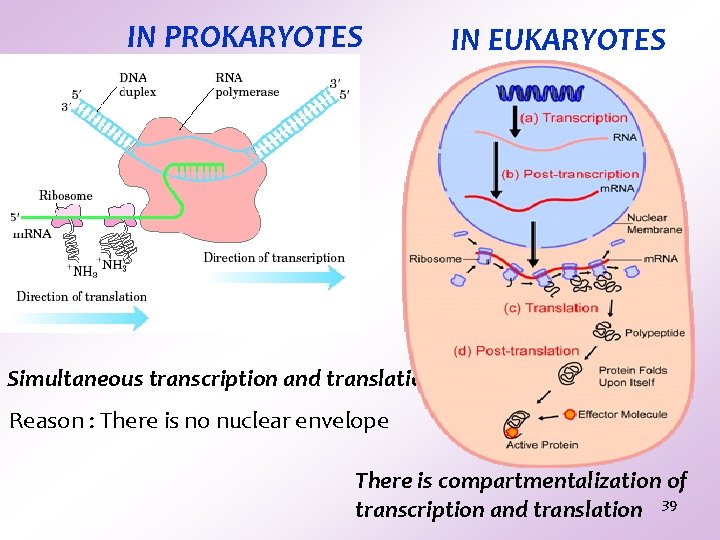
IN PROKARYOTES IN EUKARYOTES Simultaneous transcription and translation Reason : There is no nuclear envelope There is compartmentalization of transcription and translation 39

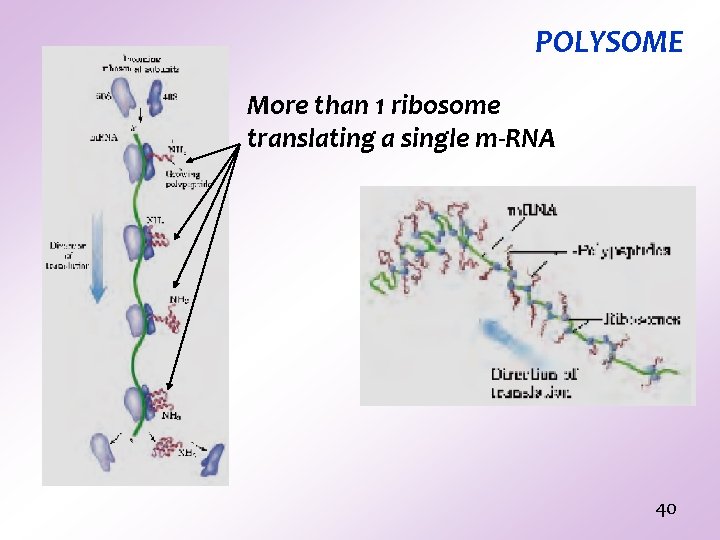
POLYSOME More than 1 ribosome translating a single m-RNA 40

41

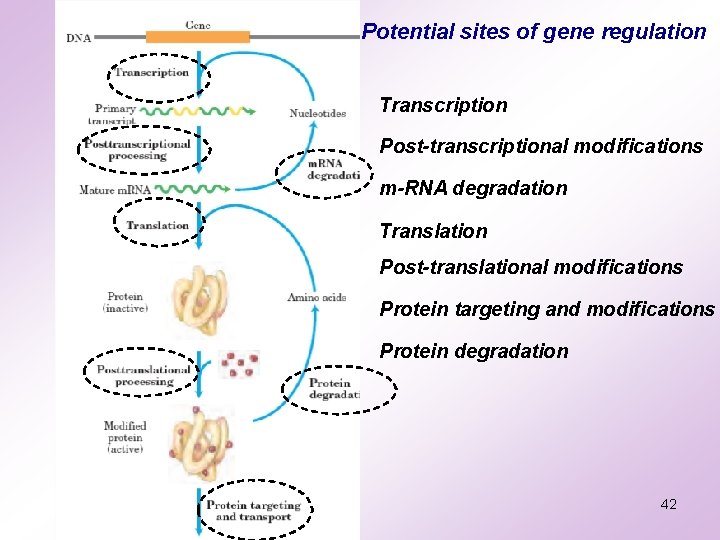
Potential sites of gene regulation Transcription Post-transcriptional modifications m-RNA degradation Translation Post-translational modifications Protein targeting and modifications Protein degradation 42

Types of genes & gene expression: Unvarying expression of genes is called constitutive expression genes which are expressed at the same rate through out the life eg : Hexokinase of the cell - House-keeping genes Cellular levels of gene products rise and fall in response to molecular signals – Regulated gene expression Gene products which in response to molecular signals – Inducible genes Process – Induction Eg : High levels of DNA damage induces the enzymes of DNA repair Gene product which in response to molecular signals – Repressible genes Process – Repression Eg : in bacteria presence of Tryptophan represses the enzymes which synthesize 43 Tryptophan.

Transcription is mediated and regulated by DNA – protein interactions DNA - RNA Polymerase interaction PROMOTER Is a site on the DNA close to the site where the transcription begins Types of promoters : TATA box, Pribnow box, Protein factors which bind to the promoter: 3 types Specificity factors : Alter specificity of the enzyme for the promoter Eg : Sigma factor for RNA polymerase Activators : enhance the RNA polymerase –promoter interaction Repressors : impede access of the enzyme to the promoter OPERATORS : specific location on the DNA where the repressors bind 44

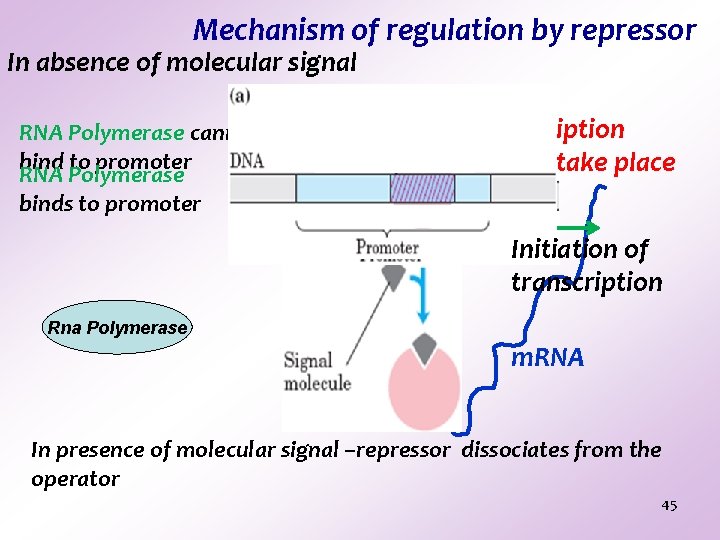
Mechanism of regulation by repressor In absence of molecular signal RNA Polymerase cannot bind to promoter RNA Polymerase binds to promoter REPRESSOR Transcription cannot take place Initiation of transcription Rna Polymerase m. RNA In presence of molecular signal –repressor dissociates from the operator 45

Metabolism of Prokaryotes Normal - source of carbon is Glucose Glycolysis Energy In absence of Glucose Other sugars : Lactose, Arabinose converted to intermediates of glycolysis More enzymes are needed for these extra steps 46

In Bacteria : OPERON Genes of PROTEINS that participate in a set of related processes are clustered together on the chromosome. • They have a common operator • They are transcribed together as a polycistronic m. RNA Gene cluster + Operator + other elements which help in regulation - OPERON 47

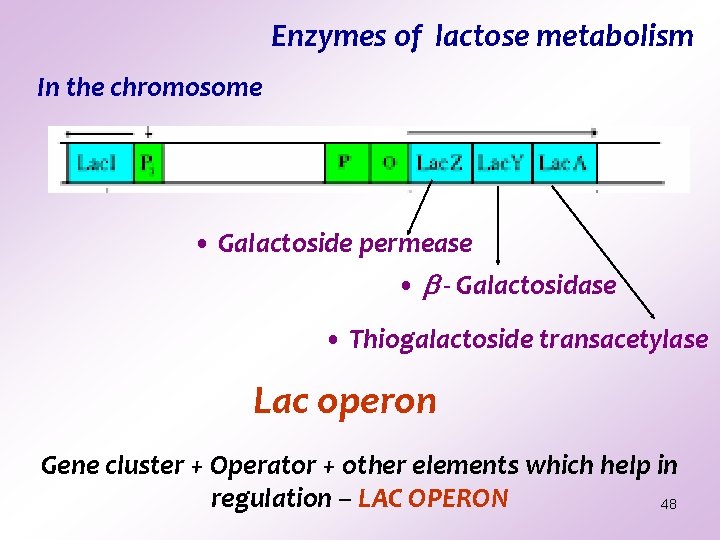
Enzymes of lactose metabolism In the chromosome • Galactoside permease • - Galactosidase • Thiogalactoside transacetylase Lac operon Gene cluster + Operator + other elements which help in regulation – LAC OPERON 48

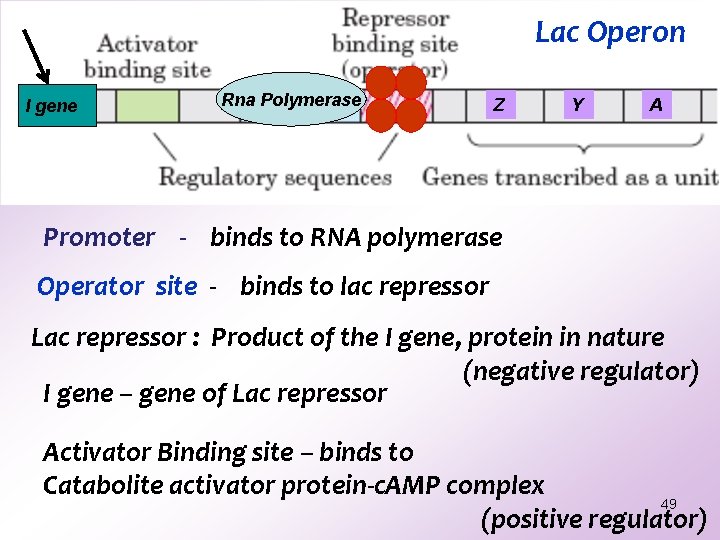
Lac Operon I gene Rna Polymerase Z Y A Promoter - binds to RNA polymerase Operator site - binds to lac repressor Lac repressor : Product of the I gene, protein in nature (negative regulator) I gene – gene of Lac repressor Activator Binding site – binds to Catabolite activator protein-c. AMP complex 49 (positive regulator)

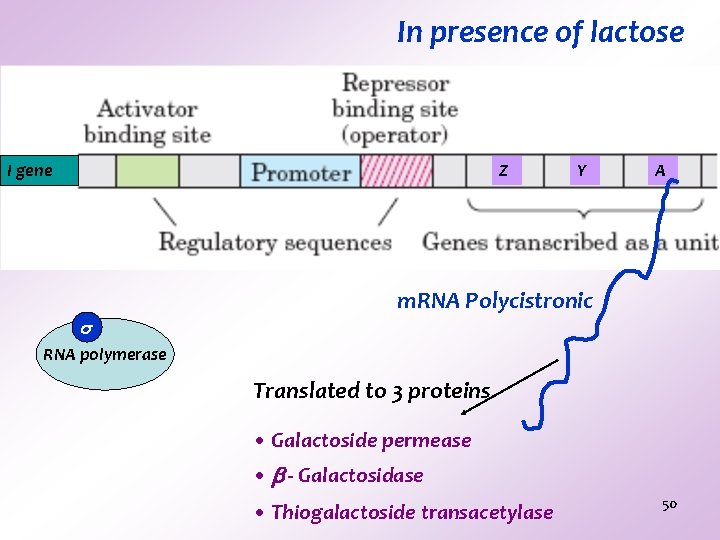
In presence of lactose I gene Z Y A m. RNA Polycistronic RNA polymerase Translated to 3 proteins • Galactoside permease • - Galactosidase • Thiogalactoside transacetylase 50

• When glucose and lactose both are present • Glucose is used up first • The Genes of lactose metabolism are not transcribed 51

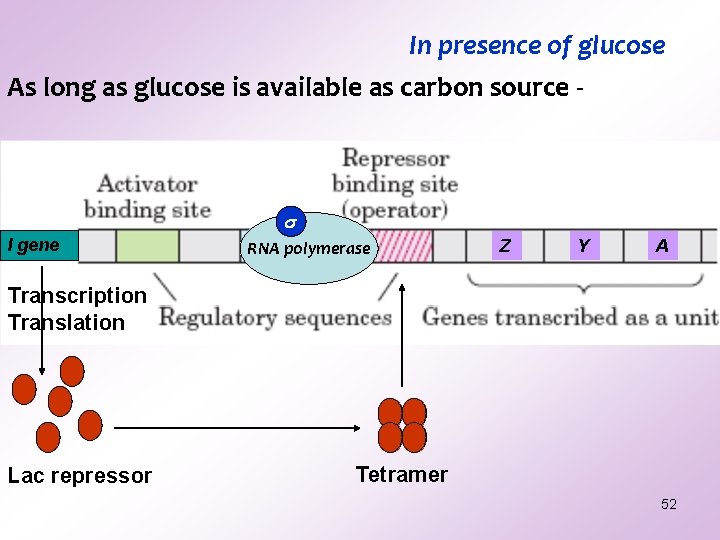
In presence of glucose As long as glucose is available as carbon source - I gene RNA polymerase Z Y A Transcription Translation Lac repressor Tetramer 52

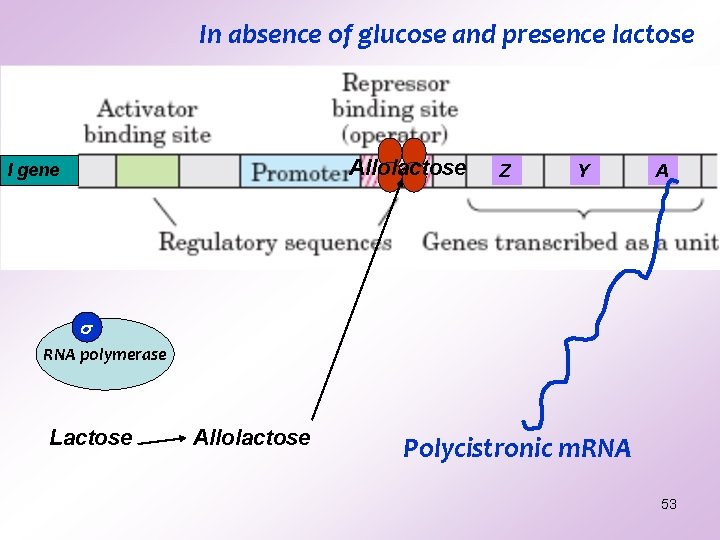
In absence of glucose and presence lactose Allolactose I gene Z Y A RNA polymerase Lactose Allolactose Polycistronic m. RNA 53

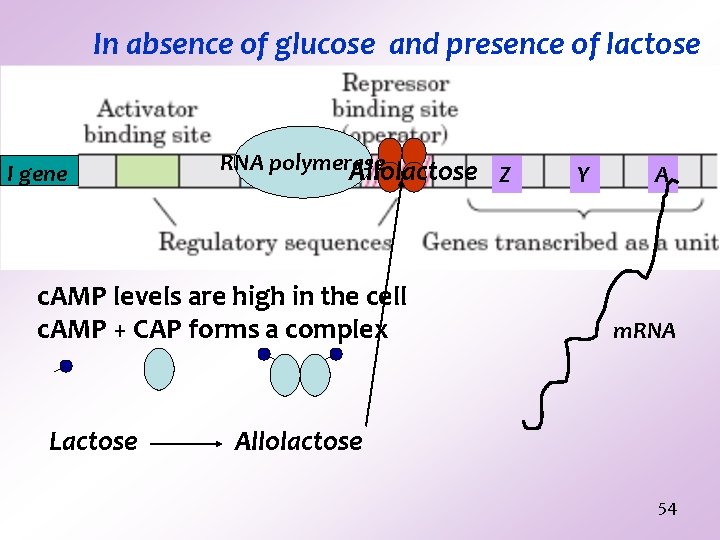
In absence of glucose and presence of lactose I gene RNA polymerase Allolactose c. AMP levels are high in the cell c. AMP + CAP forms a complex Lactose Z Y A m. RNA Allolactose 54

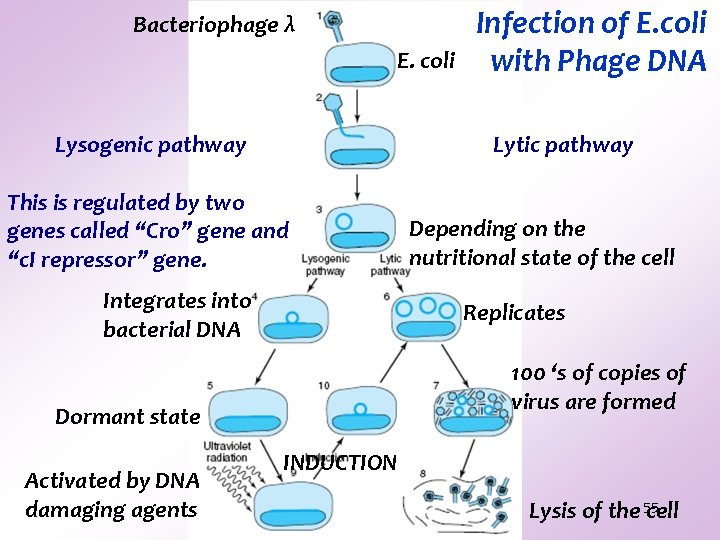
Bacteriophage λ E. coli Lysogenic pathway Lytic pathway This is regulated by two genes called “Cro” gene and “c. I repressor” gene. Integrates into bacterial DNA Depending on the nutritional state of the cell Replicates 100 ‘s of copies of virus are formed Dormant state Activated by DNA damaging agents Infection of E. coli with Phage DNA INDUCTION Lysis of the 55 cell

Regulation of gene expression in eukaryotes 56

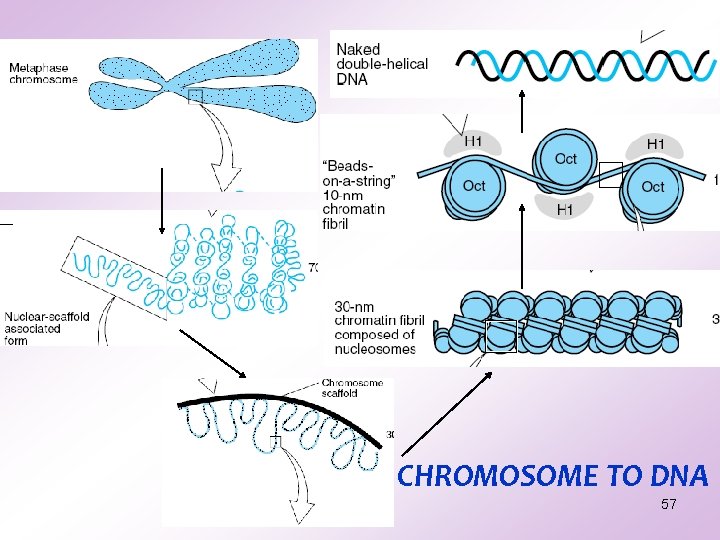
CHROMOSOME TO DNA 57

Special features of eukaryotic chromatin remodelling • Chromatin structure Heterochromatin – Euchromatin – • densely packed • Less condensed regions • transcriptionally inactive • Nucleosomes are absent • 10% of the total DNA • transcriptionally active • Associated with centromeres • Sensitive to DNAse enzyme acetylation De-acetylation

Special features of eukaryotic chromatin remodelling 1. Histone acetylation / de-acetylation – chromatin remodelling Acetylation of histones disrupts nucleosome structure and makes it transcriptionally active. De-acetylation - makes it transcriptionally inactive. 2. Methylation of deoxycytidine residues prevents active transcription of genes. 59

Regulation by binding to other molecules 3. Binding of specific transcription factors Disrupts nucleosome structure and increases access to transcription elements 60

Regulation by binding to other molecules 4. DNA elements which modulate eukaryotic genes SILENCERS : sequences on the DNA that the rate of transcription on binding to signaling molecules. ENHANCERS: sequences on the DNA that the rate of transcription on binding to signaling molecules. HORMONE RESPONSE ELEMENTS: DNA sequences which enhance or decrease the gene expression after binding to hormones. 61

• PROPERTIES of DNA elements which modulate eukaryotic genes • Located upstream or downstream to the start site. • Located long away from start site • Oriented in either way • Bind with transcription factors. • Locus control regions (LCR’s) on the DNA Associated with a group of proteins and regulates transcription of genes 62

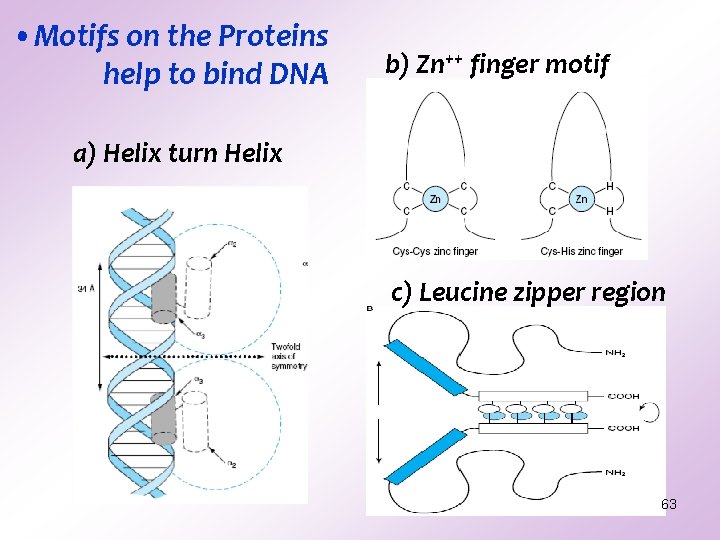
• Motifs on the Proteins help to bind DNA b) Zn++ finger motif a) Helix turn Helix c) Leucine zipper region 63

Differences in gene regulation in Prokaryotes and Eukaryotes Regulation can occur at 3 levels • Transcription • Nuclear m-RNA processing • M-RNA stability 64

REGULATION AT m. RNA PROCESSING 1. Alternative start sites: Ex: Mouse amylase Myosin light chain Rat glucokinase 2. Alternative Polyadenylation sites: Ig M heavy chain – 2700 bases ----membrane bound antibody Ig M heavy chain – 2400 bases ----secretory antibody 65

3. Alternative Splicing Ex: Seven tropomyosin m. RNA molecules in different tissues 66

In addition • Gene amplification • Gene rearrangement • Gene amplification : No. of copies of the gene is increased. Drug resistance is because of several 100 fold increase in the copies of target genes y gene amplification in response to heavy doses of drugs 67

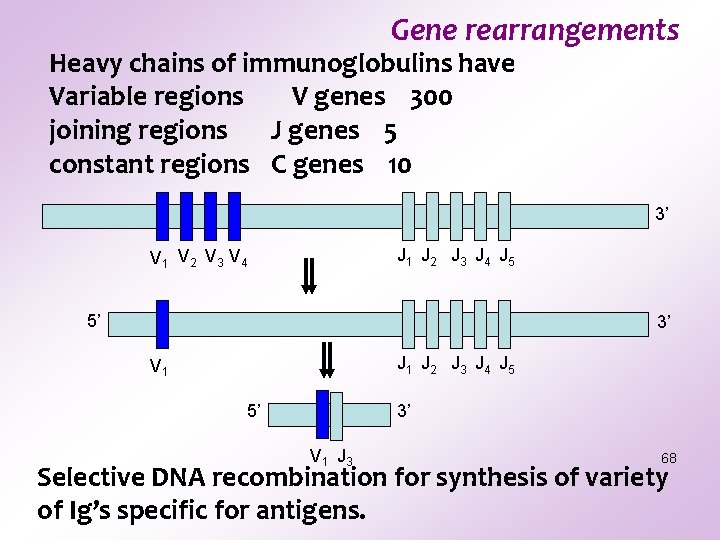
Gene rearrangements Heavy chains of immunoglobulins have Variable regions V genes 300 joining regions J genes 5 constant regions C genes 10 3’ J 1 J 2 J 3 J 4 J 5 V 1 V 2 V 3 V 4 5’ 3’ J 1 J 2 J 3 J 4 J 5 V 1 5’ 3’ V 1 J 3 68 Selective DNA recombination for synthesis of variety of Ig’s specific for antigens.

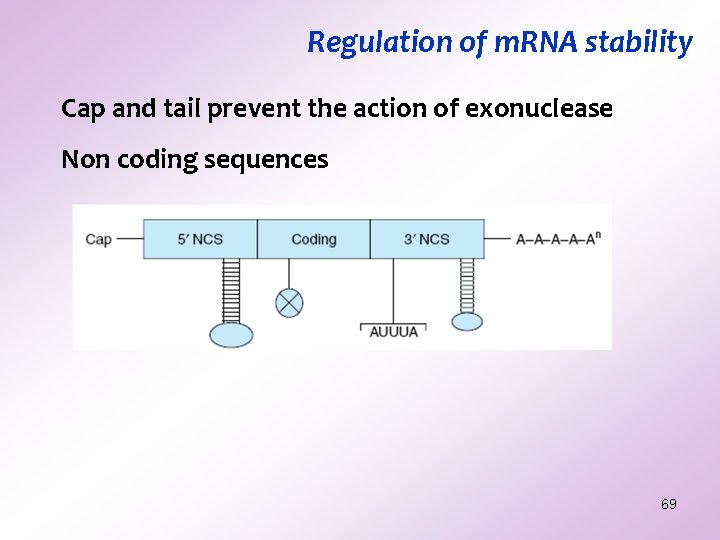
Regulation of m. RNA stability Cap and tail prevent the action of exonuclease Non coding sequences 69

Thank you 70