1 2 Biomanufacturing Defined The production of pharmaceutical
1
2

Biomanufacturing Defined The production of pharmaceutical proteins using genetically engineered cells 3

Expression Choices Cell type: • E. coli • Yeast • Mammalian – CHO 4
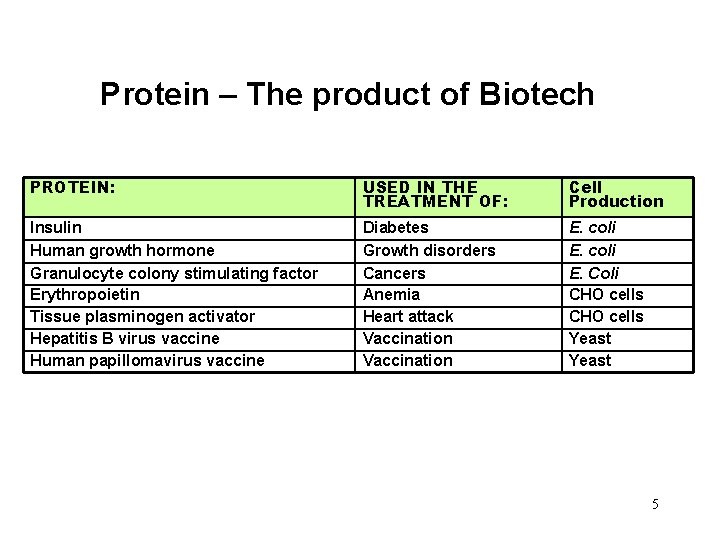
Protein – The product of Biotech PROTEIN: USED IN THE TREATMENT OF: Cell Production Insulin Human growth hormone Granulocyte colony stimulating factor Erythropoietin Tissue plasminogen activator Hepatitis B virus vaccine Human papillomavirus vaccine Diabetes Growth disorders Cancers Anemia Heart attack Vaccination E. coli E. Coli CHO cells Yeast 5

The Value of Proteins Price Per Gram Bovine Growth Hormone $14 Gold $48 Insulin $60 Growth Hormone $227, 000 Granulocyte Colony Stimulating Factor (GCSF) $1, 357, 000 *Prices in 2011 US Dollars 6
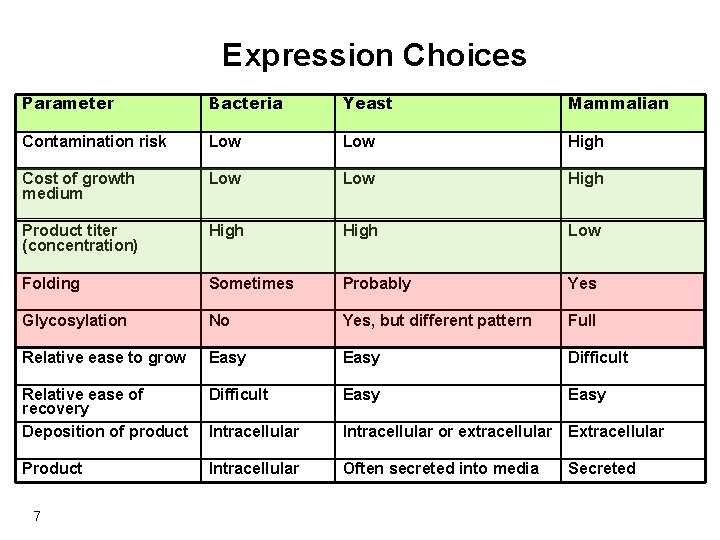
Expression Choices Parameter Bacteria Yeast Mammalian Contamination risk Low High Cost of growth medium Low High Product titer (concentration) High Low Folding Sometimes Probably Yes Glycosylation No Yes, but different pattern Full Relative ease to grow Easy Difficult Relative ease of recovery Deposition of product Difficult Easy Intracellular or extracellular Extracellular Product Intracellular Often secreted into media Secreted 7
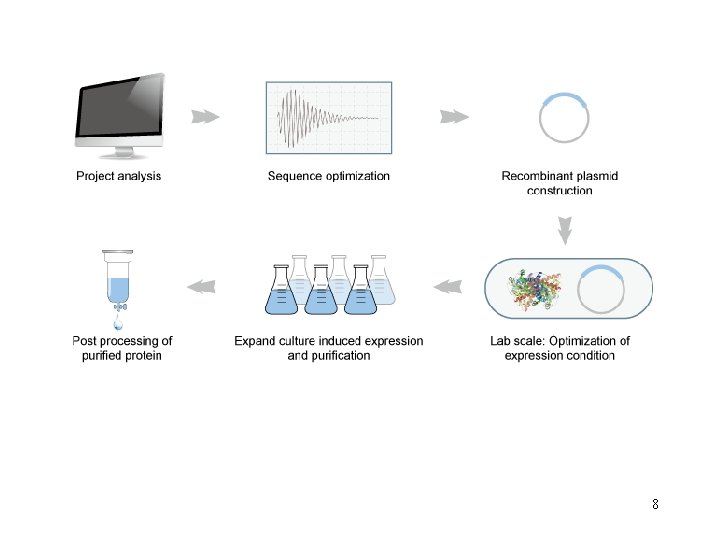
8

Choice of host Each host system has its own advantages and disadvantages, and it is important to consider these before final selection of host. Expression in different parts of the cell will lead to varying amounts of cellular (contaminant) proteins that will need to be removed to obtain a pure target protein. 9

10

Induction Biotech companies genetically engineer plasmids to place genes behind inducible promoters 11
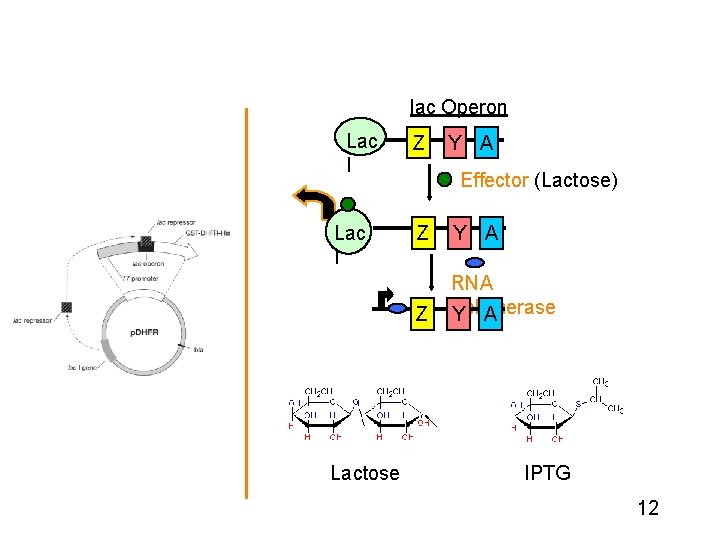
lac Operon Lac I Transcriptional Regulation in the p. DHFR system Lactose Z Y A Effector (Lactose) Z Y A Z RNA Polymerase Y A IPTG 12
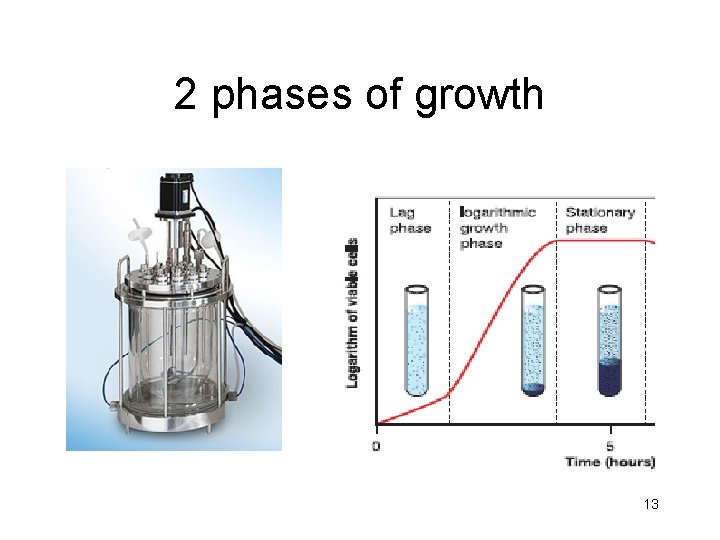
2 phases of growth 13
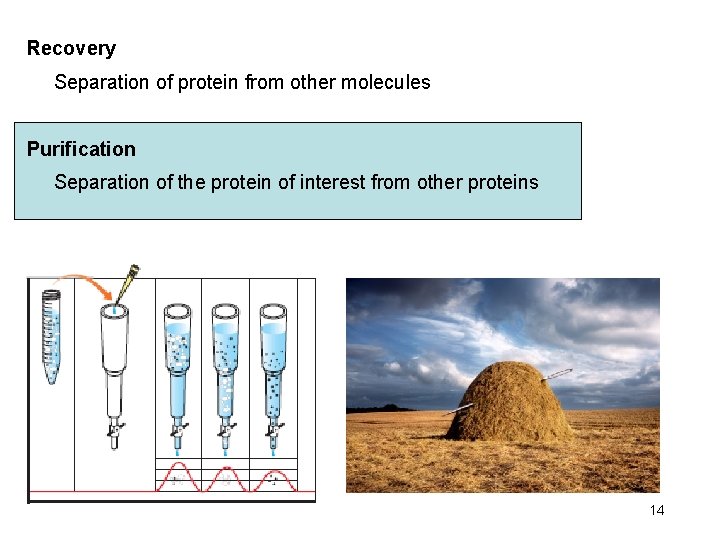
Recovery Separation of protein from other molecules Purification Separation of the protein of interest from other proteins 14

Steps in Recombinant Protein Purification 1. Design expression plasmid, transform, select 2. Grow culture of positive clone, induce expression 3. Lyse cells 4. Centrifuge to isolate protein-containing fraction 5. Column Chromatography—collect fractions 6. Assess purity on SDS-PAGE
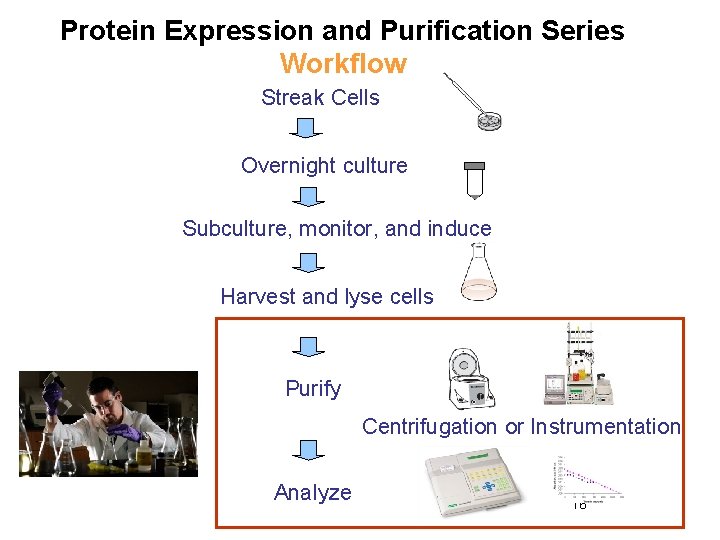
Protein Expression and Purification Series Workflow Streak Cells Overnight culture Subculture, monitor, and induce Harvest and lyse cells Purify Centrifugation or Instrumentation Analyze 16

Cell lysis: rupture cell wall / plasma membrane, --> release contents (organelles, proteins…) 1. Physical means 2. Sonication 3. Osmotic shock

Chromatography Basics • Mobile phase (solvent and the molecules to be separated) • Stationary phase (through which the mobile phase travels) – paper (in paper chromatography) – glass, resin, or ceramic beads (in column chromatography) • Molecules travel through the stationary phase at 18 different rates because of their chemistry.

Performing the chromatographic separation • Gravity Chromatography • Spin Column Chromatography • Chromatography Instrumentation • Small scale • Biomanufacturing scale (bioreactors) 19

20

Protein purification – column chromatography -Protein mixture applied to column -Solvent (buffer) applied to top, flowed through column - Different proteins interact with matrix to different extents, flow at different rates -Proteins collected separately in different fractions

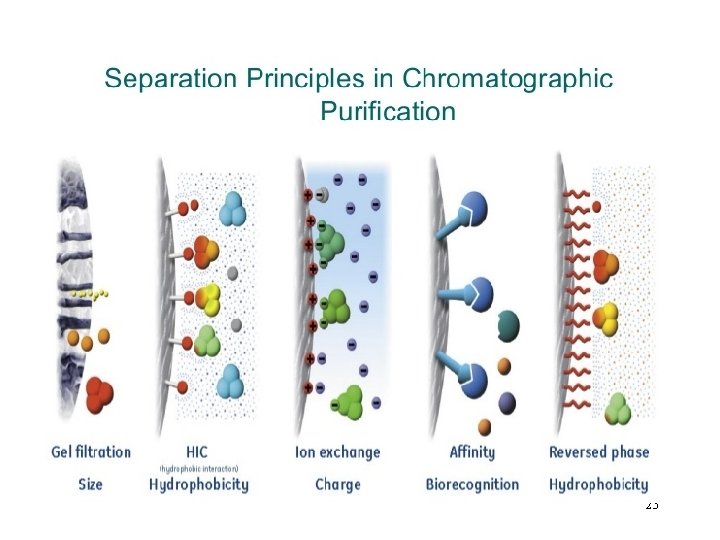
Column Chromatography Molecules can be separated on the basis of: 1. SIZE—Gel filtration 2. CHARGE—Ion exchange 3. SPECIFIC BINDING—Affinity
23

Types of Column Chromatography • Ion Exchange (protein charge) • Size Exclusion (separates on size) • Hydrophobic Interaction (hydrophobicity) • Affinity: • Protein A • His-tagged • Glutathione-s-transferase • …. . 24
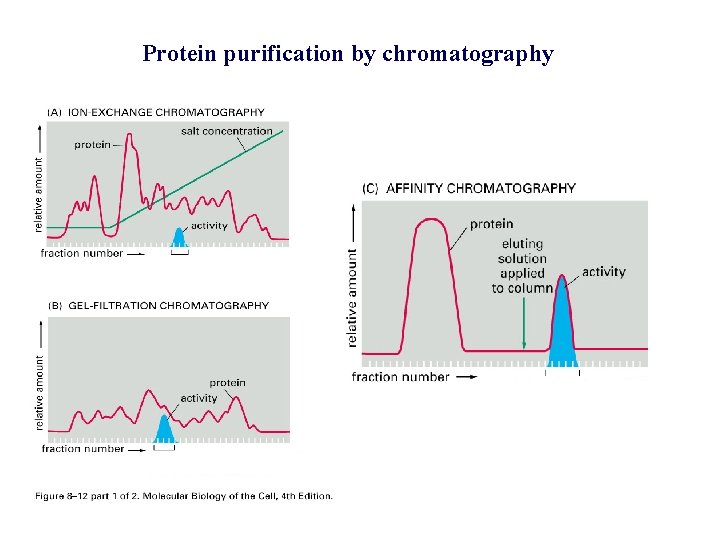
Protein purification by chromatography
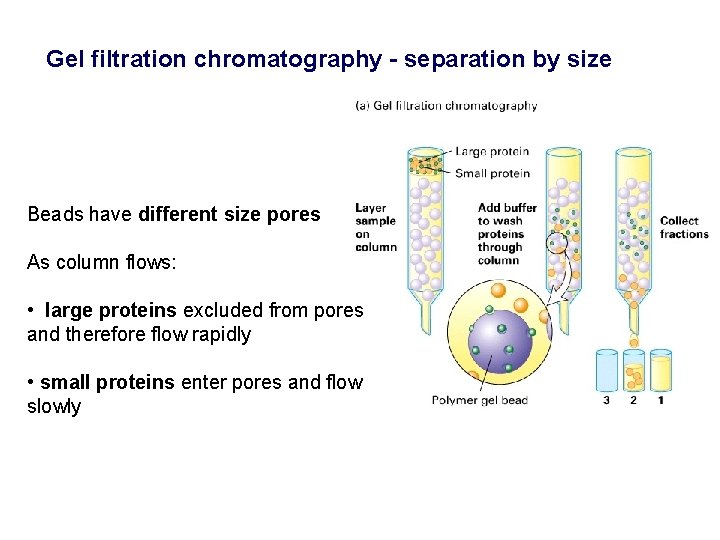
Gel filtration chromatography - separation by size Beads have different size pores As column flows: • large proteins excluded from pores and therefore flow rapidly • small proteins enter pores and flow slowly

What is Affinity chromatography (AC) ? • Affinity chromatography (AC) is a technique enabling purification of a biomolecule with respect to biological function or individual chemical structure. • AC is designed to purify a particular molecule from a mixed sample.
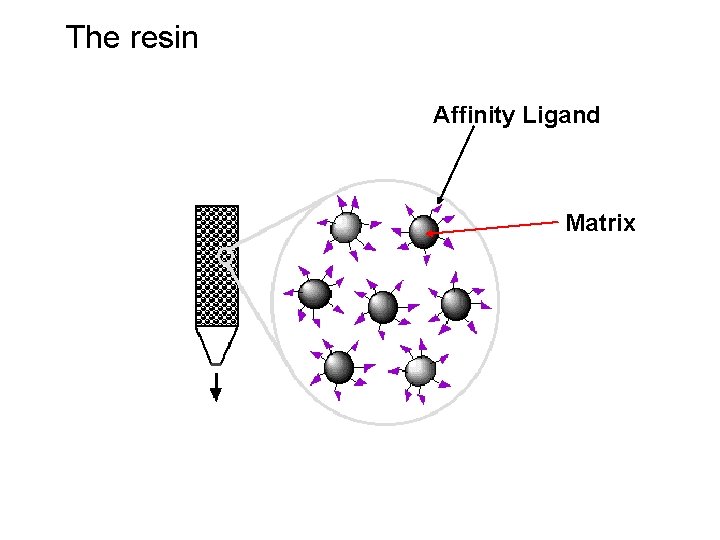
The resin Affinity Ligand Matrix

Examples of tags and ligands • • • His-tag FLAGTM peptide Strep-tag GST tag Maltose binding protein fusion (MBP) • Calmodulin binding protein fusion • • • Transition metal ion Monoclonal antibody Biotin Glutathione Amylose Ca 2+ There are situations where you don’t need a tag.
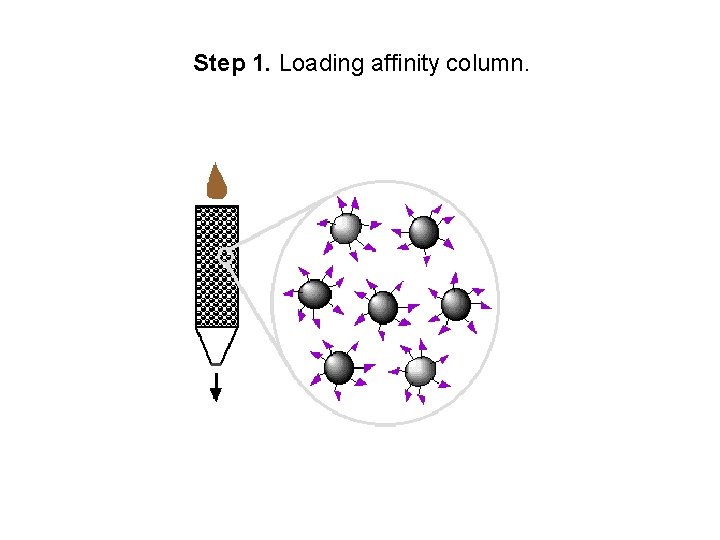
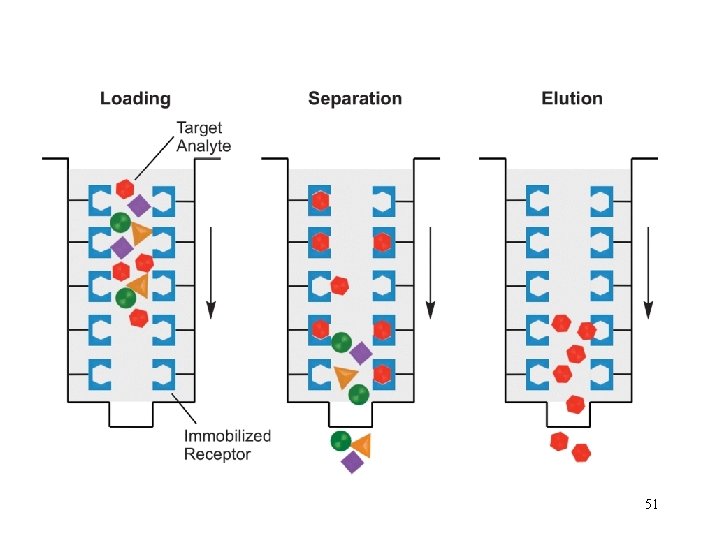
Step 1. Loading affinity column.
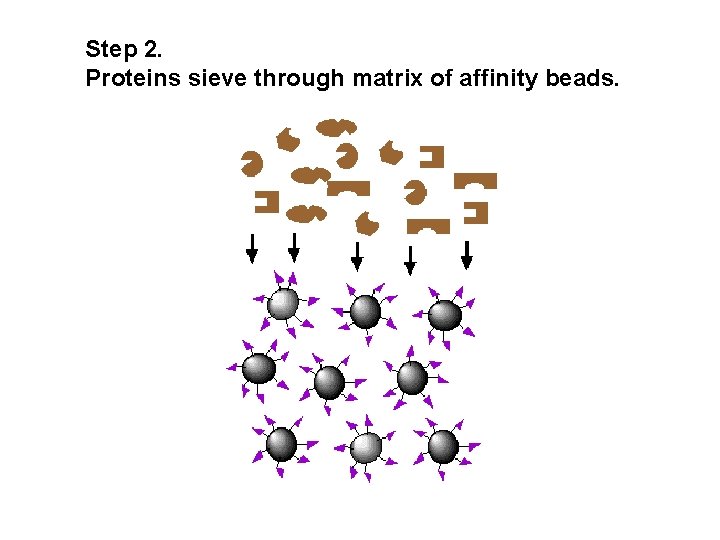
Step 2. Proteins sieve through matrix of affinity beads.
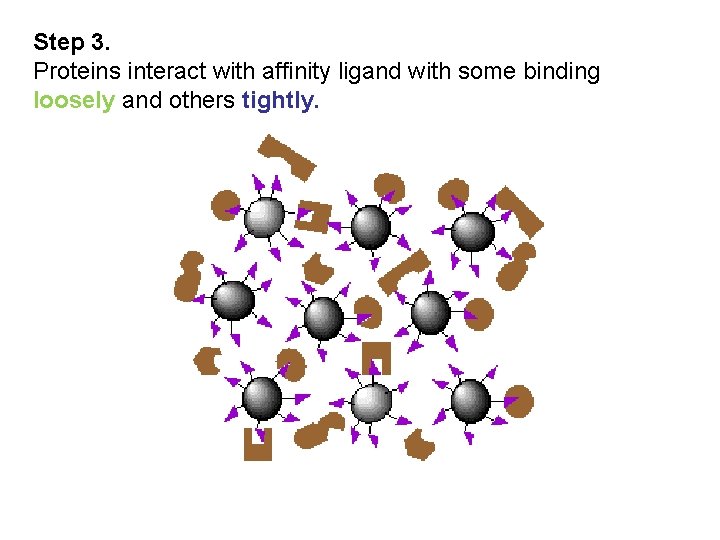
Step 3. Proteins interact with affinity ligand with some binding loosely and others tightly.
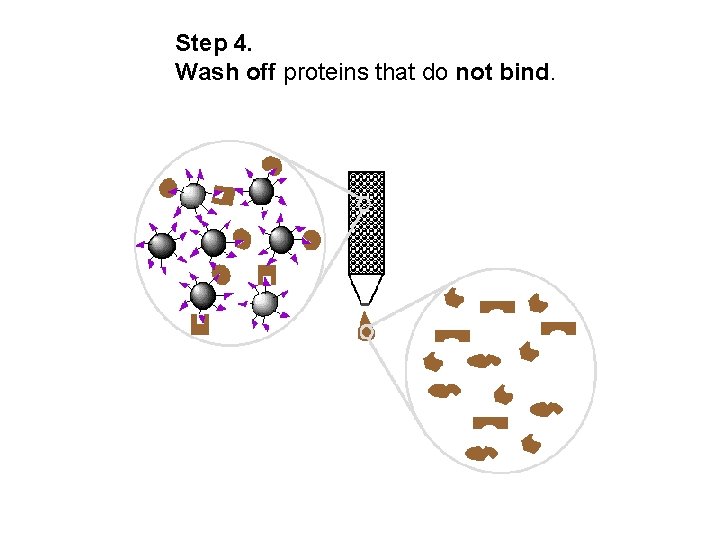
Step 4. Wash off proteins that do not bind.
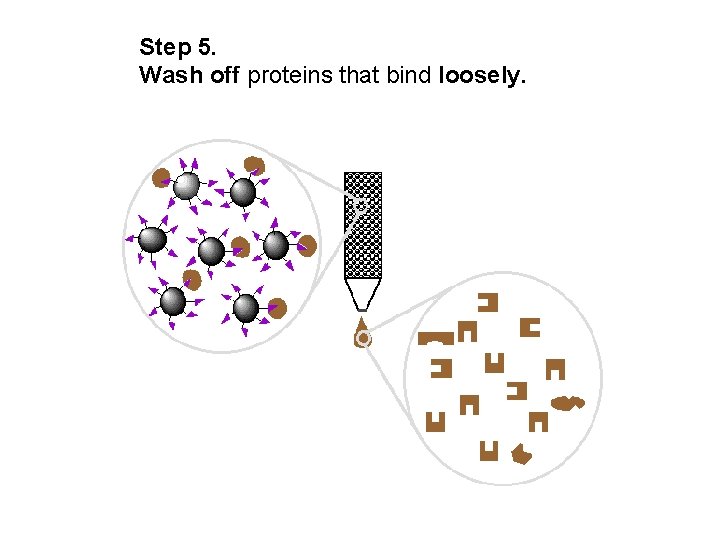
Step 5. Wash off proteins that bind loosely.
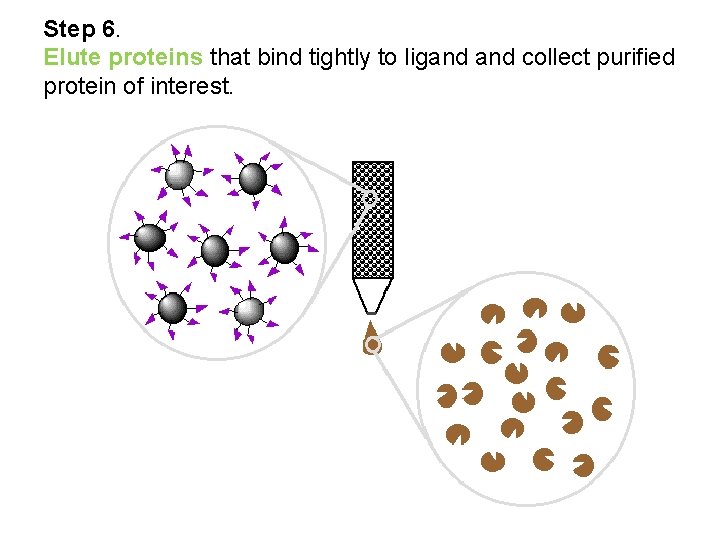
Step 6. Elute proteins that bind tightly to ligand collect purified protein of interest.
36
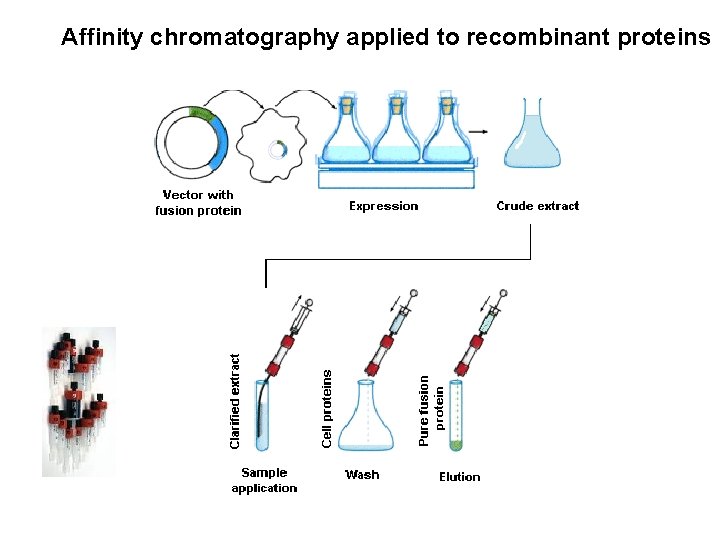
Affinity chromatography applied to recombinant proteins
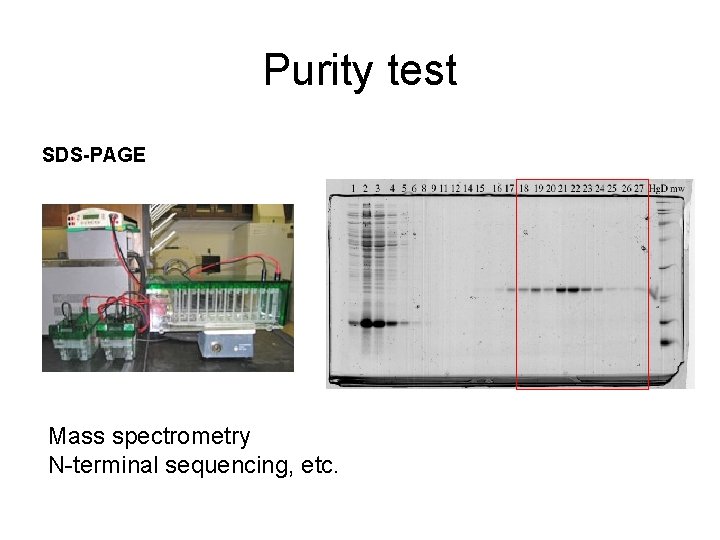
Purity test SDS-PAGE Mass spectrometry N-terminal sequencing, etc.

Downstream of protein purification • • • Biophysical characterization Biochemical analysis of activities Physiological relevance Pathological mechanisms etc.

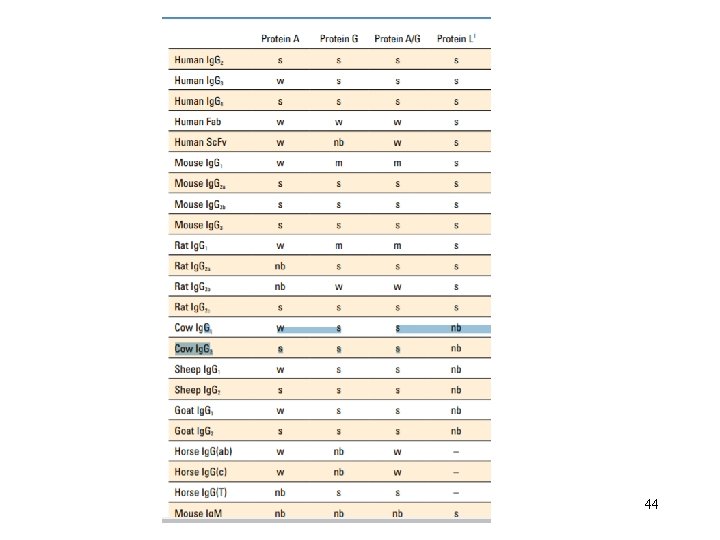
Antibody Purification Methods Antiserum from an immunized animal can be used directly for certain applications, but more often some form of antibody purification is required to obtain an antibody probe that is effective for multiple types of detection methods. • Physicochemical fractionation —differential precipitation, size-exclusion or solid-phase binding of immunoglobulins based on size, charge or other shared chemical characteristics of antibodies in typical samples. This isolates a subset of sample proteins that includes the immunoglobulins. • Antigen-specific affinity • —affinity purification of only those antibodies in a sample that bind to a particular antigen molecule through their specific antigen-binding domains. • This purifies all antibodies that bind the antigen without regard to antibody class or isotype. 40
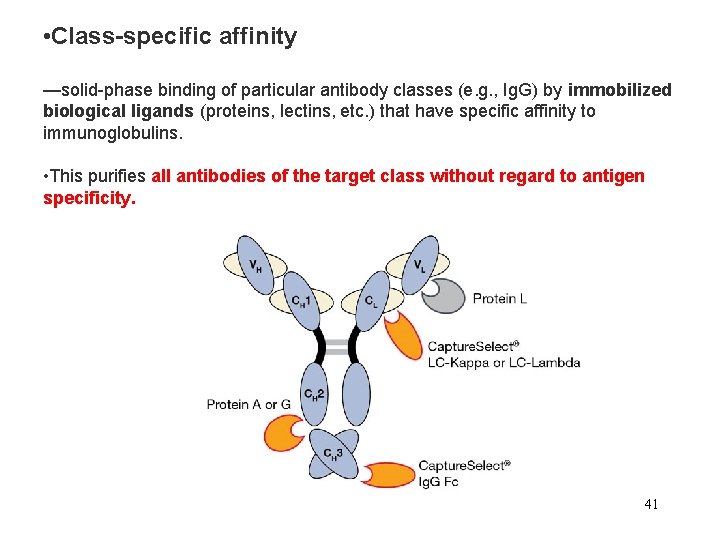
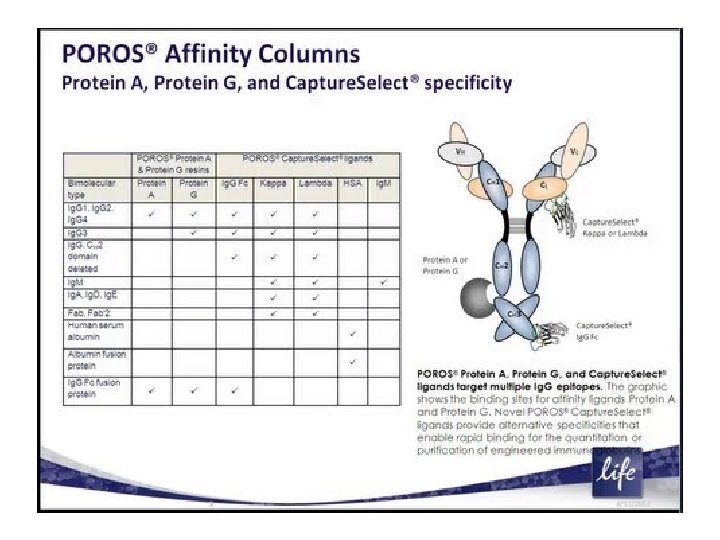
• Class-specific affinity —solid-phase binding of particular antibody classes (e. g. , Ig. G) by immobilized biological ligands (proteins, lectins, etc. ) that have specific affinity to immunoglobulins. • This purifies all antibodies of the target class without regard to antigen specificity. 41
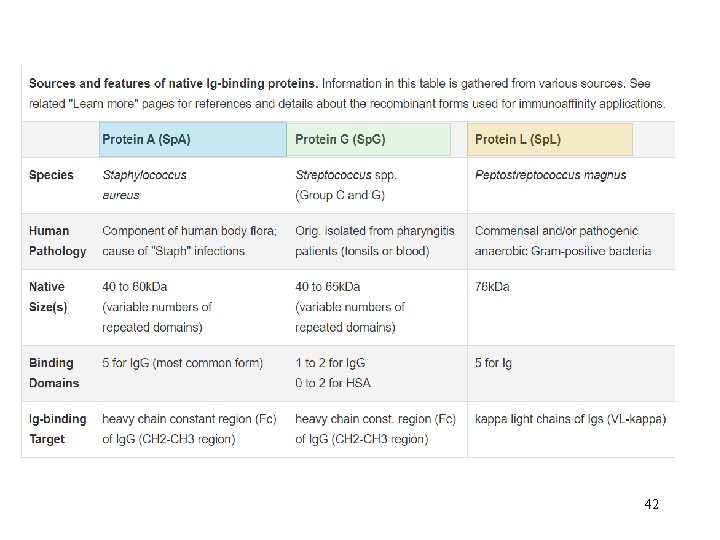
42

1. Bind: Add a clarified, physiologic-buffered (p. H 7 to 8) sample of rabbit serum to the column and allow it to slowly pass through or mix with the Protein A resin to allow the Ig. G to bind to the immobilized ligand. 2. Wash: Add phosphate-buffered saline (PBS) and allow it to pass through the column to wash away nonbound serum components. Use a volume of wash buffer equivalent to 5 to 10 times the resin volume. 3. Elute: Add acidic elution buffer (e. g. , 0. 1 M glycine-HCl, p. H 2. 8), and collect small fractions of solution that pass from the column. The low-p. H condition dissociates the antibody from the immobilized Protein A, and the Ig. G is recovered in its purified state in one or several of the collected fractions. 4. Neutralize or exchange buffer: Use a protein assay or other means to identify and combine elution fractions that contain the purified antibody. Then add 1/10 th volume of 1 M Tris-HCl (p. H 8. 5) to neutralize the buffer. 43
44
45
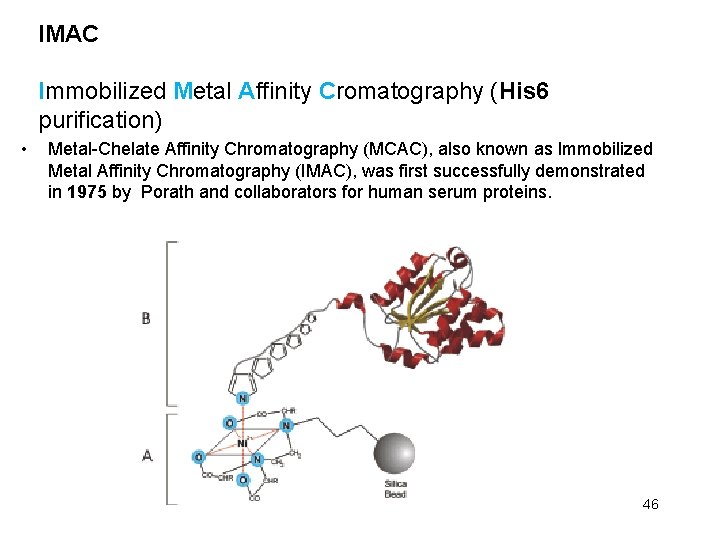
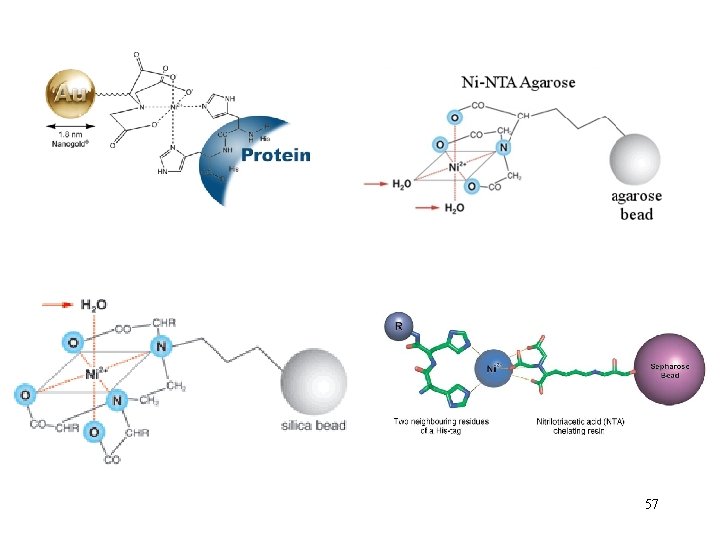
IMAC Immobilized Metal Affinity Cromatography (His 6 purification) • Metal-Chelate Affinity Chromatography (MCAC), also known as Immobilized Metal Affinity Chromatography (IMAC), was first successfully demonstrated in 1975 by Porath and collaborators for human serum proteins. 46
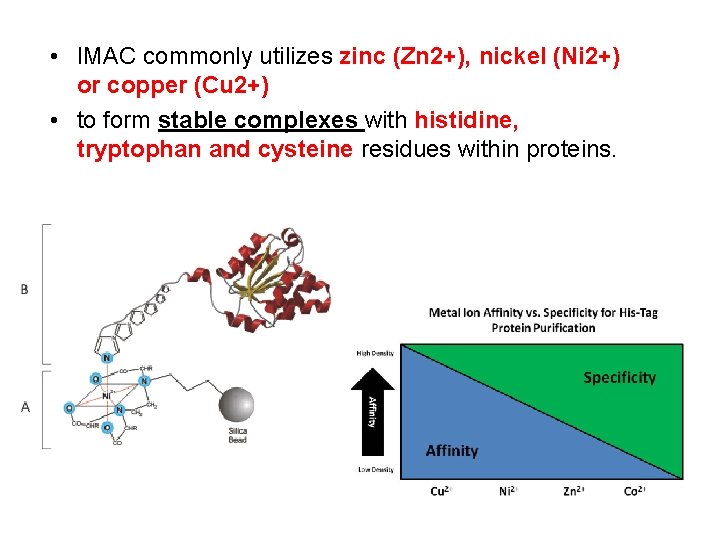
• IMAC commonly utilizes zinc (Zn 2+), nickel (Ni 2+) or copper (Cu 2+) • to form stable complexes with histidine, tryptophan and cysteine residues within proteins.
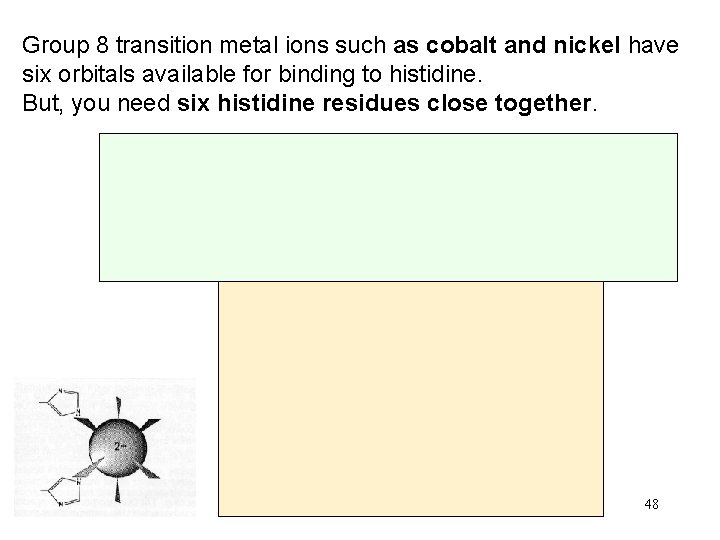
Group 8 transition metal ions such as cobalt and nickel have six orbitals available for binding to histidine. But, you need six histidine residues close together. 48
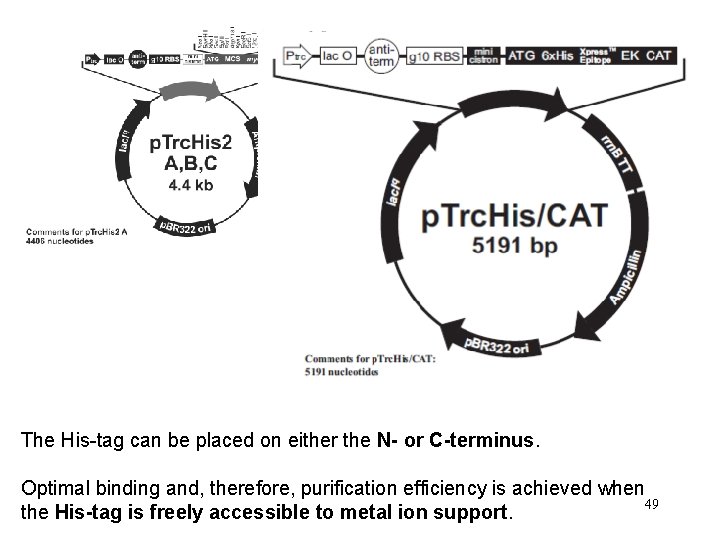
The His-tag can be placed on either the N- or C-terminus. Optimal binding and, therefore, purification efficiency is achieved when 49 the His-tag is freely accessible to metal ion support.
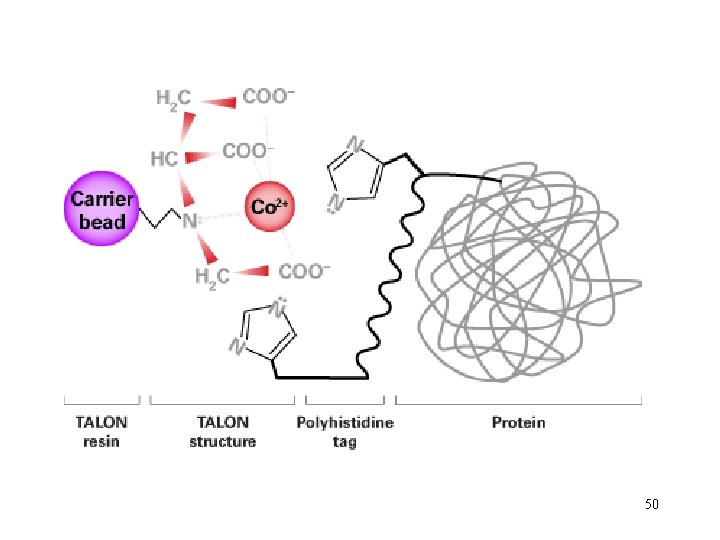
50
51

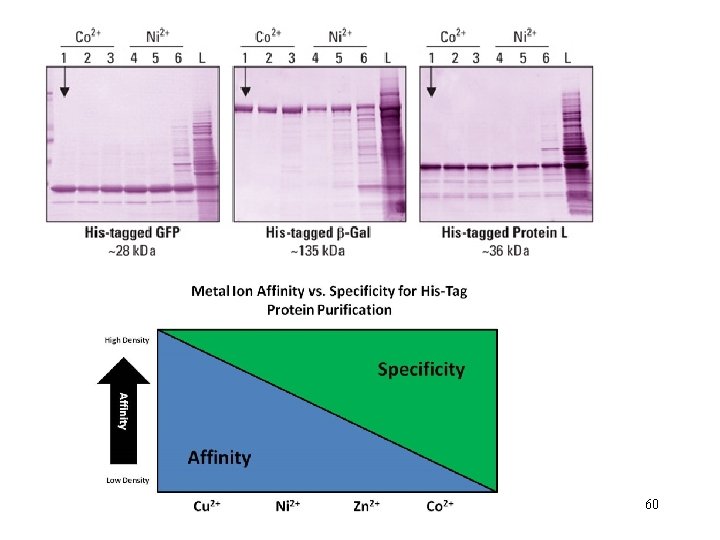
Key Parameters for the Operation of IMAC • The p. H is critical for initial binding and subsequent elution of bound proteins. • binding occurs at neutral or slightly alkali p. H (6. 5 - 8. 0) • elution generally occurs under acidic environments (< 6. 0). Chelating agents, such as ethylenediaminetetracetic acid (EDTA) and ethylene glycolbis(β-aminoethyl ether) N, N, N’, -tetraacetic acid (EGTA), must be excluded from all solutions because they will strip the metal ions from the matrix.
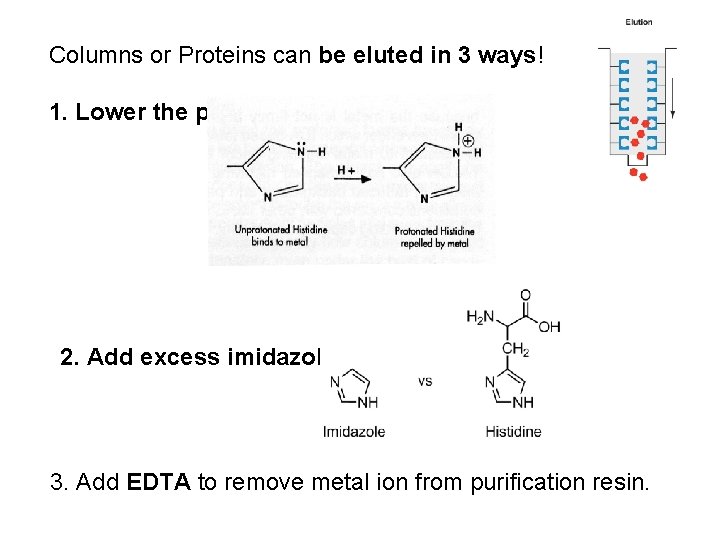
Columns or Proteins can be eluted in 3 ways! 1. Lower the p. H. 2. Add excess imidazole 3. Add EDTA to remove metal ion from purification resin.
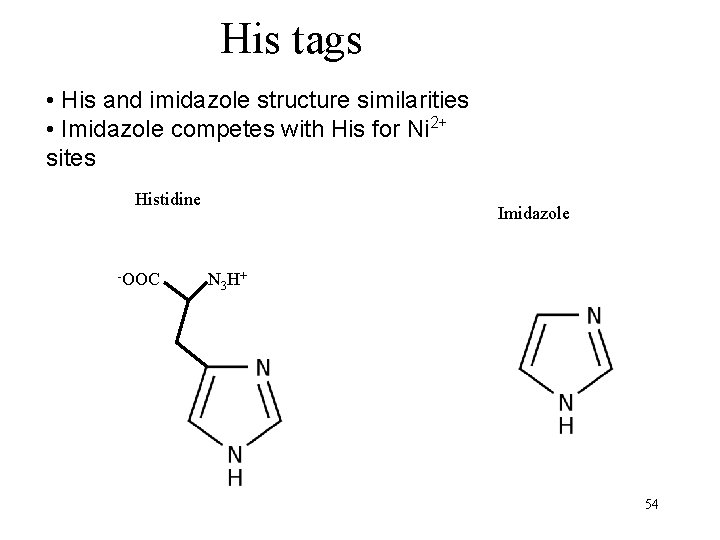
His tags • His and imidazole structure similarities • Imidazole competes with His for Ni 2+ sites Histidine -OOC Imidazole N 3 H+ 54
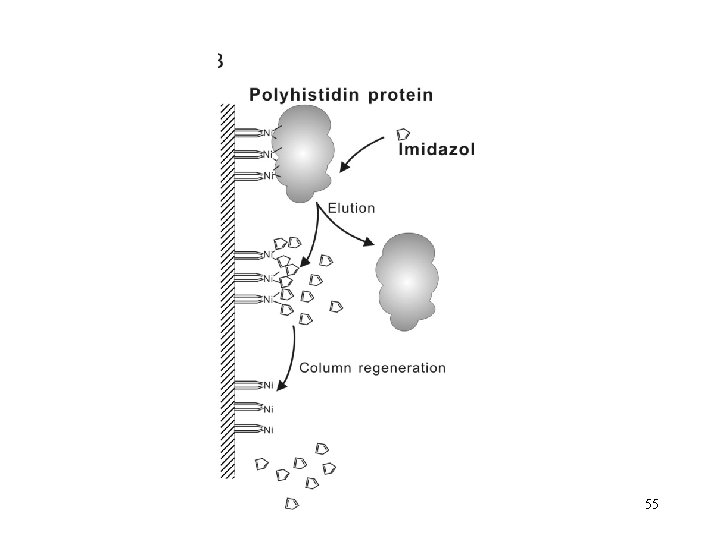
55
56
57

58

59
60

IMAC ADVANTAGES - the small size of the affinity ligand. Due to the small size, it has minimal effects on the folding of the protein. - if the His-tag is placed on the N-terminal end of the protein, it can easily be removed using an endoprotease. - Another advantage of using His-tag purification methods is that polyhistidine tags can bind proteins under both native and denaturing conditions. The use of denaturing conditions becomes important when proteins are found in inclusion bodies and must be denatured so they can be solubilized. 61

IMAC DISADVANTAGES - potential degradation of the Histag when a few histidine residues are proteolytically degraded, the affinity of the tagged protein is greatly reduced leading to a decrease in the protein yield. - dimer and tetramer formation, once a protein has a His-tag added to its structure, it has the potential to form dimers and tetramers in the presence of metal ions. While this is often not a large problem, it can lead to inaccurate molecular mass estimates of the tagged protein. - coelution of other histidine-containing proteins. coelution of proteins that naturally have two or more adjacent histidine residues. 62

63

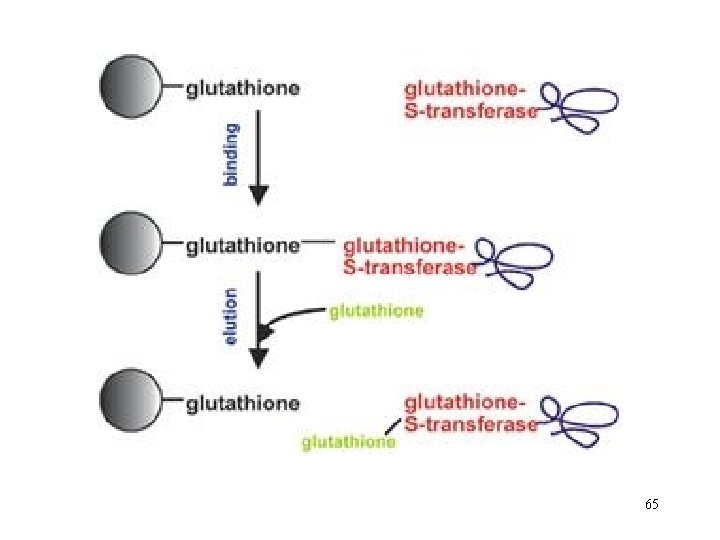
Glutathione S-transferase (GST) Gene Fusion System The Glutathione S-transferase (GST) Gene Fusion System is a versatile system for the expression, purification, and detection of fusion proteins produced in E. coli. The system is based on inducible, high-level expression of genes or gene fragments as fusions with Schistosoma japonicum GST occurs naturally as a Mr 26. 000 protein that can be expressed in E. coli with full enzymatic activity Expression in E. coli yields fusion proteins with the GST moiety at the amino terminus and the protein of interest at the carboxyl terminus. The protein accumulates within the cell’s cytoplasm. 64
65
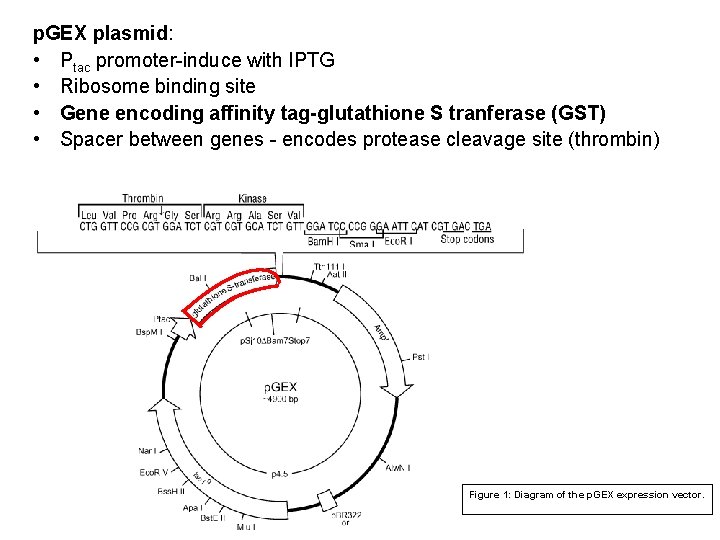
p. GEX plasmid: • Ptac promoter-induce with IPTG • Ribosome binding site • Gene encoding affinity tag-glutathione S tranferase (GST) • Spacer between genes - encodes protease cleavage site (thrombin) Figure 1: Diagram of the p. GEX expression vector.
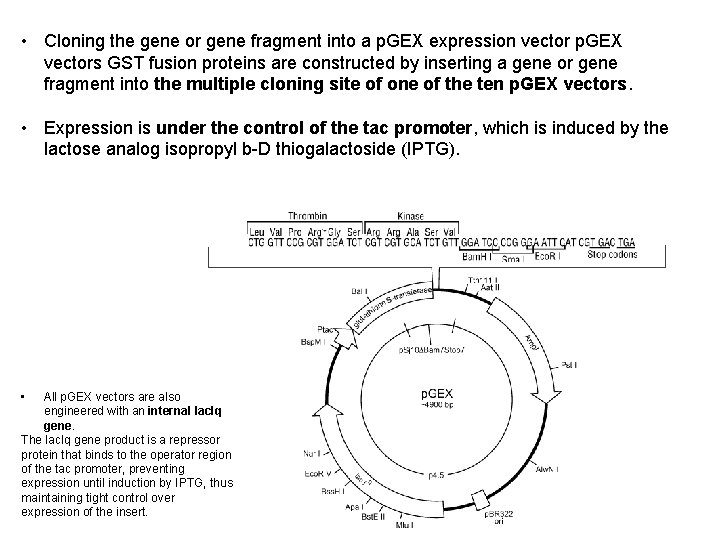
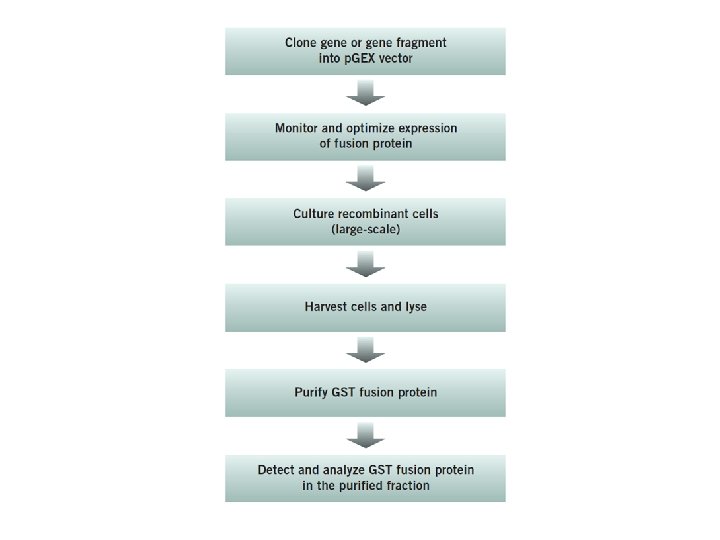
• Cloning the gene or gene fragment into a p. GEX expression vector p. GEX vectors GST fusion proteins are constructed by inserting a gene or gene fragment into the multiple cloning site of one of the ten p. GEX vectors. • Expression is under the control of the tac promoter, which is induced by the lactose analog isopropyl b-D thiogalactoside (IPTG). • All p. GEX vectors are also engineered with an internal lac. Iq gene. The lac. Iq gene product is a repressor protein that binds to the operator region of the tac promoter, preventing expression until induction by IPTG, thus maintaining tight control over expression of the insert. 67
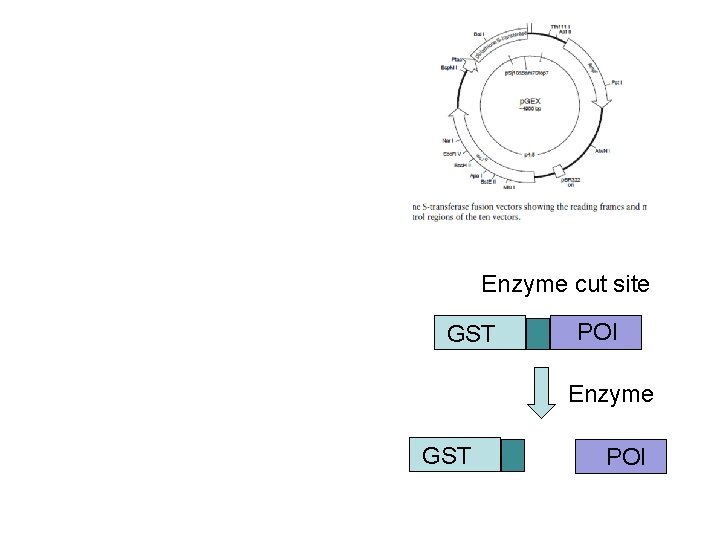
Enzyme cut site GST POI Enzyme GST POI
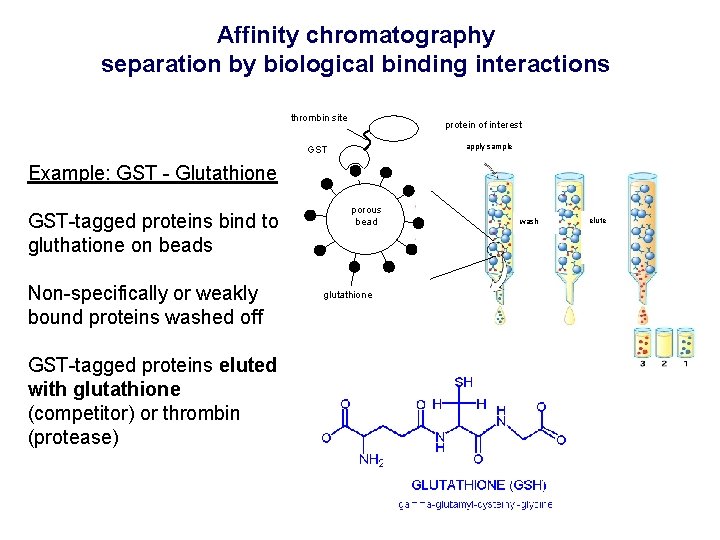
Affinity chromatography separation by biological binding interactions thrombin site protein of interest apply sample GST Example: GST - Glutathione GST-tagged proteins bind to gluthatione on beads Non-specifically or weakly bound proteins washed off GST-tagged proteins eluted with glutathione (competitor) or thrombin (protease) porous bead glutathione wash elute

70

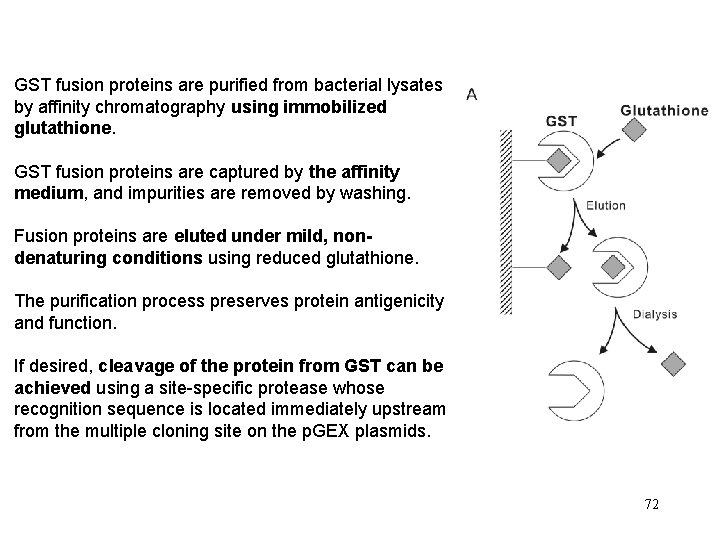
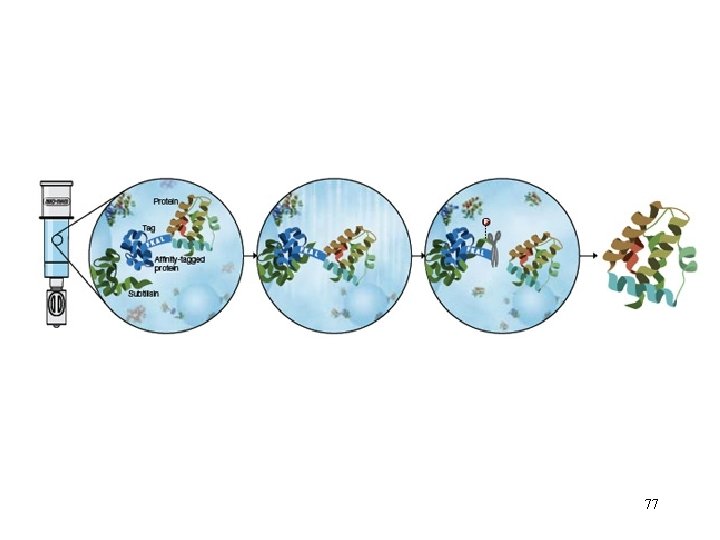
GST fusion proteins are purified from bacterial lysates by affinity chromatography using immobilized glutathione. GST fusion proteins are captured by the affinity medium, and impurities are removed by washing. Fusion proteins are eluted under mild, nondenaturing conditions using reduced glutathione. The purification process preserves protein antigenicity and function. If desired, cleavage of the protein from GST can be achieved using a site-specific protease whose recognition sequence is located immediately upstream from the multiple cloning site on the p. GEX plasmids. 72


74

75

77
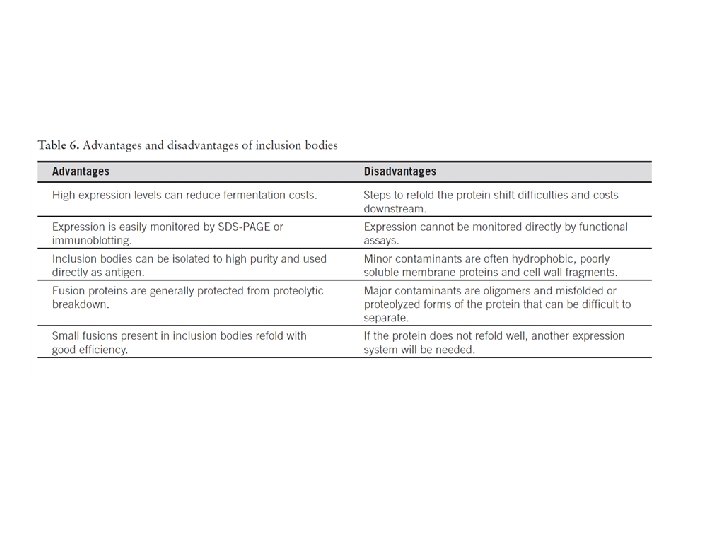
80
81
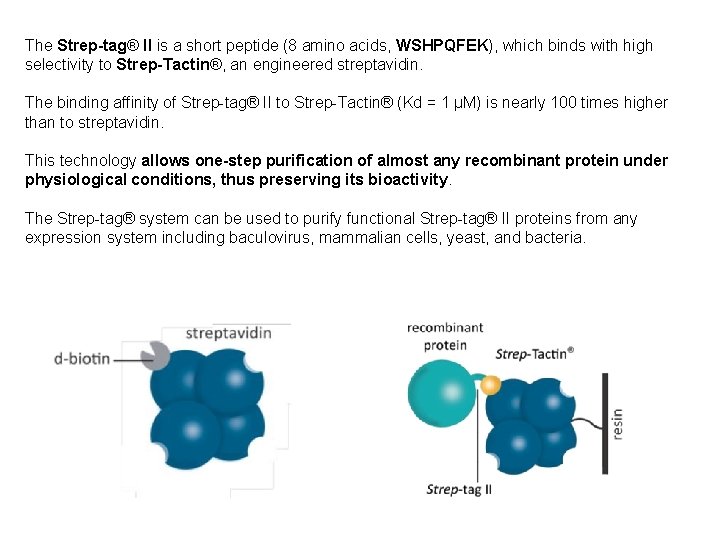
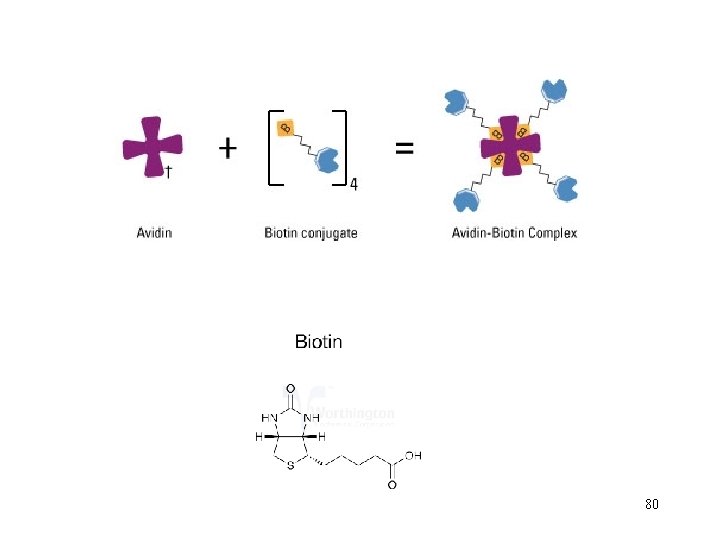
The Strep-tag® II is a short peptide (8 amino acids, WSHPQFEK), which binds with high selectivity to Strep-Tactin®, an engineered streptavidin. The binding affinity of Strep-tag® II to Strep-Tactin® (Kd = 1 µM) is nearly 100 times higher than to streptavidin. This technology allows one-step purification of almost any recombinant protein under physiological conditions, thus preserving its bioactivity. The Strep-tag® system can be used to purify functional Strep-tag® II proteins from any expression system including baculovirus, mammalian cells, yeast, and bacteria. 82
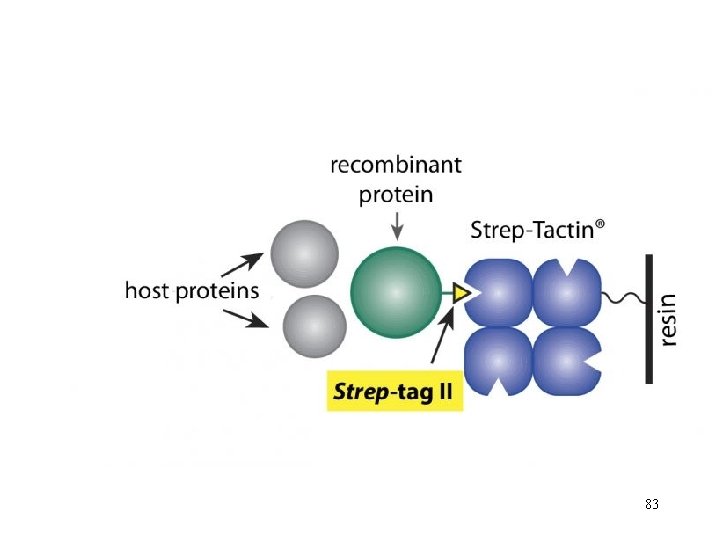
83

After application of the crude extract on a Strep-Tactin® column and a short washing step, gentle elution of purified recombinant protein is performed by addition of 84 low concentrations (2. 5 m. M) desthiobiotin.
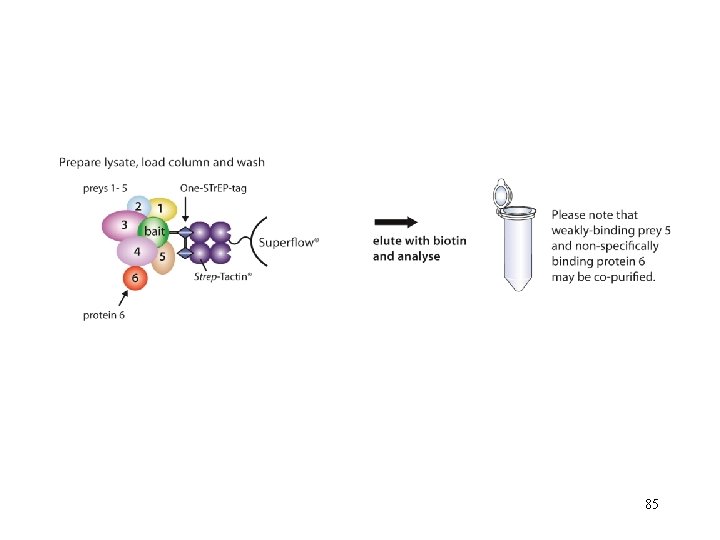
85

FLAG TAG FLAG-tag, or FLAG octapeptide, or FLAG epitope, is a polypeptide protein tag that can be added to a protein using recombinant DNA technology, having the sequence motif DYKDDDDK. It has been used for studying proteins in living cells and for protein purification by affinity chromatography. Asp. Tyr. Lys. Asp. Lys
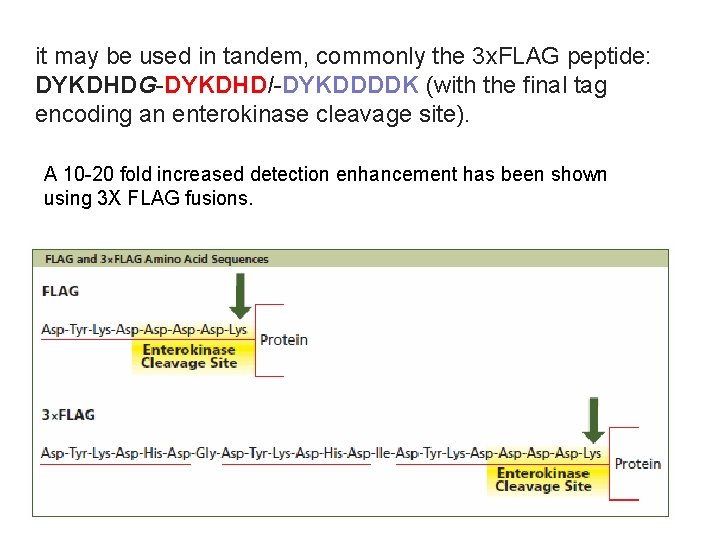
it may be used in tandem, commonly the 3 x. FLAG peptide: DYKDHDG-DYKDHDI-DYKDDDDK (with the final tag encoding an enterokinase cleavage site). A 10 -20 fold increased detection enhancement has been shown using 3 X FLAG fusions. 87

Some commercially available antibodies (e. g. , M 1/4 E 11) recognize the epitope only when it is present at the Nterminus. However, other available antibodies (e. g. , M 2) are positioninsensitive. 88

90

Bio. Logic™ LP Instrumentation Bio. Logic LP Demo Bio. Logic Duo. Flow™ 91

Scaling up of the process developed during research and development Biomanufacturing 92
- Slides: 92