1 Bioinformatics Principles Jong Bhak KOGIC UNIST Ulsan

1 Bioinformatics Principles 박종화 Jong Bhak 朴鍾和 KOGIC UNIST Ulsan Korea jongbhak@gmail. com 20160509

Bio 감사의말 • Researchers who are honest and passionate in doing science • People who support scientific research by paying tax • MRC, Harvard, KAIST, KOBIC, TBI & Genome Research Foundation. 테라젠 고진업대표이사. • NCC, 이연수, 이진수박사, • 국가참조표준센터(채균식, 김창근박사) • • 해양연 이정현박사와 동료들 한양대학교 (류성언교수, 김덕수교수, 고인송교수) UNIST, (조무제, 정무영 총장), BME교수, 지역지원자들 UNIST students
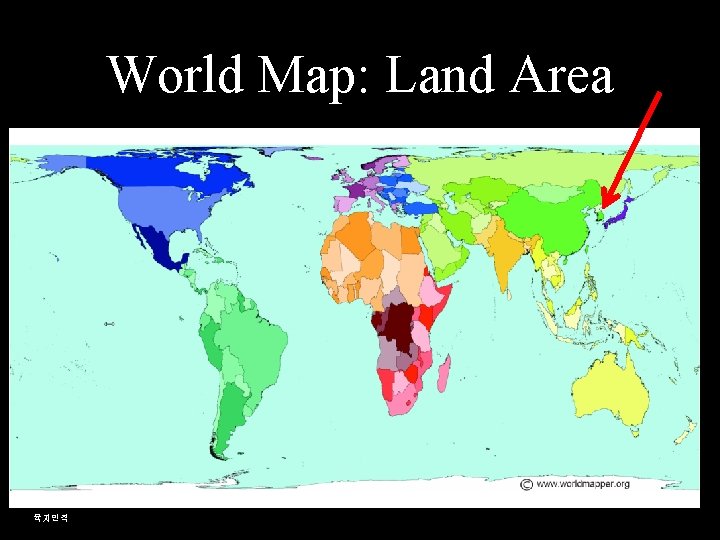
World Map: Land Area 육지면적
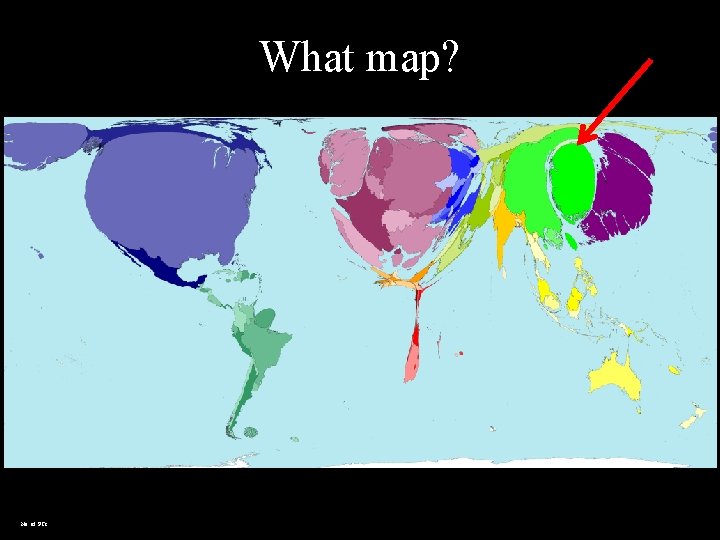
What map? No. of PCs
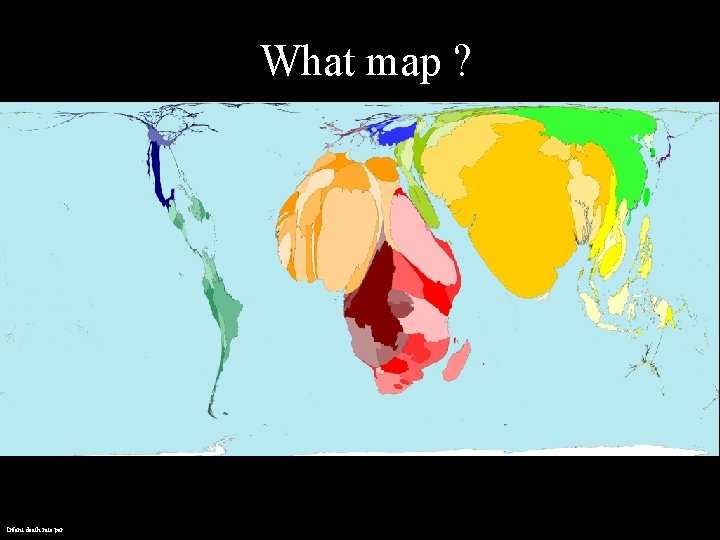
What map ? Infant death rate per

Bioinformatics Is about Mapping X is IEP Y is Size Old map Is not Accurate. However It helps People to Explore. Gangnido 1402 Data

Lesson • Depending on the parameters we use, the world and problems can be interpreted differently. 2021 -10 -18 • Bioinformatics is mapping the biological universe using certain bioinformatics parameters 7

First principles first • Biological systems are within the universe 2021 -10 -18 • It is necessary to understand the universe to understand life 8

The origin of the universe 2021 -10 -18 9
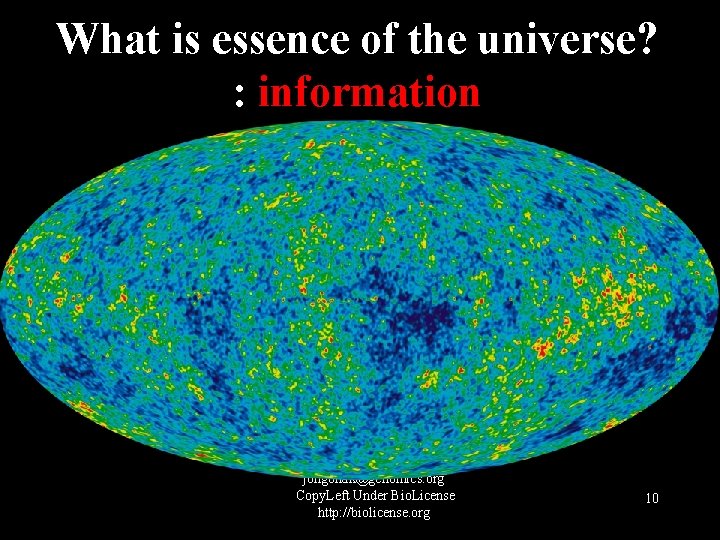
What is essence of the universe? : information jongbhak@genomics. org Copy. Left Under Bio. License http: //biolicense. org 10

Assumption/Hypothesis: • The essence of the universe is switching. The existences are the simulatenous instances of essence. 2021 -10 -18 11

Assumption/Hypothesis: • The fundamental elements of the universe are of information. • Physical objects are representations, reflections, or virtuals of information. • Basic entities are of inforamtion: energy/matters, space, and time are derivatives of information. 2021 -10 -18 12

Assumption/Hypothesis: • The physical reality is the product of information. • The physical world is numerical and mathematical representation of information. • Information precedes three dimentional physical world. • The universe is perfectly computable as it is an instance of computing. 2021 -10 -18 13

So, what is the universe? • The universe is the largest set of switches 2021 -10 -18 14

“Para-Programmed” Meta-Programmed Universe? 15 jongbhak@genomics. org Copy. Left Under Bio. License

Physical universe is an instance of the informational universe 2021 -10 -18 16
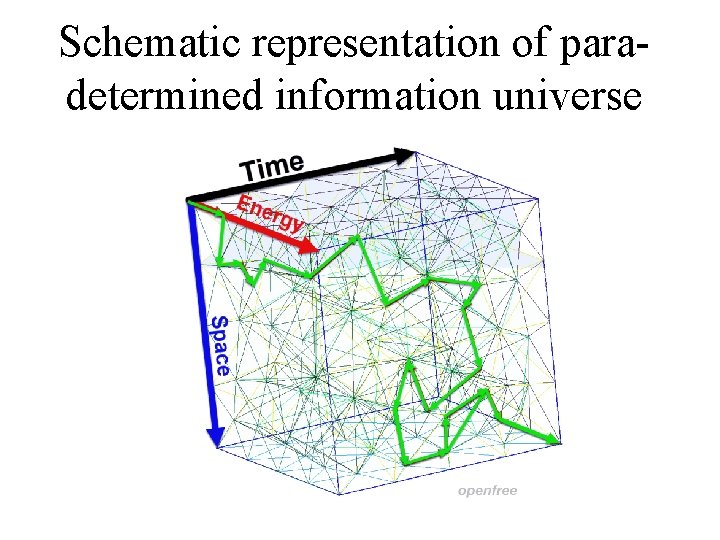
Schematic representation of paradetermined information universe 2021 -10 -18 17
What is Life? Life is a set of switches 18 jongbhak@genomics. org Copy. Left Under Bio. License
Switching State Change • Switching is the result of state change – The concept of time invented (Dynamics) jongbhak@genomics. org Copy. Left Under Bio. License 19
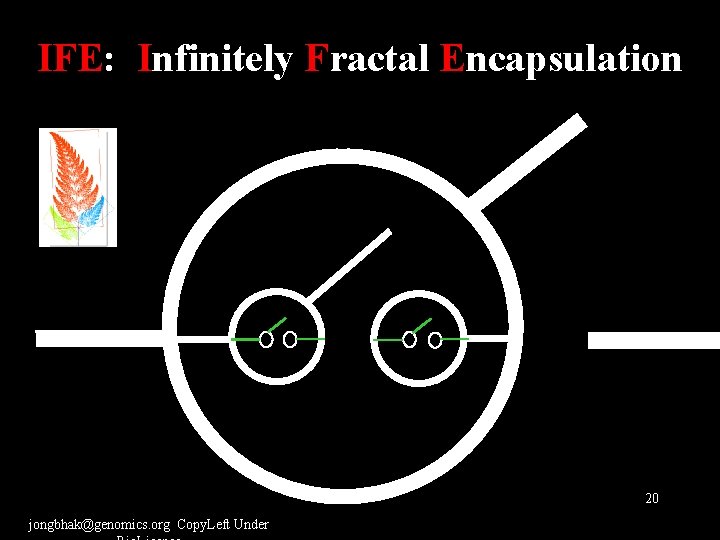
IFE: Infinitely Fractal Encapsulation 20 jongbhak@genomics. org Copy. Left Under
Information Hierarchy 2021 -10 -18 21

Reproducing chemicals Human • We could be all computers • The Earth is a gigantic computer jongbhak@genomics. org Copy. Left Under Bio. License 22

Fundamentals for bioinformatics • Required: Thoughts on Philosophy, Society, Science, and Biology in general.

Life is Complex(? ) • Complex Homo sapiens live in layers of complex systems: multi-cellular, multi-organismal, multisocietal and multi-cultural. – Similar patterns occur again in different layers? • The main problem is what the general patterns are in the infinite number of biological layers. • Bioinformatics is the Study of Complexity

The core of Biology • Biology is an information science over energy metabolism • Two important things: – ENERGY and INFORMATION • Johann G. Mendel’s (1822 -1884) genetic work on peas (Pisum) bioinformatic analysis, modelling and prediction. • Perhaps he is the first well-known classical bioinformatist in history.

Evolution, • Charles Darwin’s is one of very few general principle biology has. The process of evolution is often applied to technology nowadays. Virtually every aspect of Biological information processing is concerned about evolution. Evolutionary theories also provide third element in Biology: time. • Bioinformatics deals with evolution in Biology, all the time. 2021 -10 -18 [1809 - 1882] 26

Long Definition of Bioinformatics Is a discipline of Science that analyses, seeks understanding and models Life as an Information Processing phoenomenon over Energy with methods derived from philosophy, mathematics and computer science using biological experimental data. - Jong Bhak, 2000

Short Definition of Bioinformatics • Bioinformatics is Biology is Bioinformatics - Jong Bhak, 2000 and

Brief history of Biology • • • • Darwin Mendel 3 D Proteins DNA model Sequencing(Sanger) Cloning Recombination Amplification Technologies Human Reference Genome Next Gen. Sequencing/Personal Genomics Diagnostics using Genomic data Synthetic Biology Genome Engineering Cancer Cure 2022 Aging Cure 2042
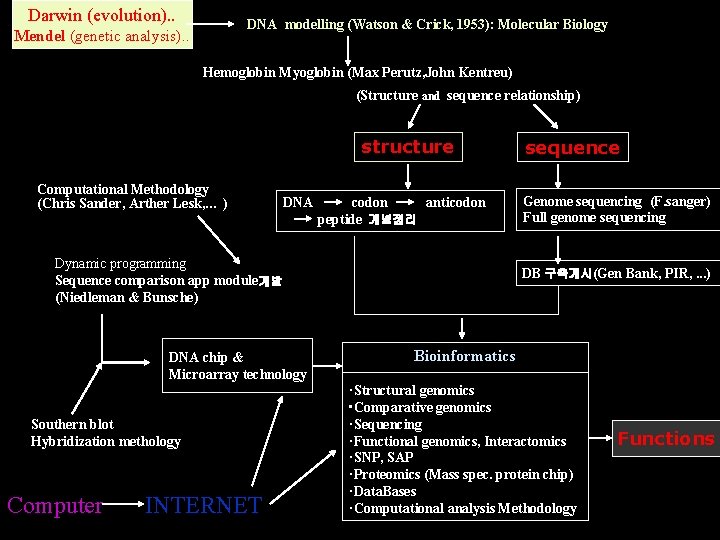
Darwin (evolution). . DNA modelling (Watson & Crick, 1953): Molecular Biology Mendel (genetic analysis). . Hemoglobin Myoglobin (Max Perutz, John Kentreu) (Structure and sequence relationship) structure Computational Methodology (Chris Sander, Arther Lesk, … ) DNA codon anticodon peptide 개념정리 Dynamic programming Sequence comparison app module개발 (Niedleman & Bunsche) DNA chip & Microarray technology Southern blot Hybridization methology Computer INTERNET sequence Genome sequencing (F. sanger) Full genome sequencing DB 구축개시(Gen Bank, PIR, . . . ) Bioinformatics ·Structural genomics • Comparative genomics ·Sequencing ·Functional genomics, Interactomics ·SNP, SAP ·Proteomics (Mass spec. protein chip) ·Data. Bases ·Computational analysis Methodology Functions

Bioinformatics in time • Last decades: heavily driven by structural studies such as protein folding problem and structural comparisons/classifications/molecular analyses. • A recent shift toward sequence, databases, software, computation, commercialization and functions of proteins. Mid 1990 s. – A leap of life : Bio. Internet early 1990 s. Life managed to connect humans as neurons did some time ago.

Bioinformatics in time • The most important contemporary problem: – Explaining complex systems of biology functionally and evolutionarily. Major fields of Bioinformatics: next page.

Major Domains of Bioinformatics • Sequence • Structure • Expression • Interaction • Function

Bioinformatics • Sequence – Genomics, Comparative Genomics • Structure – Structural Genomics, Structural Proteomics – Biophysics • Expression – Functional Genomics, Proteomics • Interaction – Proteomics, Interactomics • Function – Physiomics, Metabolomics

Major Parts of Bioinformatics: – Computing – (1) Structural studies, (2) sequence analysis, (3) molecular interactions (4) functional analysis of genes, proteins and their ligands (Large scale expression analysis: DNA chips, microarray ) – (5) Algorithm development ( Mathematical and physical calculation programs. Bioperl, Bio. Java, Bio. XML, Bio. Python, Bio. CPP, CGI programming ), Network and middleware programs. Bio. Infrastructure – (6) Database construction (Relational databases, Object oriented databases). Medical informatics – (7) large scale data mining (artificial intelligence approach), – (8) Complex systems and network analysis – (9) Various prediction methods. – (10) Visualization of large and complex data. – (11) Large computer systems construction (hardware) and administration. – (12) OS, Compiler, Microprocessor optimization for bioanalyses – (13) Socio-economic modelling of life – (14) neuronal and psychological description of complex organisms – (15) designing and engineering cells and organisms

Applied Fields of Bioinformatics • Sequencing related – Gene prediction, gene mapping, annotation, visualization • Genomics – Structural Genomics, – Functional Genomics (proteomics, interactomics) – Comparative genomics – SNP (single nucleotide polymorphism) , SAP (single amio…) • Proteomics • • – Mass spec, Protein Chip, Protein Interaction Interactomics (Network Biology) Complex systems (Network Biology) approach Neuroinformatics (neurological informatics) Medinformatics (medical informatics)
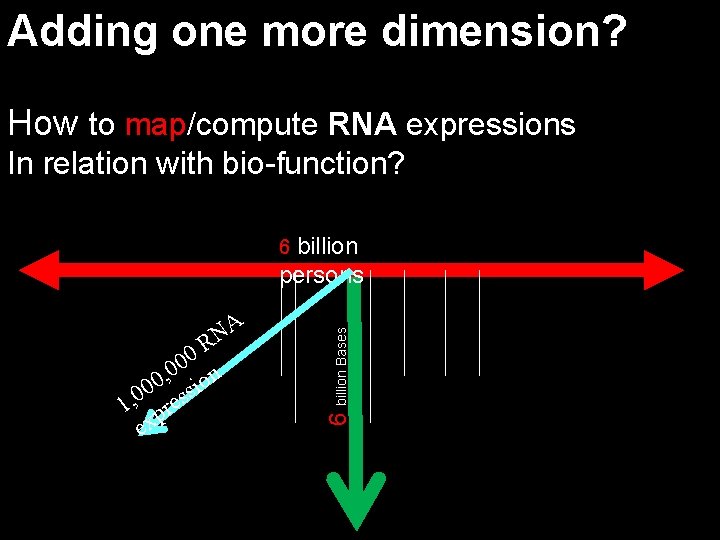
Adding one more dimension? How to map/compute RNA expressions In relation with bio-function? billion persons A N R 0 0 n 0 , 0 sio 0 0 1, pres ex 6 billion Bases 6
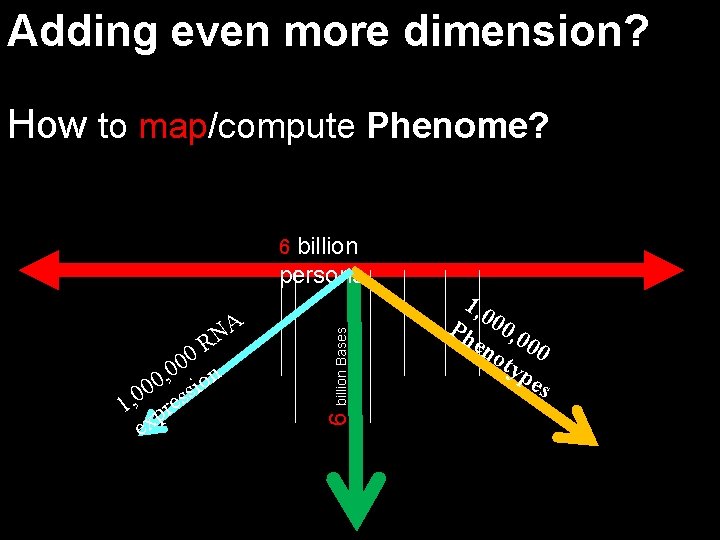
Adding even more dimension? How to map/compute Phenome? billion persons A N R 0 0 n 0 , 0 sio 0 0 1, pres ex 6 billion Bases 6 1, 0 Ph 00, 0 en 00 oty pe s
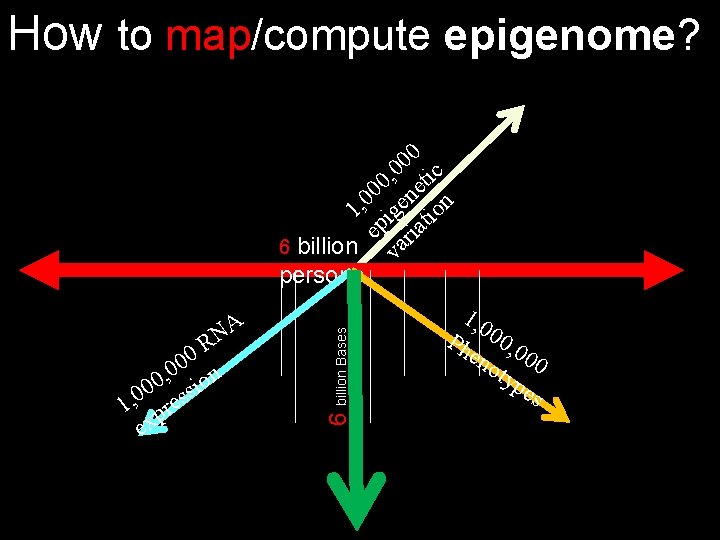
How to map/compute epigenome? A N R 0 0 n 0 , 0 sio 0 0 1, pres ex 6 billion Bases 0 0 , 0 tic 0 0 ne n 0 1, ige tio ep ria 6 billion va persons 1, 0 Ph 00, 0 en oty 00 pe s
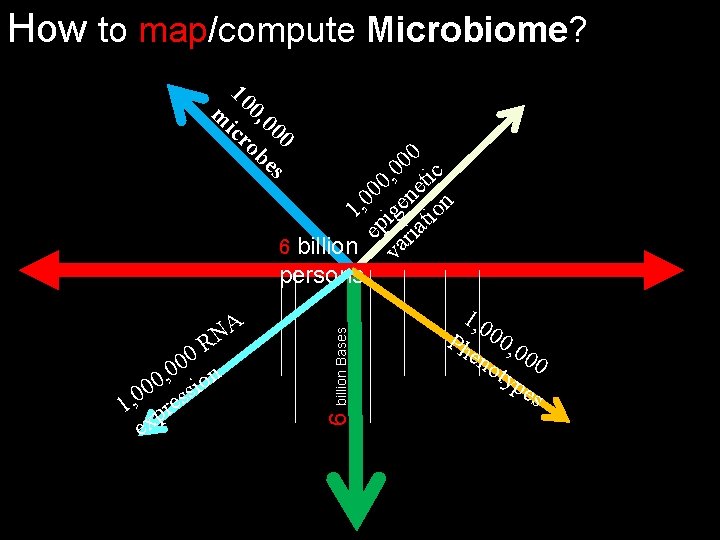
How to map/compute Microbiome? 10 m 0, 0 ic 00 ro be s A N R 0 0 n 0 , 0 sio 0 0 1, pres ex 6 billion Bases 0 0 , 0 tic 0 0 ne n 0 1, ige tio ep ria 6 billion va persons 1, 0 Ph 00, 0 en oty 00 pe s
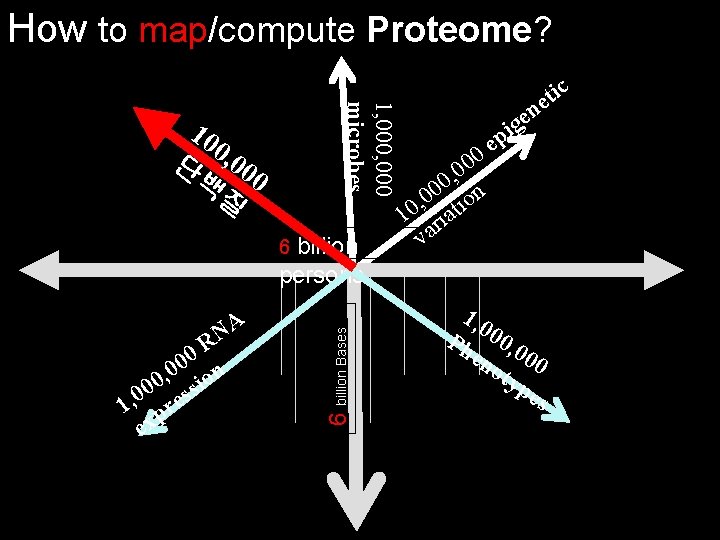
How to map/compute Proteome? 1, 000 microbes 10 0 단 , 00 백 0 질 billion persons A N R 0 0 , 0 ion 0 0 ss 0 , 1 pre ex 6 billion Bases 6 c i t e n e g 0 0 , 0 0 0 on 0 , 10 riati va i p e 1, 0 Ph 00, 0 en 00 oty pe s

Bioinformatic problems boil down to: • Representation of data.

Ways of representing Bio. Entities • • • Sequence Structure Expression levels Pathways Function • Networks

Computer • The universe is computable? 2021 -10 -18 44

Very Basic information for nonbiologists. • Elementary biological information on proteins etc. • Only for non-biologists!

Proteins • Proteins: The central processing molecules of life. (15% of the mass of the average person) • Minium 20 different kinds of amino acids: Alanine ala a CH 3 -CH(NH 2)-COOH Arginine arg r HN=C(NH 2)-NH-(CH 2)3 -CH(NH 2)-COOH Asparagine asn n H 2 N-CO-CH 2 -CH(NH 2)-COOH Aspartic acid asp d HOOC-CH 2 -CH(NH 2)-COOH Cysteine cys c HS-CH 2 -CH(NH 2)-COOH Glutamine gln q H 2 N-CO-(CH 2)2 -CH(NH 2)-COOH Glutamic acid glu e HOOC-(CH 2)2 -CH(NH 2)-COOH Glycine gly g NH 2 -COOH Histidine his h NH-CH=N-CH=C-CH 2 -CH(NH 2)-COOH Isoleucine ile i CH 3 -CH 2 -CH(CH 3)-CH(NH 2)-COOH Leucine leu l (CH 3)2 -CH-CH 2 -CH(NH 2)-COOH Lysine lys k H 2 N-(CH 2)4 -CH(NH 2)-COOH Methionine met m CH 3 -S-(CH 2)2 -CH(NH 2)-COOH Phenylalanine phe f Ph-CH 2 -CH(NH 2)-COOH Proline pro p NH-(CH 2)3 -CH-COOH Serine ser s HO-CH 2 -CH(NH 2)-COOH Threonine thr t CH 3 -CH(OH)-CH(NH 2)-COOH Tryptophan trp w Ph-NH-CH=C-CH 2 -CH(NH 2)-COOH Tyrosine tyr y HO-p-Ph-CH 2 -CH(NH 2)-COOH Valine val v (CH 3)2 -CH-CH(NH 2)-COOH http: //www. nyu. edu/pages/mathmol/library/life 1. html
Amino Acids (L-form)

Types of Amino Acids Amino acids can be grouped into 4 -5 different groups for Bioinformatic analysis. Most important distinctions: Hydrophobic and Hydrophilic groups Big side chain groups and Small side chain groups Cysteine for disulphide bonding. (well conserved) Proline structurally important Histidine important for switching • Aliphatic - alanine glycine isoleucine proline valine • Aromatic - phenylalanine tryptophan tyrosine • Acidic - aspartic acid glutamic acid • Basic - arginine histidine lysine • Hydroxylic - serine threonine • Amidic (containing amide group) - asparagine glutamine • http: //chemistry. gsu. edu/glactone/PDB/Amino_Acids/aa. html
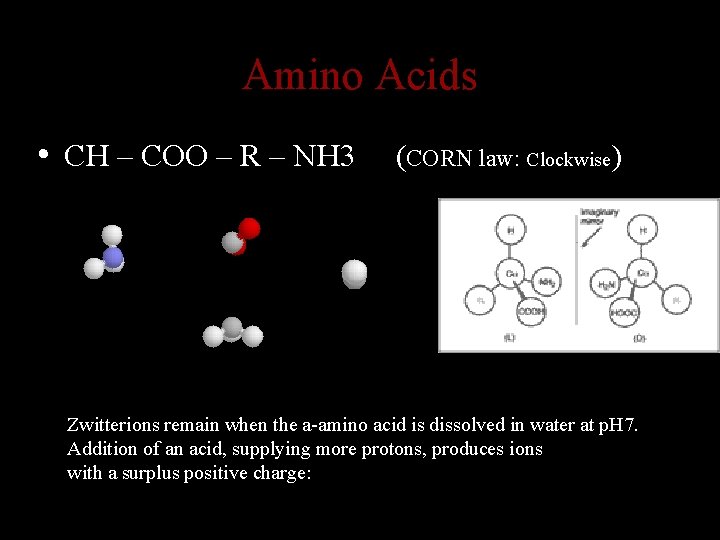
Amino Acids • CH – COO – R – NH 3 (CORN law: Clockwise) Zwitterions remain when the a-amino acid is dissolved in water at p. H 7. Addition of an acid, supplying more protons, produces ions with a surplus positive charge:

Peptide Bond
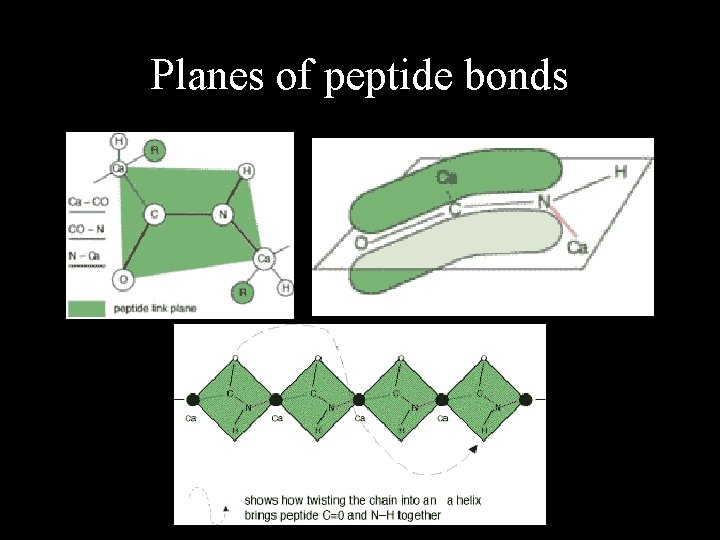
Planes of peptide bonds
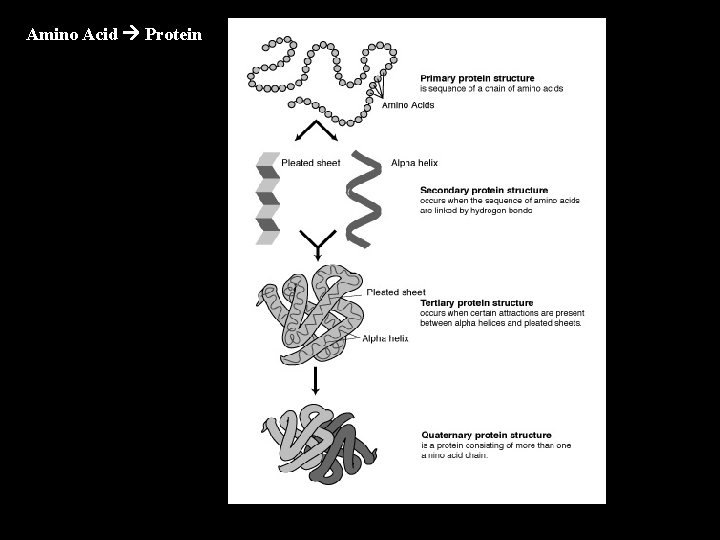
Amino Acid Protein
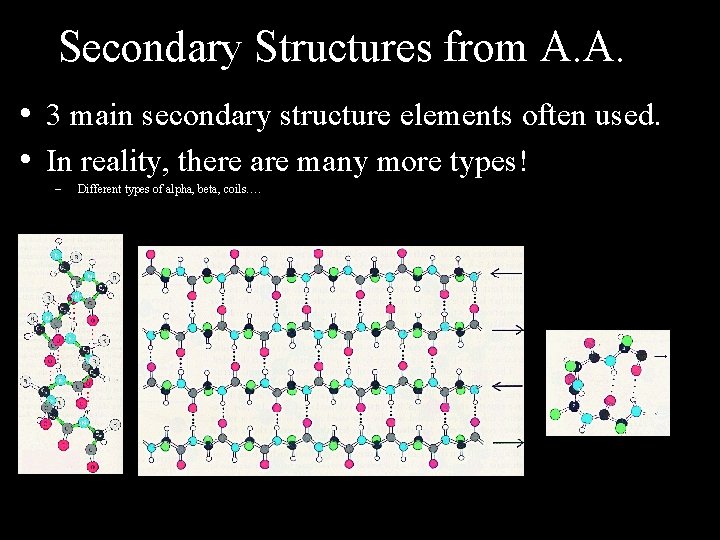
Secondary Structures from A. A. • 3 main secondary structure elements often used. • In reality, there are many more types! – Different types of alpha, beta, coils….
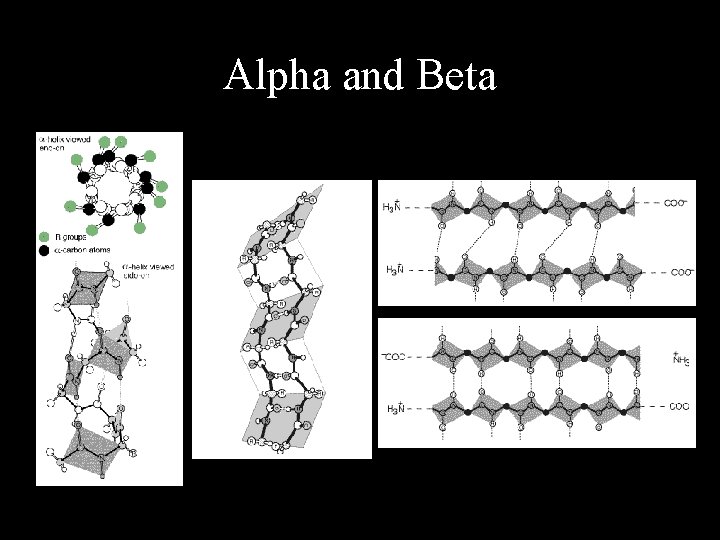
Alpha and Beta
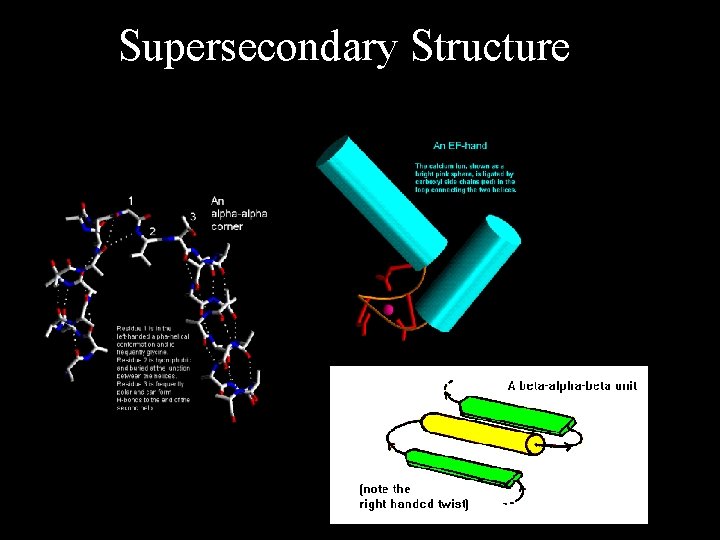
Supersecondary Structure

Basic knowledge for Bioinformatics (focusing on proteins). • Some basic points to be understood by biologists and non-biologists.

Protein • Life is a huge chunk of protein with various ligands attached. • Protein level is very efficient to work with for us a Naturally distinct unit. So, favoured by bioinformatic computing.

Only 1, 000 Protein shapes? Year 2001 • There are less than 1, 000 types of distinct shapes of protein structures known so far (called Folds)

Only 1, 000 Protein superfamilies? Year 2001 • There around 1, 000 evolutionarily distinct protein shapes (called Super. Family)

e. g) 11 very common superfamilies

Only around 10, 000 proteins? • Perhaps not much more than 10, 000 different types (representative) protein sequences in nature. *Then, where are the complexity and diversity of life come from? : Network of interactions among them. Then, where all the bio-funcitons come from? next slide

Functional diversity organizational and regulatory differences • Chimps and Humans are the same in terms of genetic components. Yet different species. • The English and Koreans: same genome but different sub-species. – Organization and regulation of information are different. Also developmental diversity. – Somehow cells ‘self-organize’ data very efficiently and effectively.

Organising Structures and Sequences bioinformatically • A technical challenge we have is how to organize the structures and sequences of proteins • There are many different ways to organize protein sequences and structures. • PDB 1976, SCOP, CATH, FSSP, , , • Swissprot, PIR, Genbank, EST, SNP, Tremble, Enssembl, … (over 500 major biological DBs) Now we have very large scale data next

Mass production of data (High throughput technology) • • • DNA/RNA Microarray Structural genomics (structure factories) Proteomics (Mass spectrometry) Interactomics (Y 2 H, protein chips based) Sequencing (5 minutes for sequencing whole human genome)

Present Bioinformatic The challenges: • • Interaction between genes Interaction between proteins Interaction between genes, proteins and ligands Networks of Networks… Wholistic understanding of the networks. Engineering genes, proteins, ligands and the whole genome.

66 Basics: Representations of protein sequences and their applications Jong Bhak Next 02/06/2001 :

Amino Acids Representation Ala alanine Met methionine Asp aspartate Phe phenylalanine Arg arginine Pro proline Asn asparagine Ser serine Cys cysteine Thr threonine Glu glutamate Trp tryptophan Gln glutamine Tyr tyrosine Gly glycine Val valine Glx glutamate or glutamine *** any His histidine --- gap of indeterminate length Ileu isoleucine TGA translation stop Lys lysine TAG translation stop Leu leucine TAA translation stop

Single Sequence representations • There are several commonly used pure sequence representation formats in “flat files” – FASTA (most commonly used for raw sequence data) – PIR • Representations in Databases (such as My. SQL) – As columns and rows • Representations in programs or objects @codons = $my. Codon. Table->revtranslate('A'); Flat file FASTA format > gi|532319|pir|TVFV 2 E envelope protein CCTCTCGGAGCTGGAAATGCAGCTATTGAGATCTTCGAATGCTGC AGCTGGAGGCAGCTGGGGAGGTCCGAGCGATGTGACC GGCCGCCATCGCTCGTCTCTTCCTCTCTCCTGCCGCCTCCTGTGT CGAAAATAACTTTTTTAGTCTAAAGAAAG >gi|532319|pir|TVFV 2 E envelope protein ELRLRYCAPAGFALLKCNDADYDGFKTNCSNVSVVHCTNLMNTTLLL SYSENRTAPTEVRRYTGGHERQKRVPFVXXXXXXX

Accessing Bioperl Codon. Table (from object oriented module) • use Bio: : Tools: : Codon. Table; • • • # defaults to ID 1 "Standard" $my. Codon. Table = Bio: : Tools: : Codon. Table->new(); $my. Codon. Table 2 = Bio: : Tools: : Codon. Table -> new ( -id => 3 ); • • # change codon table $my. Codon. Table->id(5); • • • # examine codon table print join (' ', "The name of the codon table no. ", $my. Codon. Table->id(4), "is: ", $my. Codon. Table->name(), "n"); • • # translate a codon $aa = $my. Codon. Table->translate('ACU'); $aa = $my. Codon. Table->translate('act'); $aa = $my. Codon. Table->translate('ytr'); • • • # reverse translate an amino acid @codons = $my. Codon. Table->revtranslate('A'); @codons = $my. Codon. Table->revtranslate('Ser'); @codons = $my. Codon. Table->revtranslate('Glx'); @codons = $my. Codon. Table->revtranslate('c. YS', 'rna');
- Slides: 69