1 Central Dogma Review of Transcription and Translation







- Slides: 13

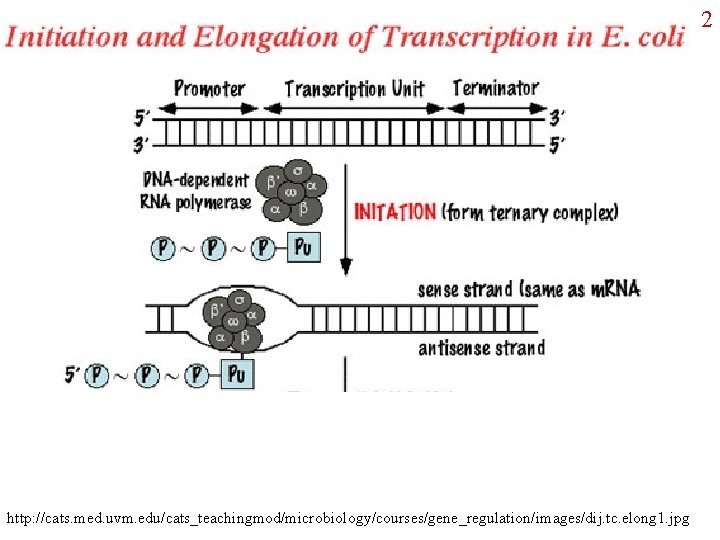
1 Central Dogma: Review of Transcription and Translation in bacteria

2 http: //cats. med. uvm. edu/cats_teachingmod/microbiology/courses/gene_regulation/images/dij. tc. elong 1. jpg

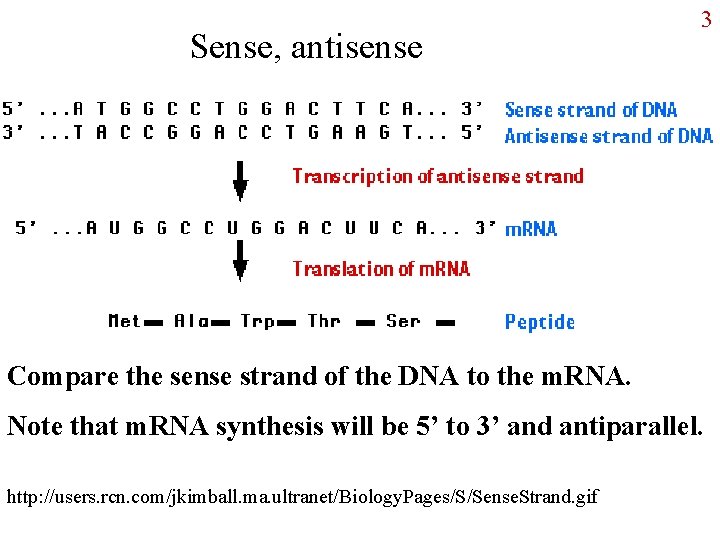
Sense, antisense 3 Compare the sense strand of the DNA to the m. RNA. Note that m. RNA synthesis will be 5’ to 3’ and antiparallel. http: //users. rcn. com/jkimball. ma. ultranet/Biology. Pages/S/Sense. Strand. gif

The Process of Transcription-2 4 • RNA synthesis continues (Elongation), only one DNA strand (template) is transcribed. • RNA nucleotides, complementary to bases on DNA strand, are connected to make m. RNA • Termination: must be a stop sign, right? – In bacteria, hairpin loop followed by run of U’s in the RNA. Of course, the DNA must code for complementary bases and a run of A’s. See next. Most common. OR – Termination factor “rho”. Enzyme. Forces RNA polymerase off the DNA.

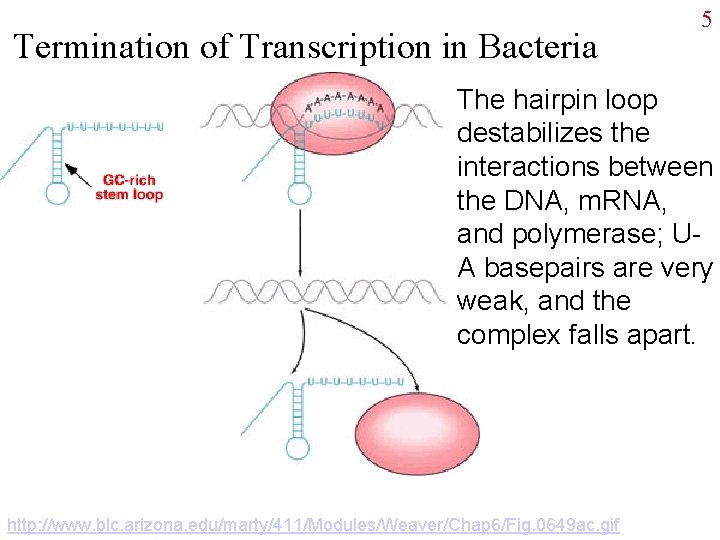
Termination of Transcription in Bacteria 5 The hairpin loop destabilizes the interactions between the DNA, m. RNA, and polymerase; UA basepairs are very weak, and the complex falls apart. http: //www. blc. arizona. edu/marty/411/Modules/Weaver/Chap 6/Fig. 0649 ac. gif

Transcription in prokaryotes • As m. RNA is made, it is ready to use. • Info from more than one gene is typically found on one m. RNA molecule. • Simpler process than in eukaryotes – no introns to remove – no cap or poly-A tail – no nuclear membrane to transport through • Transcription is expensive: each NTP leaves behind 2 Pi; like spending 2 ATP for every base used. 6

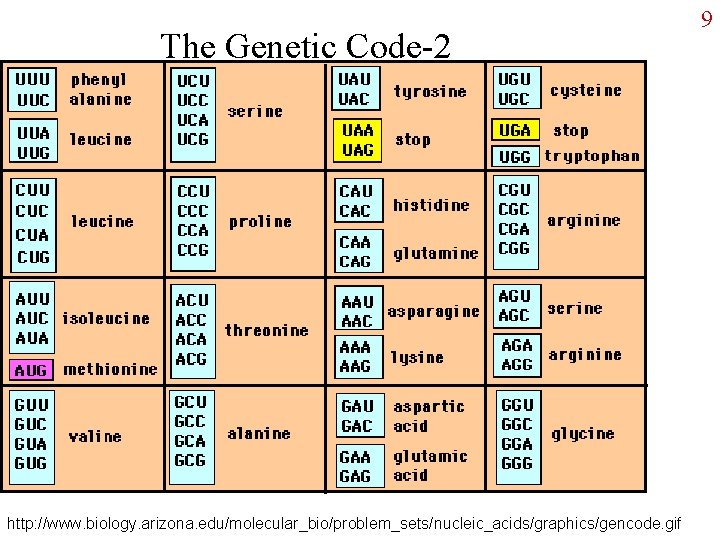
The Genetic Code 7 • Four bases taken how many at a time? Need to code for 20 different amino acids. – Each base = 1 amino acid: only 4 – Every 2 bases = 1 a. a. : 16 combinations, 4 short. – Every 3 bases: 64 combinations, enough. • Every 3 bases of RNA nucleotides: codon – Each codon is complementary to 3 bases in one strand of DNA

Properties of the Genetic Code • Code is unambiguous: 1 codon always specifies only 1 amino acid. • Code is degenerate: although unambiguous, an amino acid can be coded for by more than one codon. • Punctuated: certain codons specify “start” and “stop”. • Universal: by viruses, both prokaryotic domains, and eukaryotes (except for some protozoa, mitochondria). • Ordered: similar codons specify the same amino acid; see especially the 1 st two bases in the codon. 8

The Genetic Code-2 http: //www. biology. arizona. edu/molecular_bio/problem_sets/nucleic_acids/graphics/gencode. gif 9

Bacterial ribosomes 10 • Prokaryotic ribosomes are 70 S; eukaryotic are 80 S – S is Svedberg unit, how fast a particle travels during centrifugation. Affected by both mass and shape. • Large subunit: 50 S – 33 polypeptides, 5 S RNA, 23 S RNA • Small subunit: 30 S – 21 polypeptides, 16 S RNA • Note that 30 + 50 is not 70 • Ribosome structure and differences between prokaryotes and eukaryotes are important. – r. RNAs important in taxonomy to be discussed later – Differences are the basis for success of many antibiotics

Translation 11 • Literally, information translated from language of nucleotides to that of amino acids • Ribosomes (large and small subunits), m. RNA, t. RNAs, amino acids, and source of energy. – And various protein factors • Ribosomes attach to m. RNA, t. RNAs read codons, match amino acid to codon and ribosome connects amino acids to make proteins. • m. RNA has start codon AUG and stop codons. • Look for animations on line • http: //www. ncc. gmu. edu/dna/ANIMPROT. htm; http: //www. stolaf. edu/people/giannini/flashanimat/molgenetics/translation. swf

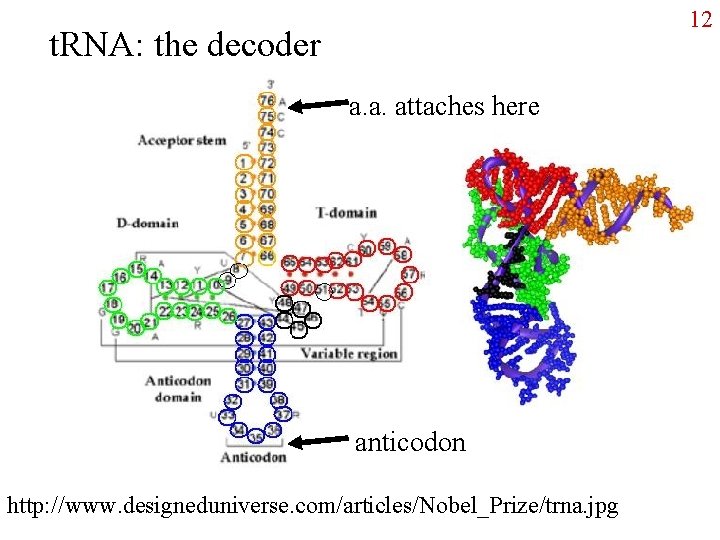
12 t. RNA: the decoder a. a. attaches here anticodon http: //www. designeduniverse. com/articles/Nobel_Prize/trna. jpg

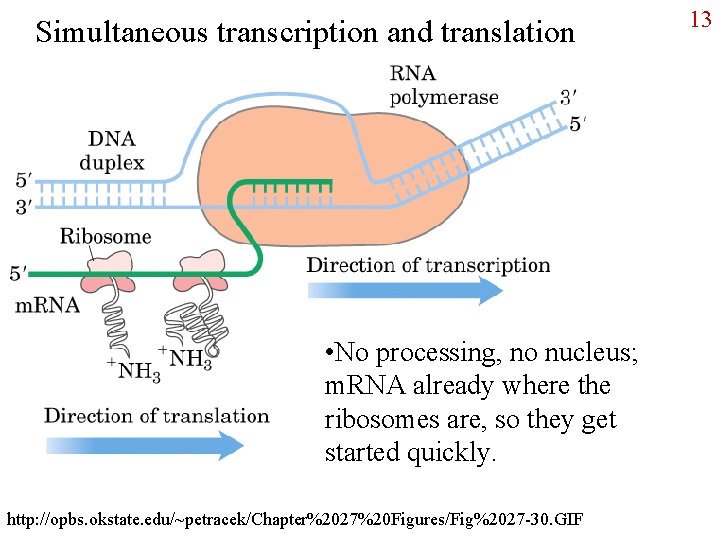
Simultaneous transcription and translation • No processing, no nucleus; m. RNA already where the ribosomes are, so they get started quickly. http: //opbs. okstate. edu/~petracek/Chapter%2027%20 Figures/Fig%2027 -30. GIF 13